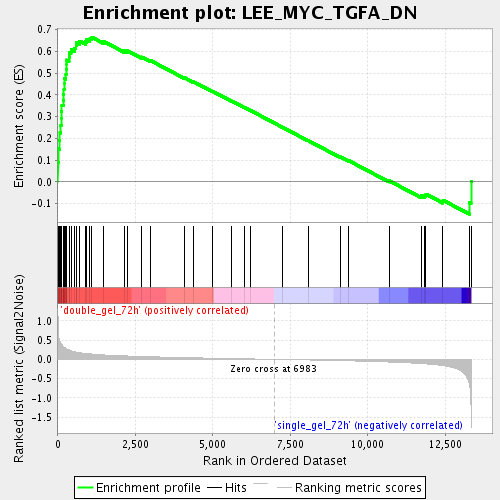
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | LEE_MYC_TGFA_DN |
| Enrichment Score (ES) | 0.6649148 |
| Normalized Enrichment Score (NES) | 2.211613 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.8297805E-4 |
| FWER p-Value | 0.0050 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 2 | 1.112 | 0.0898 | Yes |
| 2 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 9 | 0.779 | 0.1524 | Yes |
| 3 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 44 | 0.506 | 0.1907 | Yes |
| 4 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 66 | 0.461 | 0.2264 | Yes |
| 5 | KHK | KHK Entrez, Source | ketohexokinase (fructokinase) | 93 | 0.422 | 0.2586 | Yes |
| 6 | BHMT | BHMT Entrez, Source | betaine-homocysteine methyltransferase | 106 | 0.415 | 0.2912 | Yes |
| 7 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 110 | 0.403 | 0.3236 | Yes |
| 8 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 129 | 0.383 | 0.3532 | Yes |
| 9 | C4BPA | C4BPA Entrez, Source | complement component 4 binding protein, alpha | 176 | 0.331 | 0.3765 | Yes |
| 10 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 185 | 0.321 | 0.4019 | Yes |
| 11 | AQP8 | AQP8 Entrez, Source | aquaporin 8 | 195 | 0.312 | 0.4265 | Yes |
| 12 | PXMP2 | PXMP2 Entrez, Source | peroxisomal membrane protein 2, 22kDa | 202 | 0.309 | 0.4510 | Yes |
| 13 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 207 | 0.307 | 0.4755 | Yes |
| 14 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 258 | 0.285 | 0.4948 | Yes |
| 15 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 266 | 0.281 | 0.5170 | Yes |
| 16 | KLF15 | KLF15 Entrez, Source | Kruppel-like factor 15 | 278 | 0.275 | 0.5384 | Yes |
| 17 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 282 | 0.274 | 0.5604 | Yes |
| 18 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 358 | 0.245 | 0.5746 | Yes |
| 19 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 370 | 0.242 | 0.5933 | Yes |
| 20 | HAL | HAL Entrez, Source | histidine ammonia-lyase | 423 | 0.224 | 0.6075 | Yes |
| 21 | IGFALS | IGFALS Entrez, Source | insulin-like growth factor binding protein, acid labile subunit | 533 | 0.196 | 0.6152 | Yes |
| 22 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 583 | 0.188 | 0.6267 | Yes |
| 23 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 597 | 0.186 | 0.6408 | Yes |
| 24 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 695 | 0.174 | 0.6475 | Yes |
| 25 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 893 | 0.154 | 0.6451 | Yes |
| 26 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 933 | 0.150 | 0.6543 | Yes |
| 27 | CA5A | CA5A Entrez, Source | carbonic anhydrase VA, mitochondrial | 1034 | 0.142 | 0.6583 | Yes |
| 28 | HADH2 | HADH2 Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase, type II | 1095 | 0.137 | 0.6649 | Yes |
| 29 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 1471 | 0.115 | 0.6460 | No |
| 30 | SLC25A22 | SLC25A22 Entrez, Source | solute carrier family 25 (mitochondrial carrier: glutamate), member 22 | 2140 | 0.090 | 0.6030 | No |
| 31 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 2239 | 0.087 | 0.6026 | No |
| 32 | HLA-DMA | HLA-DMA Entrez, Source | major histocompatibility complex, class II, DM alpha | 2697 | 0.074 | 0.5742 | No |
| 33 | FABP2 | FABP2 Entrez, Source | fatty acid binding protein 2, intestinal | 2972 | 0.067 | 0.5590 | No |
| 34 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 4079 | 0.045 | 0.4795 | No |
| 35 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 4383 | 0.040 | 0.4599 | No |
| 36 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4996 | 0.029 | 0.4162 | No |
| 37 | ALAS2 | ALAS2 Entrez, Source | aminolevulinate, delta-, synthase 2 (sideroblastic/hypochromic anemia) | 5606 | 0.020 | 0.3720 | No |
| 38 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 6026 | 0.014 | 0.3416 | No |
| 39 | CRISP2 | CRISP2 Entrez, Source | cysteine-rich secretory protein 2 | 6210 | 0.011 | 0.3287 | No |
| 40 | CMAH | CMAH Entrez, Source | cytidine monophosphate-N-acetylneuraminic acid hydroxylase (CMP-N-acetylneuraminate monooxygenase) | 7234 | -0.004 | 0.2521 | No |
| 41 | ABCG2 | ABCG2 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 2 | 8071 | -0.016 | 0.1905 | No |
| 42 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9114 | -0.034 | 0.1149 | No |
| 43 | GPR37 | GPR37 Entrez, Source | G protein-coupled receptor 37 (endothelin receptor type B-like) | 9392 | -0.040 | 0.0972 | No |
| 44 | PSEN2 | PSEN2 Entrez, Source | presenilin 2 (Alzheimer disease 4) | 10691 | -0.070 | 0.0052 | No |
| 45 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 11727 | -0.110 | -0.0637 | No |
| 46 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 11842 | -0.116 | -0.0629 | No |
| 47 | TRPV2 | TRPV2 Entrez, Source | transient receptor potential cation channel, subfamily V, member 2 | 11876 | -0.118 | -0.0559 | No |
| 48 | RGN | RGN Entrez, Source | regucalcin (senescence marker protein-30) | 12427 | -0.161 | -0.0842 | No |
| 49 | LIPC | LIPC Entrez, Source | lipase, hepatic | 13275 | -0.650 | -0.0953 | No |
| 50 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13334 | -1.240 | 0.0006 | No |

