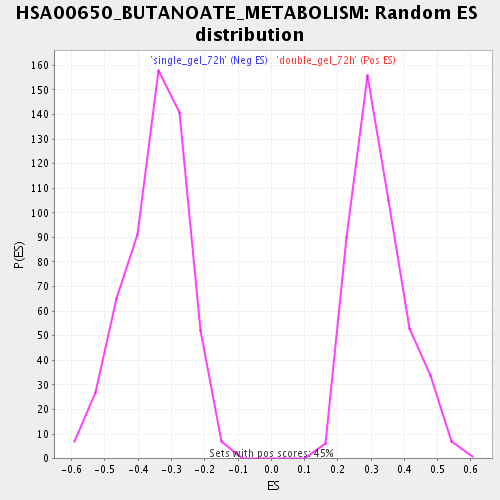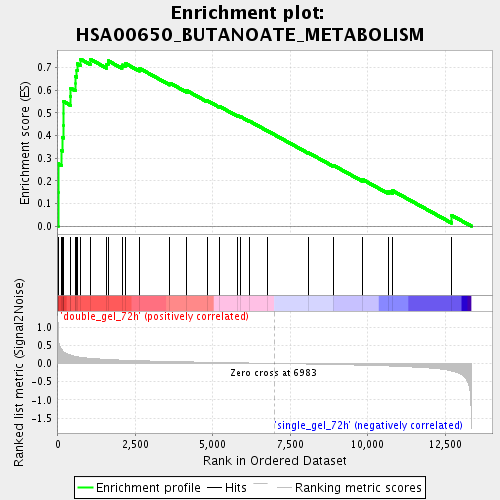
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | HSA00650_BUTANOATE_METABOLISM |
| Enrichment Score (ES) | 0.73723954 |
| Normalized Enrichment Score (NES) | 2.286015 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 6 | 0.908 | 0.1501 | Yes |
| 2 | AADAC | AADAC Entrez, Source | arylacetamide deacetylase (esterase) | 11 | 0.758 | 0.2754 | Yes |
| 3 | HMGCS2 | HMGCS2 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2 (mitochondrial) | 117 | 0.396 | 0.3333 | Yes |
| 4 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 144 | 0.362 | 0.3913 | Yes |
| 5 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 178 | 0.330 | 0.4434 | Yes |
| 6 | AACS | AACS Entrez, Source | acetoacetyl-CoA synthetase | 184 | 0.323 | 0.4967 | Yes |
| 7 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 185 | 0.321 | 0.5500 | Yes |
| 8 | DDHD1 | DDHD1 Entrez, Source | DDHD domain containing 1 | 401 | 0.231 | 0.5721 | Yes |
| 9 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 414 | 0.227 | 0.6088 | Yes |
| 10 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 560 | 0.191 | 0.6295 | Yes |
| 11 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 563 | 0.190 | 0.6610 | Yes |
| 12 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.6879 | Yes |
| 13 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.7166 | Yes |
| 14 | HMGCL | HMGCL Entrez, Source | 3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria) | 732 | 0.169 | 0.7372 | Yes |
| 15 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1044 | 0.141 | 0.7372 | No |
| 16 | BDH1 | BDH1 Entrez, Source | 3-hydroxybutyrate dehydrogenase, type 1 | 1574 | 0.110 | 0.7156 | No |
| 17 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1622 | 0.108 | 0.7300 | No |
| 18 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 2069 | 0.092 | 0.7117 | No |
| 19 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2174 | 0.089 | 0.7186 | No |
| 20 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 2625 | 0.076 | 0.6973 | No |
| 21 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 3614 | 0.053 | 0.6319 | No |
| 22 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.5980 | No |
| 23 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 4819 | 0.032 | 0.5541 | No |
| 24 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 5218 | 0.026 | 0.5285 | No |
| 25 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 5797 | 0.017 | 0.4880 | No |
| 26 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.4840 | No |
| 27 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 6175 | 0.012 | 0.4643 | No |
| 28 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6768 | 0.003 | 0.4203 | No |
| 29 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8089 | -0.016 | 0.3238 | No |
| 30 | PPME1 | PPME1 Entrez, Source | protein phosphatase methylesterase 1 | 8878 | -0.029 | 0.2695 | No |
| 31 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 9824 | -0.048 | 0.2065 | No |
| 32 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 10674 | -0.069 | 0.1541 | No |
| 33 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 10797 | -0.072 | 0.1569 | No |
| 34 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 12708 | -0.206 | 0.0476 | No |