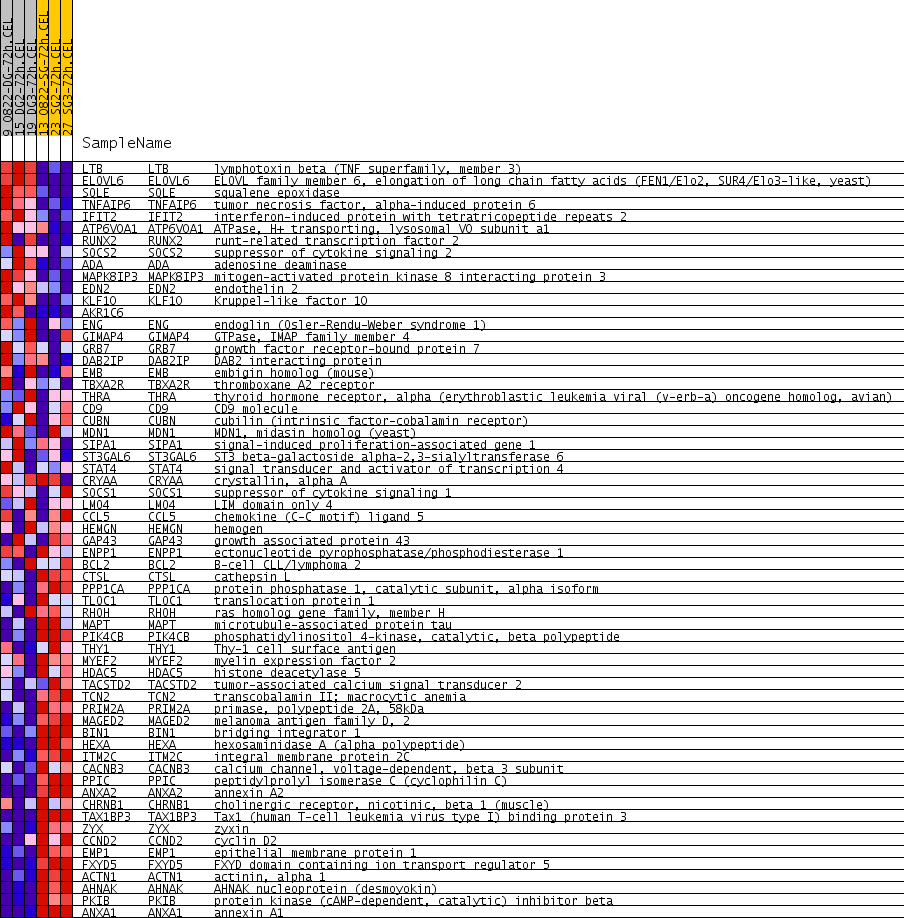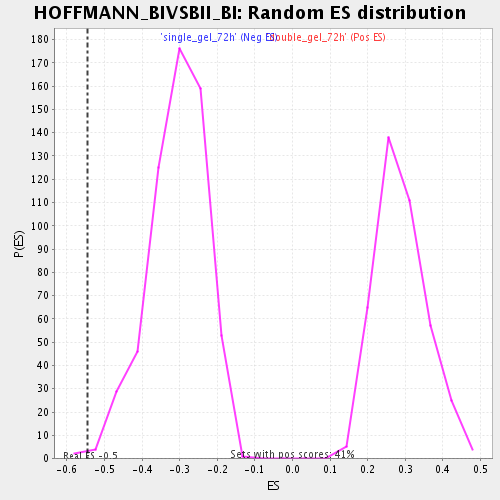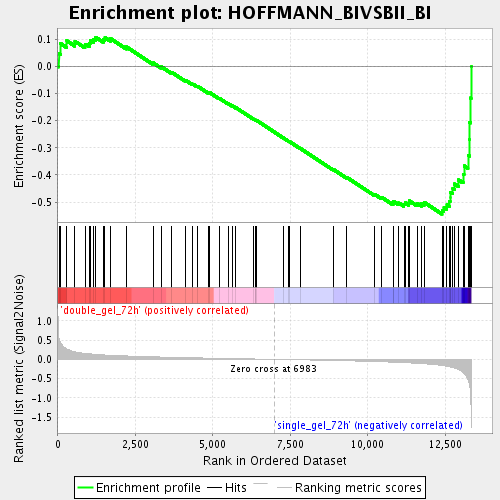
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | HOFFMANN_BIVSBII_BI |
| Enrichment Score (ES) | -0.5440254 |
| Normalized Enrichment Score (NES) | -1.7726493 |
| Nominal p-value | 0.0033613446 |
| FDR q-value | 0.04155084 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LTB | LTB Entrez, Source | lymphotoxin beta (TNF superfamily, member 3) | 35 | 0.539 | 0.0473 | No |
| 2 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 83 | 0.433 | 0.0839 | No |
| 3 | SQLE | SQLE Entrez, Source | squalene epoxidase | 287 | 0.271 | 0.0938 | No |
| 4 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 548 | 0.192 | 0.0920 | No |
| 5 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 875 | 0.155 | 0.0819 | No |
| 6 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 1017 | 0.143 | 0.0845 | No |
| 7 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 1041 | 0.141 | 0.0959 | No |
| 8 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 1150 | 0.133 | 0.1001 | No |
| 9 | ADA | ADA Entrez, Source | adenosine deaminase | 1219 | 0.129 | 0.1069 | No |
| 10 | MAPK8IP3 | MAPK8IP3 Entrez, Source | mitogen-activated protein kinase 8 interacting protein 3 | 1461 | 0.115 | 0.0994 | No |
| 11 | EDN2 | EDN2 Entrez, Source | endothelin 2 | 1518 | 0.112 | 0.1056 | No |
| 12 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 1693 | 0.105 | 0.1023 | No |
| 13 | AKR1C6 | 2209 | 0.088 | 0.0716 | No | ||
| 14 | ENG | ENG Entrez, Source | endoglin (Osler-Rendu-Weber syndrome 1) | 3072 | 0.065 | 0.0127 | No |
| 15 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 3337 | 0.059 | -0.0017 | No |
| 16 | GRB7 | GRB7 Entrez, Source | growth factor receptor-bound protein 7 | 3678 | 0.052 | -0.0224 | No |
| 17 | DAB2IP | DAB2IP Entrez, Source | DAB2 interacting protein | 4122 | 0.044 | -0.0517 | No |
| 18 | EMB | EMB Entrez, Source | embigin homolog (mouse) | 4351 | 0.040 | -0.0651 | No |
| 19 | TBXA2R | TBXA2R Entrez, Source | thromboxane A2 receptor | 4498 | 0.038 | -0.0726 | No |
| 20 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 4857 | 0.032 | -0.0966 | No |
| 21 | CD9 | CD9 Entrez, Source | CD9 molecule | 4896 | 0.031 | -0.0966 | No |
| 22 | CUBN | CUBN Entrez, Source | cubilin (intrinsic factor-cobalamin receptor) | 5212 | 0.026 | -0.1179 | No |
| 23 | MDN1 | MDN1 Entrez, Source | MDN1, midasin homolog (yeast) | 5490 | 0.022 | -0.1368 | No |
| 24 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 5630 | 0.020 | -0.1454 | No |
| 25 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 5745 | 0.018 | -0.1523 | No |
| 26 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 6316 | 0.009 | -0.1944 | No |
| 27 | CRYAA | CRYAA Entrez, Source | crystallin, alpha A | 6379 | 0.008 | -0.1983 | No |
| 28 | SOCS1 | SOCS1 Entrez, Source | suppressor of cytokine signaling 1 | 6382 | 0.008 | -0.1977 | No |
| 29 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 6402 | 0.008 | -0.1984 | No |
| 30 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 7275 | -0.004 | -0.2637 | No |
| 31 | HEMGN | HEMGN Entrez, Source | hemogen | 7451 | -0.007 | -0.2762 | No |
| 32 | GAP43 | GAP43 Entrez, Source | growth associated protein 43 | 7485 | -0.007 | -0.2780 | No |
| 33 | ENPP1 | ENPP1 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 1 | 7834 | -0.013 | -0.3031 | No |
| 34 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 8887 | -0.030 | -0.3795 | No |
| 35 | CTSL | CTSL Entrez, Source | cathepsin L | 9328 | -0.038 | -0.4091 | No |
| 36 | PPP1CA | PPP1CA Entrez, Source | protein phosphatase 1, catalytic subunit, alpha isoform | 10210 | -0.057 | -0.4702 | No |
| 37 | TLOC1 | TLOC1 Entrez, Source | translocation protein 1 | 10451 | -0.063 | -0.4824 | No |
| 38 | RHOH | RHOH Entrez, Source | ras homolog gene family, member H | 10818 | -0.073 | -0.5033 | No |
| 39 | MAPT | MAPT Entrez, Source | microtubule-associated protein tau | 10830 | -0.073 | -0.4973 | No |
| 40 | PIK4CB | PIK4CB Entrez, Source | phosphatidylinositol 4-kinase, catalytic, beta polypeptide | 11001 | -0.079 | -0.5028 | No |
| 41 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 11169 | -0.084 | -0.5076 | No |
| 42 | MYEF2 | MYEF2 Entrez, Source | myelin expression factor 2 | 11204 | -0.086 | -0.5022 | No |
| 43 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 11330 | -0.092 | -0.5031 | No |
| 44 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 11339 | -0.092 | -0.4951 | No |
| 45 | TCN2 | TCN2 Entrez, Source | transcobalamin II; macrocytic anemia | 11593 | -0.104 | -0.5046 | No |
| 46 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 11731 | -0.111 | -0.5046 | No |
| 47 | MAGED2 | MAGED2 Entrez, Source | melanoma antigen family D, 2 | 11818 | -0.115 | -0.5005 | No |
| 48 | BIN1 | BIN1 Entrez, Source | bridging integrator 1 | 12397 | -0.157 | -0.5295 | Yes |
| 49 | HEXA | HEXA Entrez, Source | hexosaminidase A (alpha polypeptide) | 12459 | -0.165 | -0.5188 | Yes |
| 50 | ITM2C | ITM2C Entrez, Source | integral membrane protein 2C | 12556 | -0.179 | -0.5094 | Yes |
| 51 | CACNB3 | CACNB3 Entrez, Source | calcium channel, voltage-dependent, beta 3 subunit | 12633 | -0.191 | -0.4974 | Yes |
| 52 | PPIC | PPIC Entrez, Source | peptidylprolyl isomerase C (cyclophilin C) | 12659 | -0.196 | -0.4812 | Yes |
| 53 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 12661 | -0.197 | -0.4630 | Yes |
| 54 | CHRNB1 | CHRNB1 Entrez, Source | cholinergic receptor, nicotinic, beta 1 (muscle) | 12746 | -0.213 | -0.4496 | Yes |
| 55 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 12802 | -0.225 | -0.4329 | Yes |
| 56 | ZYX | ZYX Entrez, Source | zyxin | 12928 | -0.262 | -0.4181 | Yes |
| 57 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 13080 | -0.340 | -0.3979 | Yes |
| 58 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 13109 | -0.363 | -0.3664 | Yes |
| 59 | FXYD5 | FXYD5 Entrez, Source | FXYD domain containing ion transport regulator 5 | 13236 | -0.522 | -0.3275 | Yes |
| 60 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13279 | -0.655 | -0.2700 | Yes |
| 61 | AHNAK | AHNAK Entrez, Source | AHNAK nucleoprotein (desmoyokin) | 13286 | -0.695 | -0.2061 | Yes |
| 62 | PKIB | PKIB Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor beta | 13324 | -0.997 | -0.1165 | Yes |
| 63 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | Yes |