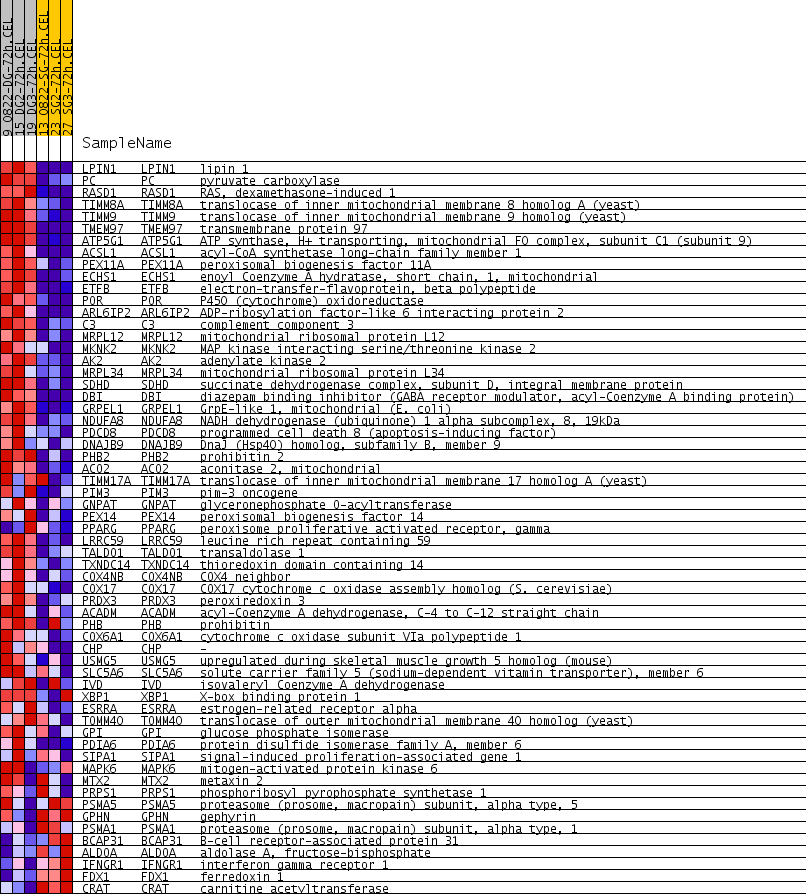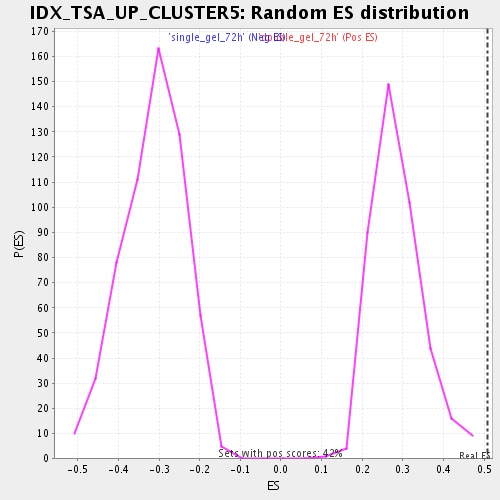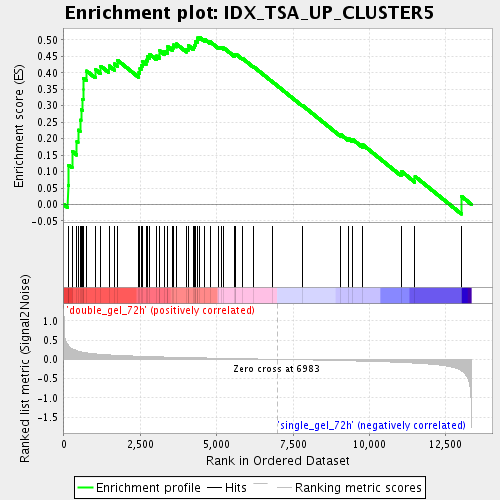
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | IDX_TSA_UP_CLUSTER5 |
| Enrichment Score (ES) | 0.50810105 |
| Normalized Enrichment Score (NES) | 1.7770141 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.019429456 |
| FWER p-Value | 0.946 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 133 | 0.378 | 0.0585 | Yes |
| 2 | PC | PC Entrez, Source | pyruvate carboxylase | 155 | 0.347 | 0.1199 | Yes |
| 3 | RASD1 | RASD1 Entrez, Source | RAS, dexamethasone-induced 1 | 269 | 0.278 | 0.1619 | Yes |
| 4 | TIMM8A | TIMM8A Entrez, Source | translocase of inner mitochondrial membrane 8 homolog A (yeast) | 425 | 0.223 | 0.1908 | Yes |
| 5 | TIMM9 | TIMM9 Entrez, Source | translocase of inner mitochondrial membrane 9 homolog (yeast) | 463 | 0.212 | 0.2264 | Yes |
| 6 | TMEM97 | TMEM97 Entrez, Source | transmembrane protein 97 | 532 | 0.196 | 0.2568 | Yes |
| 7 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 561 | 0.191 | 0.2893 | Yes |
| 8 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 609 | 0.185 | 0.3193 | Yes |
| 9 | PEX11A | PEX11A Entrez, Source | peroxisomal biogenesis factor 11A | 631 | 0.182 | 0.3507 | Yes |
| 10 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.3837 | Yes |
| 11 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 746 | 0.168 | 0.4057 | Yes |
| 12 | POR | POR Entrez, Source | P450 (cytochrome) oxidoreductase | 1030 | 0.142 | 0.4102 | Yes |
| 13 | ARL6IP2 | ARL6IP2 Entrez, Source | ADP-ribosylation factor-like 6 interacting protein 2 | 1204 | 0.130 | 0.4207 | Yes |
| 14 | C3 | C3 Entrez, Source | complement component 3 | 1475 | 0.114 | 0.4211 | Yes |
| 15 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 1652 | 0.107 | 0.4272 | Yes |
| 16 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 1755 | 0.103 | 0.4382 | Yes |
| 17 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 2453 | 0.081 | 0.4004 | Yes |
| 18 | MRPL34 | MRPL34 Entrez, Source | mitochondrial ribosomal protein L34 | 2487 | 0.080 | 0.4125 | Yes |
| 19 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 2529 | 0.079 | 0.4237 | Yes |
| 20 | DBI | DBI Entrez, Source | diazepam binding inhibitor (GABA receptor modulator, acyl-Coenzyme A binding protein) | 2576 | 0.077 | 0.4342 | Yes |
| 21 | GRPEL1 | GRPEL1 Entrez, Source | GrpE-like 1, mitochondrial (E. coli) | 2691 | 0.074 | 0.4390 | Yes |
| 22 | NDUFA8 | NDUFA8 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 8, 19kDa | 2735 | 0.073 | 0.4490 | Yes |
| 23 | PDCD8 | PDCD8 Entrez, Source | programmed cell death 8 (apoptosis-inducing factor) | 2803 | 0.071 | 0.4569 | Yes |
| 24 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 3036 | 0.066 | 0.4514 | Yes |
| 25 | PHB2 | PHB2 Entrez, Source | prohibitin 2 | 3135 | 0.064 | 0.4555 | Yes |
| 26 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 3136 | 0.064 | 0.4671 | Yes |
| 27 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 3292 | 0.060 | 0.4663 | Yes |
| 28 | PIM3 | PIM3 Entrez, Source | pim-3 oncogene | 3391 | 0.058 | 0.4695 | Yes |
| 29 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3398 | 0.058 | 0.4796 | Yes |
| 30 | PEX14 | PEX14 Entrez, Source | peroxisomal biogenesis factor 14 | 3565 | 0.054 | 0.4770 | Yes |
| 31 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 3592 | 0.054 | 0.4848 | Yes |
| 32 | LRRC59 | LRRC59 Entrez, Source | leucine rich repeat containing 59 | 3677 | 0.052 | 0.4880 | Yes |
| 33 | TALDO1 | TALDO1 Entrez, Source | transaldolase 1 | 4010 | 0.046 | 0.4714 | Yes |
| 34 | TXNDC14 | TXNDC14 Entrez, Source | thioredoxin domain containing 14 | 4067 | 0.045 | 0.4753 | Yes |
| 35 | COX4NB | COX4NB Entrez, Source | COX4 neighbor | 4072 | 0.045 | 0.4832 | Yes |
| 36 | COX17 | COX17 Entrez, Source | COX17 cytochrome c oxidase assembly homolog (S. cerevisiae) | 4243 | 0.042 | 0.4780 | Yes |
| 37 | PRDX3 | PRDX3 Entrez, Source | peroxiredoxin 3 | 4271 | 0.042 | 0.4836 | Yes |
| 38 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.4890 | Yes |
| 39 | PHB | PHB Entrez, Source | prohibitin | 4309 | 0.041 | 0.4958 | Yes |
| 40 | COX6A1 | COX6A1 Entrez, Source | cytochrome c oxidase subunit VIa polypeptide 1 | 4367 | 0.040 | 0.4988 | Yes |
| 41 | CHP | CHP Entrez, Source | - | 4368 | 0.040 | 0.5060 | Yes |
| 42 | USMG5 | USMG5 Entrez, Source | upregulated during skeletal muscle growth 5 homolog (mouse) | 4435 | 0.039 | 0.5081 | Yes |
| 43 | SLC5A6 | SLC5A6 Entrez, Source | solute carrier family 5 (sodium-dependent vitamin transporter), member 6 | 4600 | 0.036 | 0.5023 | No |
| 44 | IVD | IVD Entrez, Source | isovaleryl Coenzyme A dehydrogenase | 4783 | 0.033 | 0.4946 | No |
| 45 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 5073 | 0.028 | 0.4779 | No |
| 46 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 5141 | 0.027 | 0.4778 | No |
| 47 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 5228 | 0.026 | 0.4760 | No |
| 48 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 5582 | 0.021 | 0.4531 | No |
| 49 | PDIA6 | PDIA6 Entrez, Source | protein disulfide isomerase family A, member 6 | 5601 | 0.020 | 0.4554 | No |
| 50 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 5630 | 0.020 | 0.4569 | No |
| 51 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 5856 | 0.016 | 0.4430 | No |
| 52 | MTX2 | MTX2 Entrez, Source | metaxin 2 | 6206 | 0.011 | 0.4187 | No |
| 53 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 6821 | 0.002 | 0.3729 | No |
| 54 | PSMA5 | PSMA5 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 5 | 7801 | -0.012 | 0.3014 | No |
| 55 | GPHN | GPHN Entrez, Source | gephyrin | 9059 | -0.033 | 0.2128 | No |
| 56 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 9310 | -0.038 | 0.2009 | No |
| 57 | BCAP31 | BCAP31 Entrez, Source | B-cell receptor-associated protein 31 | 9449 | -0.041 | 0.1979 | No |
| 58 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 9779 | -0.047 | 0.1817 | No |
| 59 | IFNGR1 | IFNGR1 Entrez, Source | interferon gamma receptor 1 | 11041 | -0.080 | 0.1012 | No |
| 60 | FDX1 | FDX1 Entrez, Source | ferredoxin 1 | 11489 | -0.099 | 0.0855 | No |
| 61 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 13012 | -0.297 | 0.0248 | No |