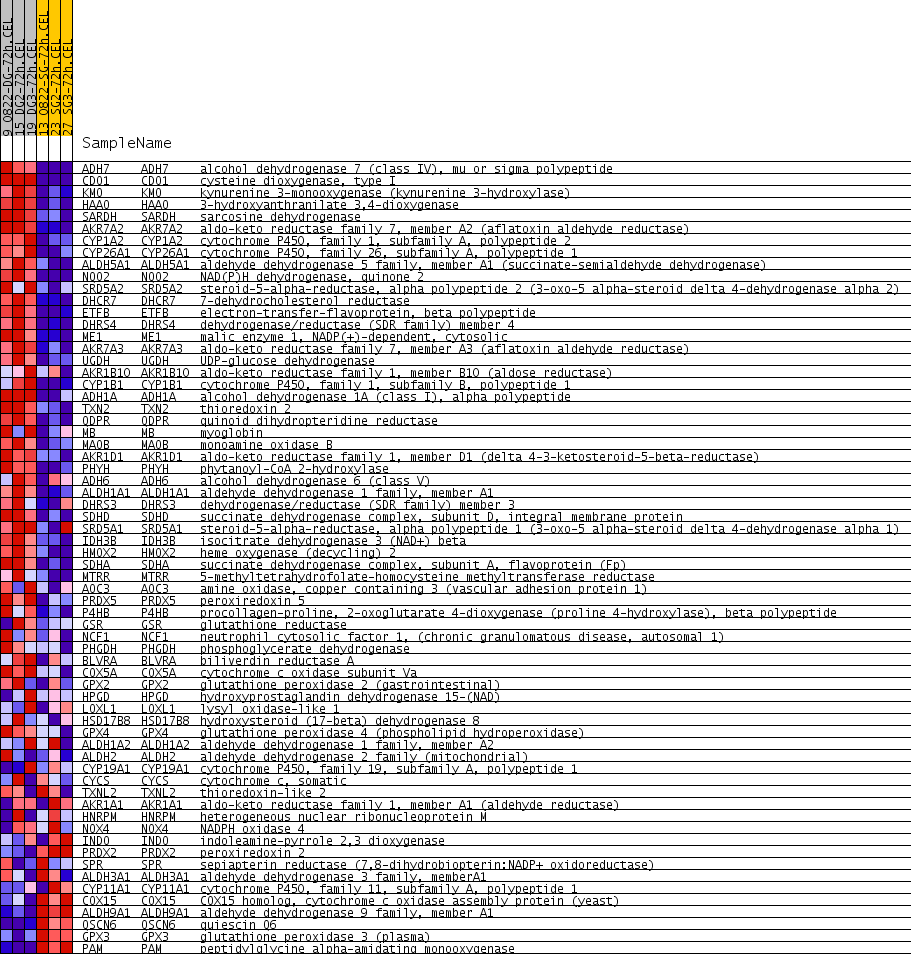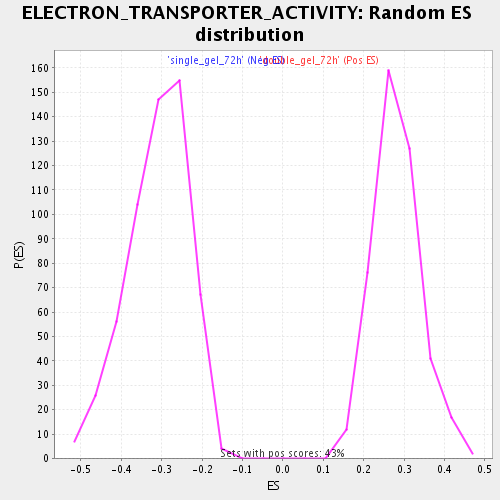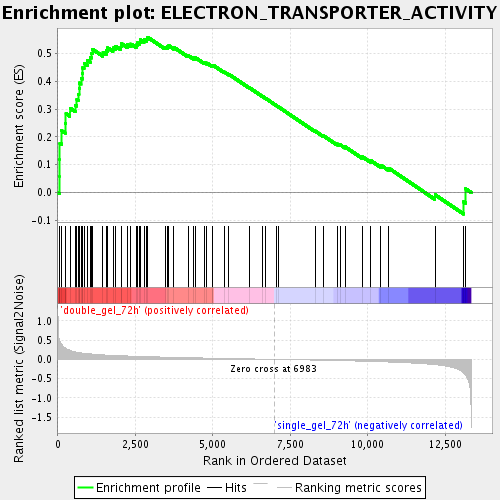
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | ELECTRON_TRANSPORTER_ACTIVITY |
| Enrichment Score (ES) | 0.5575917 |
| Normalized Enrichment Score (NES) | 1.9763005 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0038546105 |
| FWER p-Value | 0.187 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 53 | 0.495 | 0.0579 | Yes |
| 2 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 58 | 0.484 | 0.1181 | Yes |
| 3 | KMO | KMO Entrez, Source | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 64 | 0.478 | 0.1775 | Yes |
| 4 | HAAO | HAAO Entrez, Source | 3-hydroxyanthranilate 3,4-dioxygenase | 120 | 0.390 | 0.2221 | Yes |
| 5 | SARDH | SARDH Entrez, Source | sarcosine dehydrogenase | 237 | 0.291 | 0.2498 | Yes |
| 6 | AKR7A2 | AKR7A2 Entrez, Source | aldo-keto reductase family 7, member A2 (aflatoxin aldehyde reductase) | 260 | 0.283 | 0.2835 | Yes |
| 7 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 390 | 0.235 | 0.3032 | Yes |
| 8 | CYP26A1 | CYP26A1 Entrez, Source | cytochrome P450, family 26, subfamily A, polypeptide 1 | 562 | 0.191 | 0.3141 | Yes |
| 9 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.3335 | Yes |
| 10 | NQO2 | NQO2 Entrez, Source | NAD(P)H dehydrogenase, quinone 2 | 658 | 0.179 | 0.3524 | Yes |
| 11 | SRD5A2 | SRD5A2 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 2 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 2) | 681 | 0.176 | 0.3727 | Yes |
| 12 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 693 | 0.174 | 0.3936 | Yes |
| 13 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 746 | 0.168 | 0.4107 | Yes |
| 14 | DHRS4 | DHRS4 Entrez, Source | dehydrogenase/reductase (SDR family) member 4 | 788 | 0.164 | 0.4281 | Yes |
| 15 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.4476 | Yes |
| 16 | AKR7A3 | AKR7A3 Entrez, Source | aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase) | 845 | 0.158 | 0.4641 | Yes |
| 17 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.4748 | Yes |
| 18 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1044 | 0.141 | 0.4855 | Yes |
| 19 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 1078 | 0.138 | 0.5003 | Yes |
| 20 | ADH1A | ADH1A Entrez, Source | alcohol dehydrogenase 1A (class I), alpha polypeptide | 1116 | 0.136 | 0.5145 | Yes |
| 21 | TXN2 | TXN2 Entrez, Source | thioredoxin 2 | 1454 | 0.116 | 0.5036 | Yes |
| 22 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 1560 | 0.110 | 0.5095 | Yes |
| 23 | MB | MB Entrez, Source | myoglobin | 1607 | 0.109 | 0.5196 | Yes |
| 24 | MAOB | MAOB Entrez, Source | monoamine oxidase B | 1779 | 0.102 | 0.5194 | Yes |
| 25 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.5259 | Yes |
| 26 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 2036 | 0.093 | 0.5242 | Yes |
| 27 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 2053 | 0.092 | 0.5345 | Yes |
| 28 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 2239 | 0.087 | 0.5315 | Yes |
| 29 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 2353 | 0.084 | 0.5334 | Yes |
| 30 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 2529 | 0.079 | 0.5301 | Yes |
| 31 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 2561 | 0.078 | 0.5375 | Yes |
| 32 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 2648 | 0.075 | 0.5404 | Yes |
| 33 | HMOX2 | HMOX2 Entrez, Source | heme oxygenase (decycling) 2 | 2666 | 0.074 | 0.5484 | Yes |
| 34 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2780 | 0.072 | 0.5489 | Yes |
| 35 | MTRR | MTRR Entrez, Source | 5-methyltetrahydrofolate-homocysteine methyltransferase reductase | 2865 | 0.070 | 0.5513 | Yes |
| 36 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 2897 | 0.069 | 0.5576 | Yes |
| 37 | PRDX5 | PRDX5 Entrez, Source | peroxiredoxin 5 | 3457 | 0.057 | 0.5226 | No |
| 38 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 3536 | 0.055 | 0.5236 | No |
| 39 | GSR | GSR Entrez, Source | glutathione reductase | 3579 | 0.054 | 0.5272 | No |
| 40 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 3743 | 0.051 | 0.5213 | No |
| 41 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 4199 | 0.043 | 0.4924 | No |
| 42 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 4380 | 0.040 | 0.4838 | No |
| 43 | COX5A | COX5A Entrez, Source | cytochrome c oxidase subunit Va | 4442 | 0.039 | 0.4841 | No |
| 44 | GPX2 | GPX2 Entrez, Source | glutathione peroxidase 2 (gastrointestinal) | 4722 | 0.034 | 0.4673 | No |
| 45 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.4653 | No |
| 46 | LOXL1 | LOXL1 Entrez, Source | lysyl oxidase-like 1 | 4974 | 0.030 | 0.4562 | No |
| 47 | HSD17B8 | HSD17B8 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 8 | 5000 | 0.029 | 0.4580 | No |
| 48 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 5373 | 0.024 | 0.4329 | No |
| 49 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 5520 | 0.021 | 0.4246 | No |
| 50 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6175 | 0.012 | 0.3768 | No |
| 51 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 6596 | 0.005 | 0.3458 | No |
| 52 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 6707 | 0.004 | 0.3380 | No |
| 53 | TXNL2 | TXNL2 Entrez, Source | thioredoxin-like 2 | 7061 | -0.001 | 0.3115 | No |
| 54 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 7127 | -0.002 | 0.3069 | No |
| 55 | HNRPM | HNRPM Entrez, Source | heterogeneous nuclear ribonucleoprotein M | 8298 | -0.020 | 0.2213 | No |
| 56 | NOX4 | NOX4 Entrez, Source | NADPH oxidase 4 | 8566 | -0.024 | 0.2042 | No |
| 57 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 9012 | -0.032 | 0.1747 | No |
| 58 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 9109 | -0.034 | 0.1718 | No |
| 59 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 9268 | -0.037 | 0.1645 | No |
| 60 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 9824 | -0.048 | 0.1288 | No |
| 61 | CYP11A1 | CYP11A1 Entrez, Source | cytochrome P450, family 11, subfamily A, polypeptide 1 | 10102 | -0.054 | 0.1147 | No |
| 62 | COX15 | COX15 Entrez, Source | COX15 homolog, cytochrome c oxidase assembly protein (yeast) | 10424 | -0.062 | 0.0983 | No |
| 63 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 10674 | -0.069 | 0.0882 | No |
| 64 | QSCN6 | QSCN6 Entrez, Source | quiescin Q6 | 12170 | -0.136 | -0.0074 | No |
| 65 | GPX3 | GPX3 Entrez, Source | glutathione peroxidase 3 (plasma) | 13095 | -0.352 | -0.0331 | No |
| 66 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 13160 | -0.412 | 0.0137 | No |