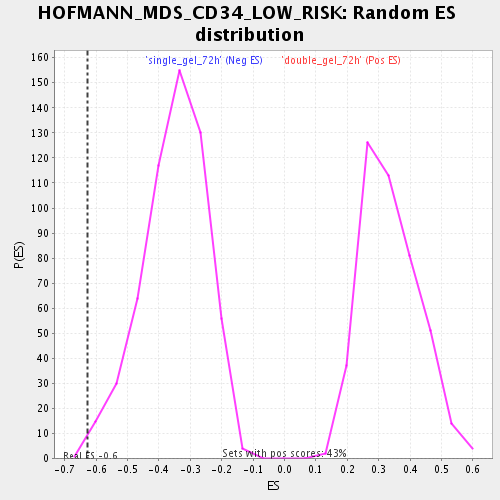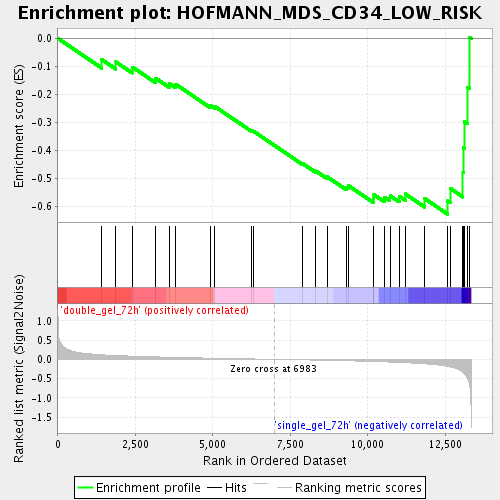
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | HOFMANN_MDS_CD34_LOW_RISK |
| Enrichment Score (ES) | -0.62746835 |
| Normalized Enrichment Score (NES) | -1.780919 |
| Nominal p-value | 0.0017482517 |
| FDR q-value | 0.040305533 |
| FWER p-Value | 0.991 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1413 | 0.118 | -0.0754 | No |
| 2 | DLK1 | DLK1 Entrez, Source | delta-like 1 homolog (Drosophila) | 1871 | 0.099 | -0.0840 | No |
| 3 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 2401 | 0.082 | -0.1023 | No |
| 4 | NCLN | NCLN Entrez, Source | nicalin homolog (zebrafish) | 3156 | 0.063 | -0.1424 | No |
| 5 | COIL | COIL Entrez, Source | coilin | 3588 | 0.054 | -0.1608 | No |
| 6 | ZMYM3 | ZMYM3 Entrez, Source | zinc finger, MYM-type 3 | 3804 | 0.050 | -0.1640 | No |
| 7 | HRAS | HRAS Entrez, Source | v-Ha-ras Harvey rat sarcoma viral oncogene homolog | 4923 | 0.030 | -0.2400 | No |
| 8 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 5059 | 0.028 | -0.2428 | No |
| 9 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 6256 | 0.010 | -0.3299 | No |
| 10 | PTPN7 | PTPN7 Entrez, Source | protein tyrosine phosphatase, non-receptor type 7 | 6306 | 0.009 | -0.3312 | No |
| 11 | DNM2 | DNM2 Entrez, Source | dynamin 2 | 7880 | -0.013 | -0.4458 | No |
| 12 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8319 | -0.020 | -0.4735 | No |
| 13 | STIP1 | STIP1 Entrez, Source | stress-induced-phosphoprotein 1 (Hsp70/Hsp90-organizing protein) | 8684 | -0.026 | -0.4940 | No |
| 14 | IER3 | IER3 Entrez, Source | immediate early response 3 | 9311 | -0.038 | -0.5311 | No |
| 15 | TSPAN1 | TSPAN1 Entrez, Source | tetraspanin 1 | 9367 | -0.039 | -0.5251 | No |
| 16 | MALL | MALL Entrez, Source | mal, T-cell differentiation protein-like | 10181 | -0.056 | -0.5716 | No |
| 17 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 10199 | -0.057 | -0.5581 | No |
| 18 | PSMD8 | PSMD8 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 8 | 10537 | -0.065 | -0.5665 | No |
| 19 | TOB2 | TOB2 Entrez, Source | transducer of ERBB2, 2 | 10718 | -0.070 | -0.5617 | No |
| 20 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 11020 | -0.079 | -0.5636 | No |
| 21 | CAPNS1 | CAPNS1 Entrez, Source | calpain, small subunit 1 | 11209 | -0.086 | -0.5553 | No |
| 22 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 11842 | -0.116 | -0.5726 | No |
| 23 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 12574 | -0.181 | -0.5804 | Yes |
| 24 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 12661 | -0.197 | -0.5357 | Yes |
| 25 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 13073 | -0.336 | -0.4791 | Yes |
| 26 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 13085 | -0.343 | -0.3907 | Yes |
| 27 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 13114 | -0.366 | -0.2974 | Yes |
| 28 | TUBB2B | TUBB2B Entrez, Source | tubulin, beta 2B | 13224 | -0.502 | -0.1748 | Yes |
| 29 | GPRC5A | GPRC5A Entrez, Source | G protein-coupled receptor, family C, group 5, member A | 13289 | -0.705 | 0.0040 | Yes |