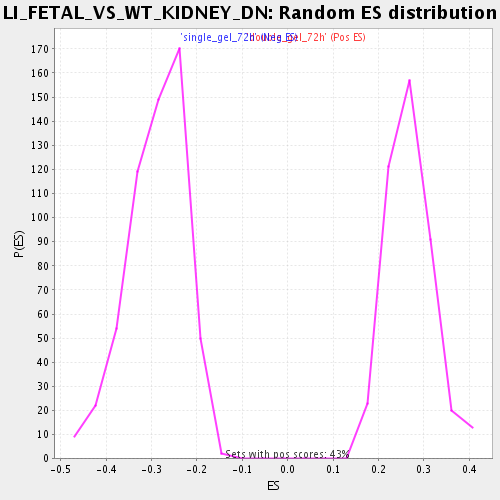
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | LI_FETAL_VS_WT_KIDNEY_DN |
| Enrichment Score (ES) | -0.55805403 |
| Normalized Enrichment Score (NES) | -1.9302919 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.014060197 |
| FWER p-Value | 0.414 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.0164 | No |
| 2 | LRP4 | LRP4 Entrez, Source | low density lipoprotein receptor-related protein 4 | 323 | 0.258 | 0.0299 | No |
| 3 | SLC16A1 | SLC16A1 Entrez, Source | solute carrier family 16, member 1 (monocarboxylic acid transporter 1) | 974 | 0.147 | -0.0054 | No |
| 4 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 984 | 0.146 | 0.0075 | No |
| 5 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 1277 | 0.125 | -0.0028 | No |
| 6 | SHMT2 | SHMT2 Entrez, Source | serine hydroxymethyltransferase 2 (mitochondrial) | 1347 | 0.122 | 0.0033 | No |
| 7 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 1393 | 0.119 | 0.0110 | No |
| 8 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.0139 | No |
| 9 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | -0.0621 | No |
| 10 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 2764 | 0.072 | -0.0678 | No |
| 11 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 2940 | 0.068 | -0.0747 | No |
| 12 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 3161 | 0.063 | -0.0854 | No |
| 13 | DKC1 | DKC1 Entrez, Source | dyskeratosis congenita 1, dyskerin | 3581 | 0.054 | -0.1120 | No |
| 14 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 3875 | 0.049 | -0.1295 | No |
| 15 | BAZ1B | BAZ1B Entrez, Source | bromodomain adjacent to zinc finger domain, 1B | 3938 | 0.048 | -0.1298 | No |
| 16 | ABCA2 | ABCA2 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 2 | 4097 | 0.045 | -0.1375 | No |
| 17 | ACP1 | ACP1 Entrez, Source | acid phosphatase 1, soluble | 4108 | 0.044 | -0.1342 | No |
| 18 | ASNS | ASNS Entrez, Source | asparagine synthetase | 4245 | 0.042 | -0.1405 | No |
| 19 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | -0.1427 | No |
| 20 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 4343 | 0.040 | -0.1402 | No |
| 21 | GARS | GARS Entrez, Source | glycyl-tRNA synthetase | 4370 | 0.040 | -0.1384 | No |
| 22 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 4390 | 0.040 | -0.1362 | No |
| 23 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 5052 | 0.028 | -0.1834 | No |
| 24 | COL2A1 | COL2A1 Entrez, Source | collagen, type II, alpha 1 (primary osteoarthritis, spondyloepiphyseal dysplasia, congenital) | 5191 | 0.026 | -0.1914 | No |
| 25 | STRAP | STRAP Entrez, Source | serine/threonine kinase receptor associated protein | 5363 | 0.024 | -0.2020 | No |
| 26 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5597 | 0.020 | -0.2177 | No |
| 27 | LIG3 | LIG3 Entrez, Source | ligase III, DNA, ATP-dependent | 5865 | 0.016 | -0.2364 | No |
| 28 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 5968 | 0.015 | -0.2427 | No |
| 29 | SV2A | SV2A Entrez, Source | synaptic vesicle glycoprotein 2A | 6094 | 0.013 | -0.2509 | No |
| 30 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 6256 | 0.010 | -0.2621 | No |
| 31 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 6995 | -0.000 | -0.3177 | No |
| 32 | DDX52 | DDX52 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 52 | 7309 | -0.005 | -0.3409 | No |
| 33 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 7542 | -0.008 | -0.3576 | No |
| 34 | CCT3 | CCT3 Entrez, Source | chaperonin containing TCP1, subunit 3 (gamma) | 7557 | -0.008 | -0.3579 | No |
| 35 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 7671 | -0.010 | -0.3655 | No |
| 36 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 7917 | -0.014 | -0.3827 | No |
| 37 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 8206 | -0.018 | -0.4027 | No |
| 38 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 8216 | -0.018 | -0.4017 | No |
| 39 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 8224 | -0.018 | -0.4005 | No |
| 40 | SRPK1 | SRPK1 Entrez, Source | SFRS protein kinase 1 | 8261 | -0.019 | -0.4014 | No |
| 41 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 8733 | -0.027 | -0.4345 | No |
| 42 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 8998 | -0.032 | -0.4514 | No |
| 43 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9129 | -0.035 | -0.4580 | No |
| 44 | CHN1 | CHN1 Entrez, Source | chimerin (chimaerin) 1 | 9568 | -0.043 | -0.4870 | No |
| 45 | TARBP2 | TARBP2 Entrez, Source | Tar (HIV-1) RNA binding protein 2 | 9589 | -0.043 | -0.4845 | No |
| 46 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 9648 | -0.045 | -0.4848 | No |
| 47 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 9737 | -0.046 | -0.4871 | No |
| 48 | PCBP2 | PCBP2 Entrez, Source | poly(rC) binding protein 2 | 10047 | -0.053 | -0.5054 | No |
| 49 | NUP188 | NUP188 Entrez, Source | nucleoporin 188kDa | 10164 | -0.056 | -0.5090 | No |
| 50 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 10720 | -0.070 | -0.5443 | No |
| 51 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 10903 | -0.075 | -0.5511 | Yes |
| 52 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 10934 | -0.076 | -0.5462 | Yes |
| 53 | CHST1 | CHST1 Entrez, Source | carbohydrate (keratan sulfate Gal-6) sulfotransferase 1 | 10942 | -0.077 | -0.5396 | Yes |
| 54 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 11071 | -0.081 | -0.5417 | Yes |
| 55 | SS18 | SS18 Entrez, Source | synovial sarcoma translocation, chromosome 18 | 11166 | -0.084 | -0.5410 | Yes |
| 56 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 11249 | -0.088 | -0.5390 | Yes |
| 57 | SPAG5 | SPAG5 Entrez, Source | sperm associated antigen 5 | 11402 | -0.095 | -0.5416 | Yes |
| 58 | FAM62A | FAM62A Entrez, Source | family with sequence similarity 62 (C2 domain containing), member A | 11541 | -0.101 | -0.5426 | Yes |
| 59 | CKAP5 | CKAP5 Entrez, Source | cytoskeleton associated protein 5 | 11647 | -0.106 | -0.5406 | Yes |
| 60 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 11649 | -0.107 | -0.5308 | Yes |
| 61 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 11676 | -0.108 | -0.5227 | Yes |
| 62 | YWHAZ | YWHAZ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide | 11724 | -0.110 | -0.5160 | Yes |
| 63 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 11731 | -0.111 | -0.5061 | Yes |
| 64 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 11752 | -0.111 | -0.4973 | Yes |
| 65 | AURKA | AURKA Entrez, Source | aurora kinase A | 11917 | -0.119 | -0.4985 | Yes |
| 66 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 12256 | -0.144 | -0.5106 | Yes |
| 67 | GPC1 | GPC1 Entrez, Source | glypican 1 | 12419 | -0.160 | -0.5079 | Yes |
| 68 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 12432 | -0.162 | -0.4938 | Yes |
| 69 | CACNB3 | CACNB3 Entrez, Source | calcium channel, voltage-dependent, beta 3 subunit | 12633 | -0.191 | -0.4911 | Yes |
| 70 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 12752 | -0.214 | -0.4801 | Yes |
| 71 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 12767 | -0.216 | -0.4610 | Yes |
| 72 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 12774 | -0.218 | -0.4412 | Yes |
| 73 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 12815 | -0.228 | -0.4230 | Yes |
| 74 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 12851 | -0.238 | -0.4035 | Yes |
| 75 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12940 | -0.266 | -0.3854 | Yes |
| 76 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12949 | -0.272 | -0.3607 | Yes |
| 77 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 13078 | -0.337 | -0.3390 | Yes |
| 78 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 13080 | -0.340 | -0.3074 | Yes |
| 79 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 13086 | -0.343 | -0.2758 | Yes |
| 80 | AURKB | AURKB Entrez, Source | aurora kinase B | 13115 | -0.366 | -0.2439 | Yes |
| 81 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 13129 | -0.381 | -0.2094 | Yes |
| 82 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 13140 | -0.392 | -0.1737 | Yes |
| 83 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13267 | -0.614 | -0.1261 | Yes |
| 84 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 13280 | -0.659 | -0.0656 | Yes |
| 85 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 13302 | -0.755 | 0.0030 | Yes |