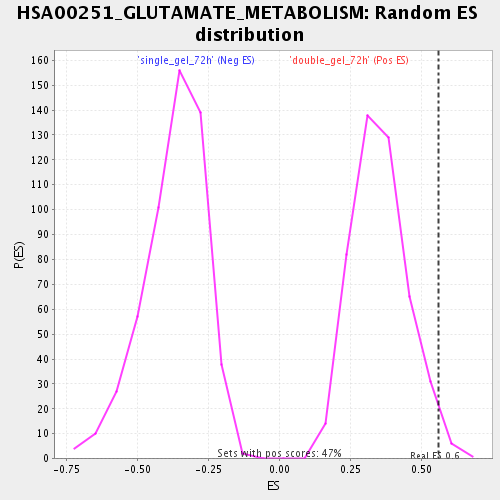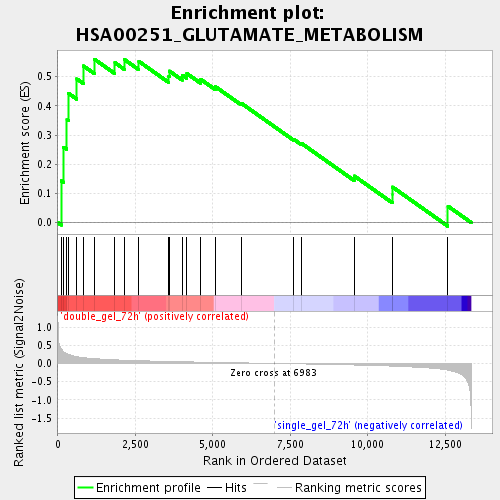
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | HSA00251_GLUTAMATE_METABOLISM |
| Enrichment Score (ES) | 0.55995756 |
| Normalized Enrichment Score (NES) | 1.5773866 |
| Nominal p-value | 0.015021459 |
| FDR q-value | 0.076018006 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 110 | 0.403 | 0.1434 | Yes |
| 2 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 185 | 0.321 | 0.2590 | Yes |
| 3 | CPS1 | CPS1 Entrez, Source | carbamoyl-phosphate synthetase 1, mitochondrial | 292 | 0.268 | 0.3519 | Yes |
| 4 | GLS2 | GLS2 Entrez, Source | glutaminase 2 (liver, mitochondrial) | 341 | 0.252 | 0.4431 | Yes |
| 5 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.4923 | Yes |
| 6 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 823 | 0.160 | 0.5369 | Yes |
| 7 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 1177 | 0.132 | 0.5600 | Yes |
| 8 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 1840 | 0.100 | 0.5478 | No |
| 9 | GPT | GPT Entrez, Source | glutamic-pyruvate transaminase (alanine aminotransferase) | 2146 | 0.090 | 0.5587 | No |
| 10 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 2613 | 0.076 | 0.5523 | No |
| 11 | GSR | GSR Entrez, Source | glutathione reductase | 3579 | 0.054 | 0.5003 | No |
| 12 | GMPS | GMPS Entrez, Source | guanine monphosphate synthetase | 3595 | 0.054 | 0.5195 | No |
| 13 | GSS | GSS Entrez, Source | glutathione synthetase | 4011 | 0.046 | 0.5057 | No |
| 14 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.5108 | No |
| 15 | NADSYN1 | NADSYN1 Entrez, Source | NAD synthetase 1 | 4609 | 0.036 | 0.4909 | No |
| 16 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 5087 | 0.028 | 0.4656 | No |
| 17 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 5916 | 0.015 | 0.4092 | No |
| 18 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 7612 | -0.009 | 0.2854 | No |
| 19 | GFPT2 | GFPT2 Entrez, Source | glutamine-fructose-6-phosphate transaminase 2 | 7852 | -0.013 | 0.2724 | No |
| 20 | NAGK | NAGK Entrez, Source | N-acetylglucosamine kinase | 9565 | -0.043 | 0.1600 | No |
| 21 | GFPT1 | GFPT1 Entrez, Source | glutamine-fructose-6-phosphate transaminase 1 | 10795 | -0.072 | 0.0950 | No |
| 22 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 10797 | -0.072 | 0.1222 | No |
| 23 | GLS | GLS Entrez, Source | glutaminase | 12588 | -0.183 | 0.0566 | No |