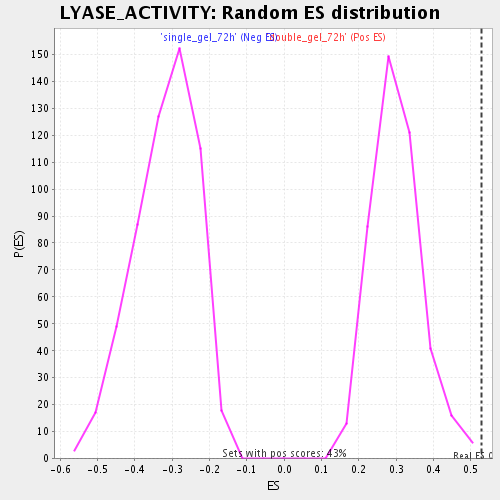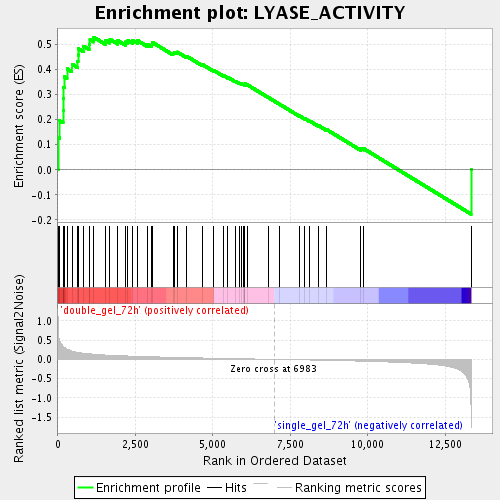
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | LYASE_ACTIVITY |
| Enrichment Score (ES) | 0.5280013 |
| Normalized Enrichment Score (NES) | 1.7615169 |
| Nominal p-value | 0.0023148148 |
| FDR q-value | 0.021993784 |
| FWER p-Value | 0.971 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 6 | 0.908 | 0.1292 | Yes |
| 2 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 54 | 0.488 | 0.1953 | Yes |
| 3 | GUCY2C | GUCY2C Entrez, Source | guanylate cyclase 2C (heat stable enterotoxin receptor) | 167 | 0.341 | 0.2356 | Yes |
| 4 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 174 | 0.334 | 0.2827 | Yes |
| 5 | ASL | ASL Entrez, Source | argininosuccinate lyase | 183 | 0.323 | 0.3283 | Yes |
| 6 | ACMSD | ACMSD Entrez, Source | aminocarboxymuconate semialdehyde decarboxylase | 208 | 0.306 | 0.3703 | Yes |
| 7 | CBS | CBS Entrez, Source | cystathionine-beta-synthase | 294 | 0.267 | 0.4021 | Yes |
| 8 | SCLY | SCLY Entrez, Source | selenocysteine lyase | 454 | 0.214 | 0.4207 | Yes |
| 9 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.4333 | Yes |
| 10 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 651 | 0.179 | 0.4576 | Yes |
| 11 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 662 | 0.178 | 0.4823 | Yes |
| 12 | ALAD | ALAD Entrez, Source | aminolevulinate, delta-, dehydratase | 826 | 0.160 | 0.4929 | Yes |
| 13 | ALDOB | ALDOB Entrez, Source | aldolase B, fructose-bisphosphate | 1007 | 0.144 | 0.4999 | Yes |
| 14 | CA5A | CA5A Entrez, Source | carbonic anhydrase VA, mitochondrial | 1034 | 0.142 | 0.5182 | Yes |
| 15 | CRYM | CRYM Entrez, Source | crystallin, mu | 1157 | 0.133 | 0.5280 | Yes |
| 16 | HDC | HDC Entrez, Source | histidine decarboxylase | 1527 | 0.112 | 0.5162 | No |
| 17 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.5200 | No |
| 18 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 1935 | 0.096 | 0.5145 | No |
| 19 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2174 | 0.089 | 0.5093 | No |
| 20 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 2259 | 0.086 | 0.5153 | No |
| 21 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 2407 | 0.082 | 0.5160 | No |
| 22 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 2566 | 0.078 | 0.5152 | No |
| 23 | ENO2 | ENO2 Entrez, Source | enolase 2 (gamma, neuronal) | 2893 | 0.069 | 0.5005 | No |
| 24 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 3035 | 0.066 | 0.4993 | No |
| 25 | GUCY1A2 | GUCY1A2 Entrez, Source | guanylate cyclase 1, soluble, alpha 2 | 3052 | 0.065 | 0.5074 | No |
| 26 | CA4 | CA4 Entrez, Source | carbonic anhydrase IV | 3716 | 0.052 | 0.4649 | No |
| 27 | GUCY2F | GUCY2F Entrez, Source | guanylate cyclase 2F, retinal | 3762 | 0.051 | 0.4688 | No |
| 28 | ENO3 | ENO3 Entrez, Source | enolase 3 (beta, muscle) | 3849 | 0.049 | 0.4693 | No |
| 29 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.4520 | No |
| 30 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.4185 | No |
| 31 | GGCX | GGCX Entrez, Source | gamma-glutamyl carboxylase | 5019 | 0.029 | 0.3968 | No |
| 32 | CA5B | CA5B Entrez, Source | carbonic anhydrase VB, mitochondrial | 5338 | 0.024 | 0.3763 | No |
| 33 | ODC1 | ODC1 Entrez, Source | ornithine decarboxylase 1 | 5487 | 0.022 | 0.3683 | No |
| 34 | ALDOC | ALDOC Entrez, Source | aldolase C, fructose-bisphosphate | 5718 | 0.018 | 0.3536 | No |
| 35 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.3454 | No |
| 36 | ADCY8 | ADCY8 Entrez, Source | adenylate cyclase 8 (brain) | 5925 | 0.015 | 0.3427 | No |
| 37 | GUCY1B3 | GUCY1B3 Entrez, Source | guanylate cyclase 1, soluble, beta 3 | 5991 | 0.014 | 0.3399 | No |
| 38 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 6026 | 0.014 | 0.3393 | No |
| 39 | GLO1 | GLO1 Entrez, Source | glyoxalase I | 6031 | 0.014 | 0.3409 | No |
| 40 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 6037 | 0.014 | 0.3425 | No |
| 41 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 6118 | 0.013 | 0.3383 | No |
| 42 | GUCY2D | GUCY2D Entrez, Source | guanylate cyclase 2D, membrane (retina-specific) | 6795 | 0.003 | 0.2878 | No |
| 43 | CA2 | CA2 Entrez, Source | carbonic anhydrase II | 7159 | -0.002 | 0.2608 | No |
| 44 | UROD | UROD Entrez, Source | uroporphyrinogen decarboxylase | 7789 | -0.012 | 0.2152 | No |
| 45 | ADCY10 | 7969 | -0.015 | 0.2038 | No | ||
| 46 | LTC4S | LTC4S Entrez, Source | leukotriene C4 synthase | 8127 | -0.017 | 0.1944 | No |
| 47 | DDC | DDC Entrez, Source | dopa decarboxylase (aromatic L-amino acid decarboxylase) | 8401 | -0.021 | 0.1769 | No |
| 48 | SDS | SDS Entrez, Source | serine dehydratase | 8683 | -0.026 | 0.1595 | No |
| 49 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 9779 | -0.047 | 0.0839 | No |
| 50 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | 0.0858 | No |
| 51 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13334 | -1.240 | 0.0006 | No |