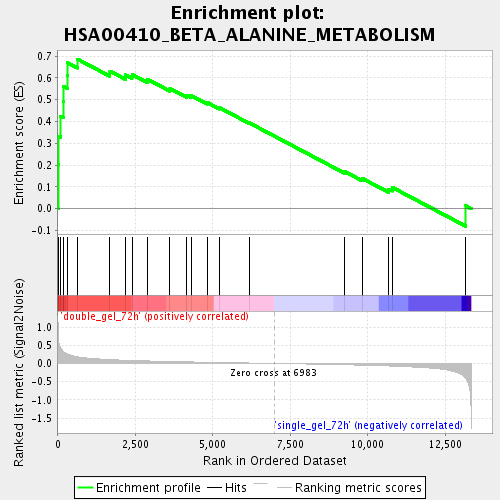
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | HSA00410_BETA_ALANINE_METABOLISM |
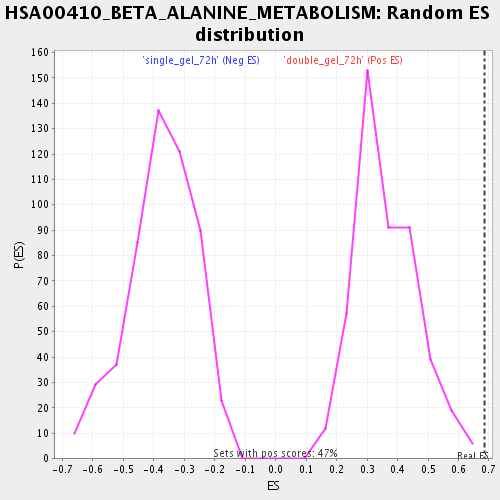
| Enrichment Score (ES) | 0.6855278 |
| Normalized Enrichment Score (NES) | 1.890684 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.007446468 |
| FWER p-Value | 0.505 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 6 | 0.908 | 0.2018 | Yes |
| 2 | ABP1 | ABP1 Entrez, Source | amiloride binding protein 1 (amine oxidase (copper-containing)) | 24 | 0.594 | 0.3328 | Yes |
| 3 | DPYS | DPYS Entrez, Source | dihydropyrimidinase | 90 | 0.425 | 0.4227 | Yes |
| 4 | UPB1 | UPB1 Entrez, Source | ureidopropionase, beta | 177 | 0.330 | 0.4899 | Yes |
| 5 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 185 | 0.321 | 0.5610 | Yes |
| 6 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 303 | 0.265 | 0.6113 | Yes |
| 7 | SRM | SRM Entrez, Source | spermidine synthase | 316 | 0.262 | 0.6687 | Yes |
| 8 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.6855 | Yes |
| 9 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.6307 | No |
| 10 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2174 | 0.089 | 0.6134 | No |
| 11 | HIBCH | HIBCH Entrez, Source | 3-hydroxyisobutyryl-Coenzyme A hydrolase | 2392 | 0.083 | 0.6155 | No |
| 12 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 2897 | 0.069 | 0.5930 | No |
| 13 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 3614 | 0.053 | 0.5512 | No |
| 14 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.5198 | No |
| 15 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.5188 | No |
| 16 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 4819 | 0.032 | 0.4870 | No |
| 17 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 5218 | 0.026 | 0.4629 | No |
| 18 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6175 | 0.012 | 0.3937 | No |
| 19 | CNDP1 | CNDP1 Entrez, Source | carnosine dipeptidase 1 (metallopeptidase M20 family) | 9246 | -0.037 | 0.1714 | No |
| 20 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 9824 | -0.048 | 0.1389 | No |
| 21 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 10674 | -0.069 | 0.0905 | No |
| 22 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 10797 | -0.072 | 0.0975 | No |
| 23 | SMS | SMS Entrez, Source | spermine synthase | 13163 | -0.420 | 0.0134 | No |