Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | FATTY_ACID_METABOLIC_PROCESS |
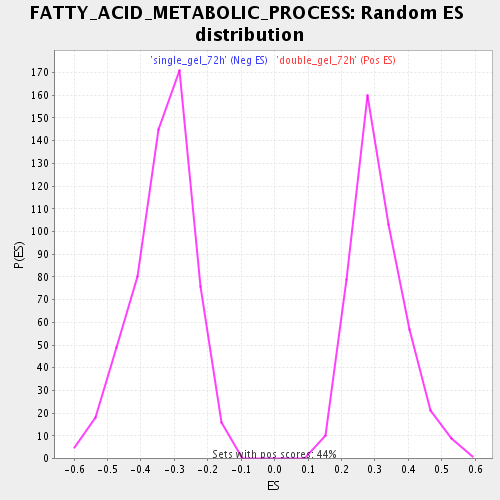
| Enrichment Score (ES) | 0.65117025 |
| Normalized Enrichment Score (NES) | 2.1093762 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.8028608E-4 |
| FWER p-Value | 0.017 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 9 | 0.779 | 0.1300 | Yes |
| 2 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 33 | 0.554 | 0.2214 | Yes |
| 3 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 54 | 0.488 | 0.3018 | Yes |
| 4 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.3466 | Yes |
| 5 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 444 | 0.218 | 0.3635 | Yes |
| 6 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 482 | 0.207 | 0.3955 | Yes |
| 7 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 536 | 0.195 | 0.4242 | Yes |
| 8 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 553 | 0.192 | 0.4552 | Yes |
| 9 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.4818 | Yes |
| 10 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 619 | 0.184 | 0.5122 | Yes |
| 11 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.5418 | Yes |
| 12 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 933 | 0.150 | 0.5445 | Yes |
| 13 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 1046 | 0.140 | 0.5597 | Yes |
| 14 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1151 | 0.133 | 0.5742 | Yes |
| 15 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 1252 | 0.127 | 0.5880 | Yes |
| 16 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 1387 | 0.119 | 0.5980 | Yes |
| 17 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1480 | 0.114 | 0.6102 | Yes |
| 18 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.6282 | Yes |
| 19 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.6449 | Yes |
| 20 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.6512 | Yes |
| 21 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 1926 | 0.096 | 0.6488 | No |
| 22 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 2496 | 0.080 | 0.6194 | No |
| 23 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | 0.6246 | No |
| 24 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 3046 | 0.066 | 0.6020 | No |
| 25 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 3063 | 0.065 | 0.6118 | No |
| 26 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3398 | 0.058 | 0.5964 | No |
| 27 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 3642 | 0.053 | 0.5870 | No |
| 28 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.5446 | No |
| 29 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | 0.5495 | No |
| 30 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.5291 | No |
| 31 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.5250 | No |
| 32 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 4887 | 0.031 | 0.5240 | No |
| 33 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 5602 | 0.020 | 0.4737 | No |
| 34 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.4572 | No |
| 35 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.4580 | No |
| 36 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | 0.3945 | No |
| 37 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.2996 | No |
| 38 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 8460 | -0.022 | 0.2716 | No |
| 39 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 9316 | -0.038 | 0.2137 | No |
| 40 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 10430 | -0.062 | 0.1404 | No |
| 41 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | 0.0887 | No |
| 42 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 11691 | -0.109 | 0.0794 | No |
| 43 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 12939 | -0.266 | 0.0303 | No |