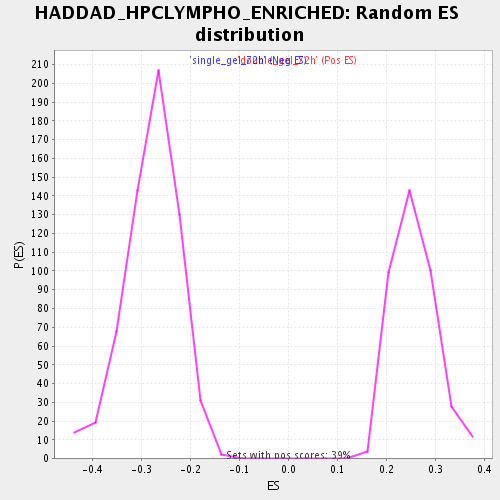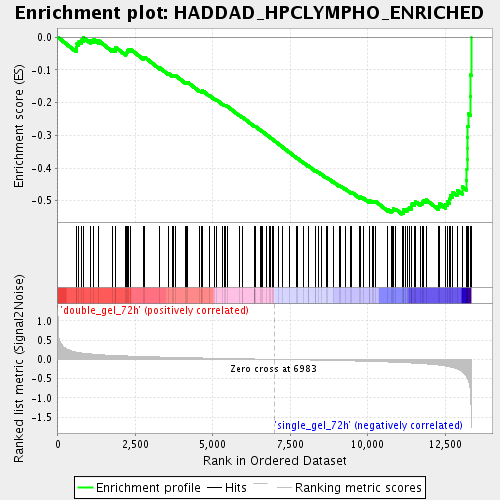
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | HADDAD_HPCLYMPHO_ENRICHED |
| Enrichment Score (ES) | -0.5408568 |
| Normalized Enrichment Score (NES) | -1.9413049 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.013291551 |
| FWER p-Value | 0.368 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 593 | 0.187 | -0.0318 | No |
| 2 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | -0.0203 | No |
| 3 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 678 | 0.176 | -0.0130 | No |
| 4 | RASGRP1 | RASGRP1 Entrez, Source | RAS guanyl releasing protein 1 (calcium and DAG-regulated) | 767 | 0.165 | -0.0081 | No |
| 5 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 829 | 0.160 | -0.0015 | No |
| 6 | CD79B | CD79B Entrez, Source | CD79b molecule, immunoglobulin-associated beta | 1051 | 0.140 | -0.0085 | No |
| 7 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 1150 | 0.133 | -0.0066 | No |
| 8 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 1307 | 0.124 | -0.0097 | No |
| 9 | QRSL1 | QRSL1 Entrez, Source | glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 | 1774 | 0.102 | -0.0379 | No |
| 10 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 1863 | 0.099 | -0.0376 | No |
| 11 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 1864 | 0.099 | -0.0307 | No |
| 12 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 2186 | 0.089 | -0.0488 | No |
| 13 | ELL3 | ELL3 Entrez, Source | elongation factor RNA polymerase II-like 3 | 2224 | 0.087 | -0.0455 | No |
| 14 | P2RX5 | P2RX5 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 5 | 2240 | 0.087 | -0.0406 | No |
| 15 | HIST1H1E | HIST1H1E Entrez, Source | histone cluster 1, H1e | 2288 | 0.085 | -0.0382 | No |
| 16 | SEC22B | SEC22B Entrez, Source | SEC22 vesicle trafficking protein homolog B (S. cerevisiae) | 2352 | 0.084 | -0.0371 | No |
| 17 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 2765 | 0.072 | -0.0632 | No |
| 18 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 2800 | 0.071 | -0.0608 | No |
| 19 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 3289 | 0.060 | -0.0935 | No |
| 20 | GH1 | GH1 Entrez, Source | growth hormone 1 | 3555 | 0.055 | -0.1097 | No |
| 21 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 3710 | 0.052 | -0.1178 | No |
| 22 | NELF | NELF Entrez, Source | nasal embryonic LHRH factor | 3738 | 0.051 | -0.1162 | No |
| 23 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 3792 | 0.050 | -0.1168 | No |
| 24 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 4112 | 0.044 | -0.1378 | No |
| 25 | LEF1 | LEF1 Entrez, Source | lymphoid enhancer-binding factor 1 | 4161 | 0.044 | -0.1384 | No |
| 26 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 4190 | 0.043 | -0.1375 | No |
| 27 | HIST1H4B | HIST1H4B Entrez, Source | histone cluster 1, H4b | 4584 | 0.036 | -0.1647 | No |
| 28 | PPM1B | PPM1B Entrez, Source | protein phosphatase 1B (formerly 2C), magnesium-dependent, beta isoform | 4644 | 0.035 | -0.1667 | No |
| 29 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 4648 | 0.035 | -0.1644 | No |
| 30 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 4666 | 0.035 | -0.1633 | No |
| 31 | CD9 | CD9 Entrez, Source | CD9 molecule | 4896 | 0.031 | -0.1785 | No |
| 32 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 5059 | 0.028 | -0.1887 | No |
| 33 | RIMS3 | RIMS3 Entrez, Source | regulating synaptic membrane exocytosis 3 | 5124 | 0.027 | -0.1917 | No |
| 34 | AKAP12 | AKAP12 Entrez, Source | A kinase (PRKA) anchor protein (gravin) 12 | 5319 | 0.024 | -0.2046 | No |
| 35 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 5391 | 0.023 | -0.2084 | No |
| 36 | P2RY14 | P2RY14 Entrez, Source | purinergic receptor P2Y, G-protein coupled, 14 | 5407 | 0.023 | -0.2079 | No |
| 37 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 5468 | 0.022 | -0.2109 | No |
| 38 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 5846 | 0.017 | -0.2383 | No |
| 39 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 5953 | 0.015 | -0.2452 | No |
| 40 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 5968 | 0.015 | -0.2453 | No |
| 41 | PPM1A | PPM1A Entrez, Source | protein phosphatase 1A (formerly 2C), magnesium-dependent, alpha isoform | 6343 | 0.009 | -0.2729 | No |
| 42 | KCNJ2 | KCNJ2 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 2 | 6345 | 0.009 | -0.2724 | No |
| 43 | SIAH2 | SIAH2 Entrez, Source | seven in absentia homolog 2 (Drosophila) | 6351 | 0.009 | -0.2722 | No |
| 44 | WFS1 | WFS1 Entrez, Source | Wolfram syndrome 1 (wolframin) | 6353 | 0.009 | -0.2716 | No |
| 45 | RPL35A | RPL35A Entrez, Source | ribosomal protein L35a | 6368 | 0.008 | -0.2721 | No |
| 46 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 6522 | 0.006 | -0.2832 | No |
| 47 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6567 | 0.006 | -0.2862 | No |
| 48 | CSH2 | CSH2 Entrez, Source | chorionic somatomammotropin hormone 2 | 6600 | 0.005 | -0.2882 | No |
| 49 | GNG7 | GNG7 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 7 | 6736 | 0.003 | -0.2982 | No |
| 50 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 6814 | 0.002 | -0.3039 | No |
| 51 | RPS11 | RPS11 Entrez, Source | ribosomal protein S11 | 6874 | 0.002 | -0.3082 | No |
| 52 | MAG | MAG Entrez, Source | myelin associated glycoprotein | 6930 | 0.001 | -0.3123 | No |
| 53 | DCK | DCK Entrez, Source | deoxycytidine kinase | 6967 | 0.000 | -0.3150 | No |
| 54 | COL5A1 | COL5A1 Entrez, Source | collagen, type V, alpha 1 | 7132 | -0.002 | -0.3273 | No |
| 55 | VDAC1 | VDAC1 Entrez, Source | voltage-dependent anion channel 1 | 7242 | -0.004 | -0.3353 | No |
| 56 | VAMP1 | VAMP1 Entrez, Source | vesicle-associated membrane protein 1 (synaptobrevin 1) | 7464 | -0.007 | -0.3515 | No |
| 57 | PPP3CC | PPP3CC Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, gamma isoform (calcineurin A gamma) | 7701 | -0.010 | -0.3686 | No |
| 58 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 7727 | -0.011 | -0.3698 | No |
| 59 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 7925 | -0.014 | -0.3837 | No |
| 60 | ABCG2 | ABCG2 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 2 | 8071 | -0.016 | -0.3935 | No |
| 61 | BLK | BLK Entrez, Source | B lymphoid tyrosine kinase | 8075 | -0.016 | -0.3927 | No |
| 62 | EFHA1 | EFHA1 Entrez, Source | EF-hand domain family, member A1 | 8302 | -0.020 | -0.4084 | No |
| 63 | TAF9B | TAF9B Entrez, Source | TAF9B RNA polymerase II, TATA box binding protein (TBP)-associated factor, 31kDa | 8303 | -0.020 | -0.4070 | No |
| 64 | DNASE2B | DNASE2B Entrez, Source | deoxyribonuclease II beta | 8411 | -0.022 | -0.4136 | No |
| 65 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8494 | -0.023 | -0.4182 | No |
| 66 | AP1B1 | AP1B1 Entrez, Source | adaptor-related protein complex 1, beta 1 subunit | 8657 | -0.026 | -0.4286 | No |
| 67 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 8704 | -0.026 | -0.4303 | No |
| 68 | PKN2 | PKN2 Entrez, Source | protein kinase N2 | 8882 | -0.030 | -0.4416 | No |
| 69 | REXO2 | REXO2 Entrez, Source | REX2, RNA exonuclease 2 homolog (S. cerevisiae) | 9073 | -0.034 | -0.4536 | No |
| 70 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9129 | -0.035 | -0.4554 | No |
| 71 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 9276 | -0.037 | -0.4638 | No |
| 72 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 9448 | -0.041 | -0.4739 | No |
| 73 | STK38 | STK38 Entrez, Source | serine/threonine kinase 38 | 9486 | -0.041 | -0.4738 | No |
| 74 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 9737 | -0.046 | -0.4895 | No |
| 75 | ZBTB10 | ZBTB10 Entrez, Source | zinc finger and BTB domain containing 10 | 9765 | -0.047 | -0.4882 | No |
| 76 | TTC13 | TTC13 Entrez, Source | tetratricopeptide repeat domain 13 | 9855 | -0.049 | -0.4915 | No |
| 77 | PCBP2 | PCBP2 Entrez, Source | poly(rC) binding protein 2 | 10047 | -0.053 | -0.5023 | No |
| 78 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 10068 | -0.053 | -0.5001 | No |
| 79 | PSPC1 | PSPC1 Entrez, Source | paraspeckle component 1 | 10148 | -0.056 | -0.5022 | No |
| 80 | PCAF | PCAF Entrez, Source | p300/CBP-associated factor | 10191 | -0.056 | -0.5014 | No |
| 81 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 10237 | -0.058 | -0.5008 | No |
| 82 | STAG3 | STAG3 Entrez, Source | stromal antigen 3 | 10647 | -0.068 | -0.5270 | No |
| 83 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 10761 | -0.072 | -0.5305 | No |
| 84 | FHIT | FHIT Entrez, Source | fragile histidine triad gene | 10806 | -0.073 | -0.5288 | No |
| 85 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 10814 | -0.073 | -0.5242 | No |
| 86 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 10903 | -0.075 | -0.5257 | No |
| 87 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 11105 | -0.082 | -0.5351 | Yes |
| 88 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 11154 | -0.084 | -0.5329 | Yes |
| 89 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 11158 | -0.084 | -0.5273 | Yes |
| 90 | NEIL1 | NEIL1 Entrez, Source | nei endonuclease VIII-like 1 (E. coli) | 11232 | -0.087 | -0.5267 | Yes |
| 91 | ELF2 | ELF2 Entrez, Source | E74-like factor 2 (ets domain transcription factor) | 11297 | -0.090 | -0.5253 | Yes |
| 92 | MOG | MOG Entrez, Source | myelin oligodendrocyte glycoprotein | 11332 | -0.092 | -0.5214 | Yes |
| 93 | SCN3A | SCN3A Entrez, Source | sodium channel, voltage-gated, type III, alpha | 11403 | -0.095 | -0.5201 | Yes |
| 94 | BLNK | BLNK Entrez, Source | B-cell linker | 11422 | -0.096 | -0.5148 | Yes |
| 95 | UXS1 | UXS1 Entrez, Source | UDP-glucuronate decarboxylase 1 | 11427 | -0.096 | -0.5083 | Yes |
| 96 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 11508 | -0.100 | -0.5074 | Yes |
| 97 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 11527 | -0.101 | -0.5018 | Yes |
| 98 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 11699 | -0.109 | -0.5071 | Yes |
| 99 | JUN | JUN Entrez, Source | jun oncogene | 11756 | -0.111 | -0.5035 | Yes |
| 100 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 11790 | -0.113 | -0.4981 | Yes |
| 101 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11886 | -0.118 | -0.4971 | Yes |
| 102 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 12294 | -0.147 | -0.5176 | Yes |
| 103 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 12319 | -0.150 | -0.5089 | Yes |
| 104 | EIF2AK3 | EIF2AK3 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 3 | 12512 | -0.172 | -0.5114 | Yes |
| 105 | HIP1R | HIP1R Entrez, Source | huntingtin interacting protein 1 related | 12571 | -0.180 | -0.5032 | Yes |
| 106 | CACNB3 | CACNB3 Entrez, Source | calcium channel, voltage-dependent, beta 3 subunit | 12633 | -0.191 | -0.4945 | Yes |
| 107 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 12657 | -0.196 | -0.4826 | Yes |
| 108 | SNN | SNN Entrez, Source | stannin | 12747 | -0.213 | -0.4744 | Yes |
| 109 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 12889 | -0.247 | -0.4678 | Yes |
| 110 | ANKRD10 | ANKRD10 Entrez, Source | ankyrin repeat domain 10 | 13047 | -0.318 | -0.4575 | Yes |
| 111 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13180 | -0.438 | -0.4369 | Yes |
| 112 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 13200 | -0.467 | -0.4057 | Yes |
| 113 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 13204 | -0.471 | -0.3731 | Yes |
| 114 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.3399 | Yes |
| 115 | DDR1 | DDR1 Entrez, Source | discoidin domain receptor family, member 1 | 13214 | -0.489 | -0.3059 | Yes |
| 116 | MME | MME Entrez, Source | membrane metallo-endopeptidase (neutral endopeptidase, enkephalinase) | 13226 | -0.505 | -0.2715 | Yes |
| 117 | NPY | NPY Entrez, Source | neuropeptide Y | 13255 | -0.563 | -0.2343 | Yes |
| 118 | CD69 | CD69 Entrez, Source | CD69 molecule | 13310 | -0.833 | -0.1802 | Yes |
| 119 | DBN1 | DBN1 Entrez, Source | drebrin 1 | 13318 | -0.939 | -0.1152 | Yes |
| 120 | CD24 | CD24 Entrez, Source | CD24 molecule | 13341 | -1.675 | 0.0001 | Yes |