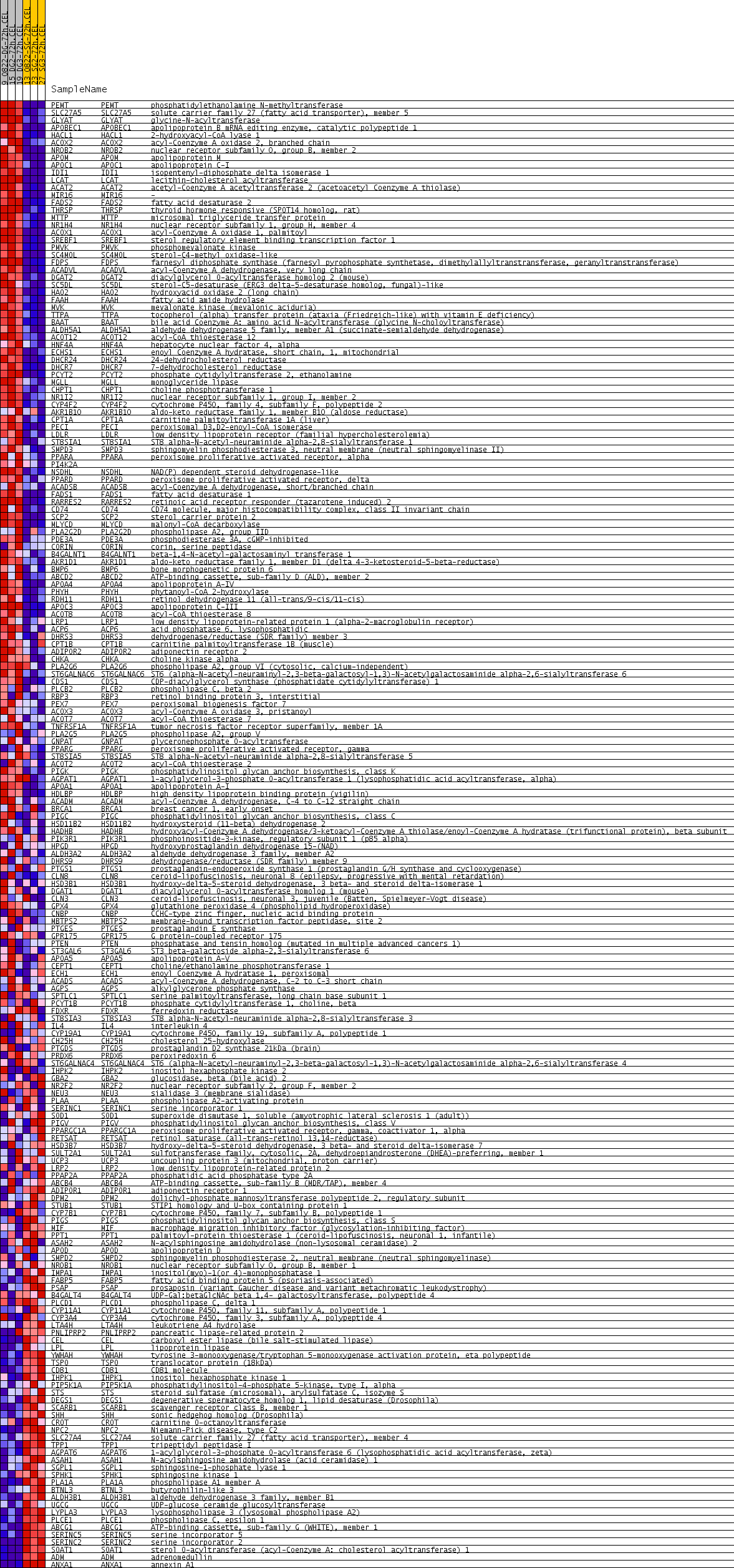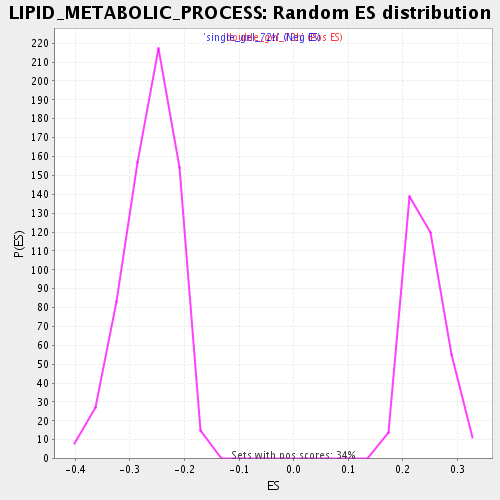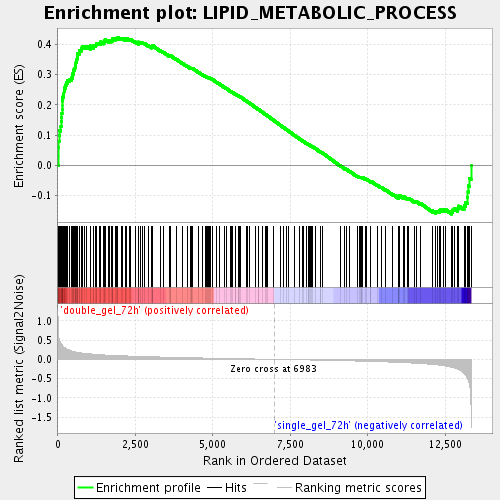
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | LIPID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.42425528 |
| Normalized Enrichment Score (NES) | 1.7673296 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.020873804 |
| FWER p-Value | 0.962 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PEMT | PEMT Entrez, Source | phosphatidylethanolamine N-methyltransferase | 8 | 0.798 | 0.0298 | Yes |
| 2 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 9 | 0.779 | 0.0595 | Yes |
| 3 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 33 | 0.554 | 0.0789 | Yes |
| 4 | APOBEC1 | APOBEC1 Entrez, Source | apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 | 40 | 0.526 | 0.0985 | Yes |
| 5 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 54 | 0.488 | 0.1161 | Yes |
| 6 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 95 | 0.419 | 0.1291 | Yes |
| 7 | NR0B2 | NR0B2 Entrez, Source | nuclear receptor subfamily 0, group B, member 2 | 104 | 0.416 | 0.1443 | Yes |
| 8 | APOM | APOM Entrez, Source | apolipoprotein M | 113 | 0.401 | 0.1590 | Yes |
| 9 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 127 | 0.384 | 0.1726 | Yes |
| 10 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 136 | 0.372 | 0.1862 | Yes |
| 11 | LCAT | LCAT Entrez, Source | lecithin-cholesterol acyltransferase | 142 | 0.365 | 0.1998 | Yes |
| 12 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 144 | 0.362 | 0.2135 | Yes |
| 13 | MIR16 | MIR16 Entrez, Source | - | 152 | 0.351 | 0.2263 | Yes |
| 14 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.2365 | Yes |
| 15 | THRSP | THRSP Entrez, Source | thyroid hormone responsive (SPOT14 homolog, rat) | 196 | 0.311 | 0.2474 | Yes |
| 16 | MTTP | MTTP Entrez, Source | microsomal triglyceride transfer protein | 218 | 0.301 | 0.2573 | Yes |
| 17 | NR1H4 | NR1H4 Entrez, Source | nuclear receptor subfamily 1, group H, member 4 | 246 | 0.288 | 0.2662 | Yes |
| 18 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 266 | 0.281 | 0.2754 | Yes |
| 19 | SREBF1 | SREBF1 Entrez, Source | sterol regulatory element binding transcription factor 1 | 310 | 0.264 | 0.2822 | Yes |
| 20 | PMVK | PMVK Entrez, Source | phosphomevalonate kinase | 385 | 0.237 | 0.2856 | Yes |
| 21 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 444 | 0.218 | 0.2895 | Yes |
| 22 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 462 | 0.212 | 0.2963 | Yes |
| 23 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 482 | 0.207 | 0.3028 | Yes |
| 24 | DGAT2 | DGAT2 Entrez, Source | diacylglycerol O-acyltransferase homolog 2 (mouse) | 503 | 0.201 | 0.3089 | Yes |
| 25 | SC5DL | SC5DL Entrez, Source | sterol-C5-desaturase (ERG3 delta-5-desaturase homolog, fungal)-like | 512 | 0.201 | 0.3160 | Yes |
| 26 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 536 | 0.195 | 0.3216 | Yes |
| 27 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 553 | 0.192 | 0.3277 | Yes |
| 28 | MVK | MVK Entrez, Source | mevalonate kinase (mevalonic aciduria) | 578 | 0.188 | 0.3331 | Yes |
| 29 | TTPA | TTPA Entrez, Source | tocopherol (alpha) transfer protein (ataxia (Friedreich-like) with vitamin E deficiency) | 581 | 0.188 | 0.3401 | Yes |
| 30 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 597 | 0.186 | 0.3461 | Yes |
| 31 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.3520 | Yes |
| 32 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 619 | 0.184 | 0.3586 | Yes |
| 33 | HNF4A | HNF4A Entrez, Source | hepatocyte nuclear factor 4, alpha | 626 | 0.183 | 0.3651 | Yes |
| 34 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.3716 | Yes |
| 35 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 685 | 0.175 | 0.3744 | Yes |
| 36 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 693 | 0.174 | 0.3805 | Yes |
| 37 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 748 | 0.168 | 0.3828 | Yes |
| 38 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 754 | 0.167 | 0.3887 | Yes |
| 39 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 778 | 0.165 | 0.3933 | Yes |
| 40 | NR1I2 | NR1I2 Entrez, Source | nuclear receptor subfamily 1, group I, member 2 | 855 | 0.157 | 0.3935 | Yes |
| 41 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 933 | 0.150 | 0.3933 | Yes |
| 42 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1044 | 0.141 | 0.3903 | Yes |
| 43 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 1046 | 0.140 | 0.3956 | Yes |
| 44 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1151 | 0.133 | 0.3928 | Yes |
| 45 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 1160 | 0.133 | 0.3972 | Yes |
| 46 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 1223 | 0.128 | 0.3974 | Yes |
| 47 | SMPD3 | SMPD3 Entrez, Source | sphingomyelin phosphodiesterase 3, neutral membrane (neutral sphingomyelinase II) | 1232 | 0.128 | 0.4017 | Yes |
| 48 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 1252 | 0.127 | 0.4051 | Yes |
| 49 | PI4K2A | 1329 | 0.123 | 0.4040 | Yes | ||
| 50 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1370 | 0.121 | 0.4055 | Yes |
| 51 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 1387 | 0.119 | 0.4088 | Yes |
| 52 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1480 | 0.114 | 0.4062 | Yes |
| 53 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.4095 | Yes |
| 54 | RARRES2 | RARRES2 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 2 | 1501 | 0.113 | 0.4134 | Yes |
| 55 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.4160 | Yes |
| 56 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 1632 | 0.107 | 0.4119 | Yes |
| 57 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.4126 | Yes |
| 58 | PLA2G2D | PLA2G2D Entrez, Source | phospholipase A2, group IID | 1734 | 0.104 | 0.4123 | Yes |
| 59 | PDE3A | PDE3A Entrez, Source | phosphodiesterase 3A, cGMP-inhibited | 1751 | 0.103 | 0.4150 | Yes |
| 60 | CORIN | CORIN Entrez, Source | corin, serine peptidase | 1759 | 0.103 | 0.4184 | Yes |
| 61 | B4GALNT1 | B4GALNT1 Entrez, Source | beta-1,4-N-acetyl-galactosaminyl transferase 1 | 1769 | 0.102 | 0.4216 | Yes |
| 62 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.4186 | Yes |
| 63 | BMP6 | BMP6 Entrez, Source | bone morphogenetic protein 6 | 1877 | 0.098 | 0.4211 | Yes |
| 64 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 1926 | 0.096 | 0.4211 | Yes |
| 65 | APOA4 | APOA4 Entrez, Source | apolipoprotein A-IV | 1934 | 0.096 | 0.4243 | Yes |
| 66 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 2036 | 0.093 | 0.4201 | No |
| 67 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 2069 | 0.092 | 0.4212 | No |
| 68 | APOC3 | APOC3 Entrez, Source | apolipoprotein C-III | 2176 | 0.089 | 0.4165 | No |
| 69 | ACOT8 | ACOT8 Entrez, Source | acyl-CoA thioesterase 8 | 2184 | 0.089 | 0.4193 | No |
| 70 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 2212 | 0.088 | 0.4206 | No |
| 71 | ACP6 | ACP6 Entrez, Source | acid phosphatase 6, lysophosphatidic | 2295 | 0.085 | 0.4176 | No |
| 72 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 2353 | 0.084 | 0.4165 | No |
| 73 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 2496 | 0.080 | 0.4087 | No |
| 74 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | 0.4040 | No |
| 75 | CHKA | CHKA Entrez, Source | choline kinase alpha | 2612 | 0.076 | 0.4059 | No |
| 76 | PLA2G6 | PLA2G6 Entrez, Source | phospholipase A2, group VI (cytosolic, calcium-independent) | 2615 | 0.076 | 0.4086 | No |
| 77 | ST6GALNAC6 | ST6GALNAC6 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 6 | 2672 | 0.074 | 0.4072 | No |
| 78 | CDS1 | CDS1 Entrez, Source | CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 1 | 2733 | 0.073 | 0.4054 | No |
| 79 | PLCB2 | PLCB2 Entrez, Source | phospholipase C, beta 2 | 2782 | 0.072 | 0.4045 | No |
| 80 | RBP3 | RBP3 Entrez, Source | retinol binding protein 3, interstitial | 2917 | 0.068 | 0.3969 | No |
| 81 | PEX7 | PEX7 Entrez, Source | peroxisomal biogenesis factor 7 | 3029 | 0.066 | 0.3910 | No |
| 82 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 3046 | 0.066 | 0.3923 | No |
| 83 | ACOT7 | ACOT7 Entrez, Source | acyl-CoA thioesterase 7 | 3055 | 0.065 | 0.3941 | No |
| 84 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 3063 | 0.065 | 0.3961 | No |
| 85 | PLA2G5 | PLA2G5 Entrez, Source | phospholipase A2, group V | 3301 | 0.060 | 0.3804 | No |
| 86 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3398 | 0.058 | 0.3753 | No |
| 87 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 3592 | 0.054 | 0.3626 | No |
| 88 | ST8SIA5 | ST8SIA5 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 5 | 3625 | 0.053 | 0.3622 | No |
| 89 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 3642 | 0.053 | 0.3630 | No |
| 90 | PIGK | PIGK Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class K | 3814 | 0.050 | 0.3519 | No |
| 91 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 4028 | 0.046 | 0.3375 | No |
| 92 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 4169 | 0.043 | 0.3285 | No |
| 93 | HDLBP | HDLBP Entrez, Source | high density lipoprotein binding protein (vigilin) | 4270 | 0.042 | 0.3225 | No |
| 94 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.3219 | No |
| 95 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | 0.3215 | No |
| 96 | PIGC | PIGC Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class C | 4338 | 0.040 | 0.3222 | No |
| 97 | HSD11B2 | HSD11B2 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 2 | 4542 | 0.037 | 0.3082 | No |
| 98 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.2994 | No |
| 99 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 4760 | 0.033 | 0.2943 | No |
| 100 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.2924 | No |
| 101 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 4819 | 0.032 | 0.2924 | No |
| 102 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 4865 | 0.031 | 0.2902 | No |
| 103 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 4887 | 0.031 | 0.2898 | No |
| 104 | CLN8 | CLN8 Entrez, Source | ceroid-lipofuscinosis, neuronal 8 (epilepsy, progressive with mental retardation) | 4936 | 0.030 | 0.2873 | No |
| 105 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 4997 | 0.029 | 0.2839 | No |
| 106 | DGAT1 | DGAT1 Entrez, Source | diacylglycerol O-acyltransferase homolog 1 (mouse) | 5118 | 0.027 | 0.2758 | No |
| 107 | CLN3 | CLN3 Entrez, Source | ceroid-lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | 5208 | 0.026 | 0.2700 | No |
| 108 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 5373 | 0.024 | 0.2584 | No |
| 109 | CNBP | CNBP Entrez, Source | CCHC-type zinc finger, nucleic acid binding protein | 5438 | 0.023 | 0.2544 | No |
| 110 | MBTPS2 | MBTPS2 Entrez, Source | membrane-bound transcription factor peptidase, site 2 | 5566 | 0.021 | 0.2456 | No |
| 111 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 5602 | 0.020 | 0.2437 | No |
| 112 | GPR175 | GPR175 Entrez, Source | G protein-coupled receptor 175 | 5625 | 0.020 | 0.2428 | No |
| 113 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 5720 | 0.018 | 0.2363 | No |
| 114 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 5745 | 0.018 | 0.2352 | No |
| 115 | APOA5 | APOA5 Entrez, Source | apolipoprotein A-V | 5815 | 0.017 | 0.2306 | No |
| 116 | CEPT1 | CEPT1 Entrez, Source | choline/ethanolamine phosphotransferase 1 | 5830 | 0.017 | 0.2302 | No |
| 117 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.2287 | No |
| 118 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.2273 | No |
| 119 | AGPS | AGPS Entrez, Source | alkylglycerone phosphate synthase | 6079 | 0.013 | 0.2132 | No |
| 120 | SPTLC1 | SPTLC1 Entrez, Source | serine palmitoyltransferase, long chain base subunit 1 | 6112 | 0.013 | 0.2112 | No |
| 121 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 6186 | 0.012 | 0.2061 | No |
| 122 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 6391 | 0.008 | 0.1909 | No |
| 123 | ST8SIA3 | ST8SIA3 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 | 6457 | 0.007 | 0.1862 | No |
| 124 | IL4 | IL4 Entrez, Source | interleukin 4 | 6466 | 0.007 | 0.1859 | No |
| 125 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 6596 | 0.005 | 0.1763 | No |
| 126 | CH25H | CH25H Entrez, Source | cholesterol 25-hydroxylase | 6694 | 0.004 | 0.1690 | No |
| 127 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | 0.1659 | No |
| 128 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6768 | 0.003 | 0.1638 | No |
| 129 | ST6GALNAC4 | ST6GALNAC4 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 | 6968 | 0.000 | 0.1487 | No |
| 130 | IHPK2 | IHPK2 Entrez, Source | inositol hexaphosphate kinase 2 | 7190 | -0.003 | 0.1320 | No |
| 131 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 7269 | -0.004 | 0.1262 | No |
| 132 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 7382 | -0.006 | 0.1179 | No |
| 133 | NEU3 | NEU3 Entrez, Source | sialidase 3 (membrane sialidase) | 7444 | -0.007 | 0.1135 | No |
| 134 | PLAA | PLAA Entrez, Source | phospholipase A2-activating protein | 7627 | -0.009 | 0.1000 | No |
| 135 | SERINC1 | SERINC1 Entrez, Source | serine incorporator 1 | 7788 | -0.012 | 0.0883 | No |
| 136 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7883 | -0.013 | 0.0817 | No |
| 137 | PIGV | PIGV Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class V | 7912 | -0.014 | 0.0801 | No |
| 138 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.0713 | No |
| 139 | RETSAT | RETSAT Entrez, Source | retinol saturase (all-trans-retinol 13,14-reductase) | 8072 | -0.016 | 0.0692 | No |
| 140 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8089 | -0.016 | 0.0686 | No |
| 141 | SULT2A1 | SULT2A1 Entrez, Source | sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 | 8105 | -0.016 | 0.0681 | No |
| 142 | UCP3 | UCP3 Entrez, Source | uncoupling protein 3 (mitochondrial, proton carrier) | 8164 | -0.017 | 0.0644 | No |
| 143 | LRP2 | LRP2 Entrez, Source | low density lipoprotein-related protein 2 | 8183 | -0.018 | 0.0637 | No |
| 144 | PPAP2A | PPAP2A Entrez, Source | phosphatidic acid phosphatase type 2A | 8226 | -0.018 | 0.0612 | No |
| 145 | ABCB4 | ABCB4 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 4 | 8306 | -0.020 | 0.0559 | No |
| 146 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 8460 | -0.022 | 0.0451 | No |
| 147 | DPM2 | DPM2 Entrez, Source | dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit | 8478 | -0.023 | 0.0447 | No |
| 148 | STUB1 | STUB1 Entrez, Source | STIP1 homology and U-box containing protein 1 | 8548 | -0.024 | 0.0404 | No |
| 149 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9114 | -0.034 | -0.0013 | No |
| 150 | PIGS | PIGS Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class S | 9263 | -0.037 | -0.0111 | No |
| 151 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 9316 | -0.038 | -0.0136 | No |
| 152 | PPT1 | PPT1 Entrez, Source | palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) | 9419 | -0.040 | -0.0199 | No |
| 153 | ASAH2 | ASAH2 Entrez, Source | N-acylsphingosine amidohydrolase (non-lysosomal ceramidase) 2 | 9653 | -0.045 | -0.0359 | No |
| 154 | APOD | APOD Entrez, Source | apolipoprotein D | 9731 | -0.046 | -0.0400 | No |
| 155 | SMPD2 | SMPD2 Entrez, Source | sphingomyelin phosphodiesterase 2, neutral membrane (neutral sphingomyelinase) | 9756 | -0.047 | -0.0400 | No |
| 156 | NR0B1 | NR0B1 Entrez, Source | nuclear receptor subfamily 0, group B, member 1 | 9783 | -0.047 | -0.0402 | No |
| 157 | IMPA1 | IMPA1 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 1 | 9804 | -0.048 | -0.0399 | No |
| 158 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 9829 | -0.048 | -0.0399 | No |
| 159 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 9922 | -0.051 | -0.0449 | No |
| 160 | B4GALT4 | B4GALT4 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 4 | 9970 | -0.052 | -0.0465 | No |
| 161 | PLCD1 | PLCD1 Entrez, Source | phospholipase C, delta 1 | 10095 | -0.054 | -0.0539 | No |
| 162 | CYP11A1 | CYP11A1 Entrez, Source | cytochrome P450, family 11, subfamily A, polypeptide 1 | 10102 | -0.054 | -0.0523 | No |
| 163 | CYP3A4 | CYP3A4 Entrez, Source | cytochrome P450, family 3, subfamily A, polypeptide 4 | 10311 | -0.059 | -0.0659 | No |
| 164 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 10430 | -0.062 | -0.0725 | No |
| 165 | PNLIPRP2 | PNLIPRP2 Entrez, Source | pancreatic lipase-related protein 2 | 10564 | -0.066 | -0.0801 | No |
| 166 | CEL | CEL Entrez, Source | carboxyl ester lipase (bile salt-stimulated lipase) | 10808 | -0.073 | -0.0958 | No |
| 167 | LPL | LPL Entrez, Source | lipoprotein lipase | 10988 | -0.078 | -0.1064 | No |
| 168 | YWHAH | YWHAH Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide | 10999 | -0.079 | -0.1042 | No |
| 169 | TSPO | TSPO Entrez, Source | translocator protein (18kDa) | 11004 | -0.079 | -0.1015 | No |
| 170 | CD81 | CD81 Entrez, Source | CD81 molecule | 11022 | -0.079 | -0.0997 | No |
| 171 | IHPK1 | IHPK1 Entrez, Source | inositol hexaphosphate kinase 1 | 11145 | -0.083 | -0.1058 | No |
| 172 | PIP5K1A | PIP5K1A Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, alpha | 11170 | -0.084 | -0.1044 | No |
| 173 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 11277 | -0.089 | -0.1091 | No |
| 174 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | -0.1091 | No |
| 175 | SCARB1 | SCARB1 Entrez, Source | scavenger receptor class B, member 1 | 11498 | -0.099 | -0.1185 | No |
| 176 | SHH | SHH Entrez, Source | sonic hedgehog homolog (Drosophila) | 11581 | -0.103 | -0.1208 | No |
| 177 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 11691 | -0.109 | -0.1249 | No |
| 178 | NPC2 | NPC2 Entrez, Source | Niemann-Pick disease, type C2 | 12096 | -0.131 | -0.1507 | No |
| 179 | SLC27A4 | SLC27A4 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 4 | 12200 | -0.139 | -0.1532 | No |
| 180 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 12262 | -0.145 | -0.1523 | No |
| 181 | AGPAT6 | AGPAT6 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 6 (lysophosphatidic acid acyltransferase, zeta) | 12308 | -0.149 | -0.1500 | No |
| 182 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 12353 | -0.153 | -0.1475 | No |
| 183 | SGPL1 | SGPL1 Entrez, Source | sphingosine-1-phosphate lyase 1 | 12434 | -0.162 | -0.1474 | No |
| 184 | SPHK1 | SPHK1 Entrez, Source | sphingosine kinase 1 | 12507 | -0.172 | -0.1464 | No |
| 185 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 12708 | -0.206 | -0.1537 | No |
| 186 | BTNL3 | BTNL3 Entrez, Source | butyrophilin-like 3 | 12739 | -0.211 | -0.1480 | No |
| 187 | ALDH3B1 | ALDH3B1 Entrez, Source | aldehyde dehydrogenase 3 family, member B1 | 12784 | -0.220 | -0.1429 | No |
| 188 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12906 | -0.254 | -0.1424 | No |
| 189 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 12939 | -0.266 | -0.1347 | No |
| 190 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 13110 | -0.363 | -0.1338 | No |
| 191 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 13162 | -0.417 | -0.1218 | No |
| 192 | SERINC5 | SERINC5 Entrez, Source | serine incorporator 5 | 13225 | -0.503 | -0.1073 | No |
| 193 | SERINC2 | SERINC2 Entrez, Source | serine incorporator 2 | 13234 | -0.520 | -0.0881 | No |
| 194 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 13248 | -0.551 | -0.0681 | No |
| 195 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | -0.0444 | No |
| 196 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | No |