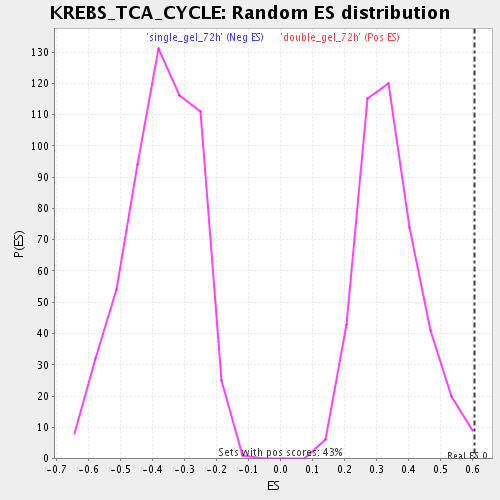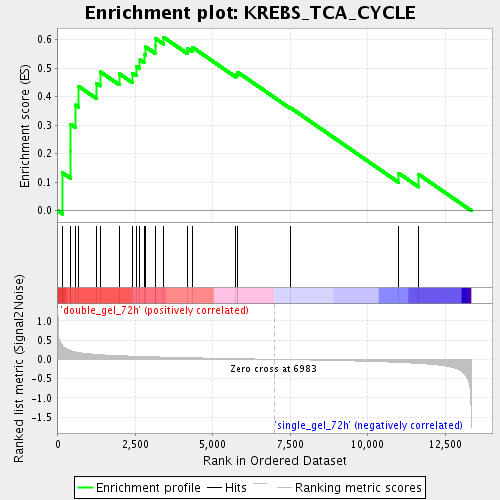
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | KREBS_TCA_CYCLE |
| Enrichment Score (ES) | 0.60801214 |
| Normalized Enrichment Score (NES) | 1.7604616 |
| Nominal p-value | 0.0070093456 |
| FDR q-value | 0.021777423 |
| FWER p-Value | 0.972 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PC | PC Entrez, Source | pyruvate carboxylase | 155 | 0.347 | 0.1328 | Yes |
| 2 | PDK4 | PDK4 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 4 | 412 | 0.227 | 0.2082 | Yes |
| 3 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 414 | 0.227 | 0.3026 | Yes |
| 4 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 563 | 0.190 | 0.3708 | Yes |
| 5 | PDK1 | PDK1 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 1 | 673 | 0.176 | 0.4359 | Yes |
| 6 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 1246 | 0.127 | 0.4458 | Yes |
| 7 | PDP2 | PDP2 Entrez, Source | - | 1362 | 0.121 | 0.4875 | Yes |
| 8 | IDH3A | IDH3A Entrez, Source | isocitrate dehydrogenase 3 (NAD+) alpha | 1985 | 0.094 | 0.4800 | Yes |
| 9 | MDH2 | MDH2 Entrez, Source | malate dehydrogenase 2, NAD (mitochondrial) | 2409 | 0.082 | 0.4825 | Yes |
| 10 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 2529 | 0.079 | 0.5063 | Yes |
| 11 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 2648 | 0.075 | 0.5286 | Yes |
| 12 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2780 | 0.072 | 0.5487 | Yes |
| 13 | PDK2 | PDK2 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 2 | 2834 | 0.071 | 0.5741 | Yes |
| 14 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 3136 | 0.064 | 0.5779 | Yes |
| 15 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 3142 | 0.063 | 0.6040 | Yes |
| 16 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 3410 | 0.058 | 0.6080 | Yes |
| 17 | CS | CS Entrez, Source | citrate synthase | 4175 | 0.043 | 0.5687 | No |
| 18 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 4335 | 0.040 | 0.5736 | No |
| 19 | SUCLG1 | SUCLG1 Entrez, Source | succinate-CoA ligase, GDP-forming, alpha subunit | 5738 | 0.018 | 0.4759 | No |
| 20 | SDHC | SDHC Entrez, Source | succinate dehydrogenase complex, subunit C, integral membrane protein, 15kDa | 5789 | 0.018 | 0.4795 | No |
| 21 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 5797 | 0.017 | 0.4862 | No |
| 22 | SUCLG2 | SUCLG2 Entrez, Source | succinate-CoA ligase, GDP-forming, beta subunit | 7511 | -0.008 | 0.3608 | No |
| 23 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 10995 | -0.078 | 0.1319 | No |
| 24 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 11644 | -0.106 | 0.1275 | No |