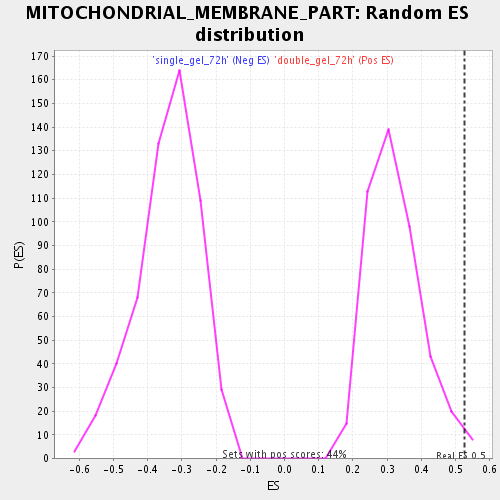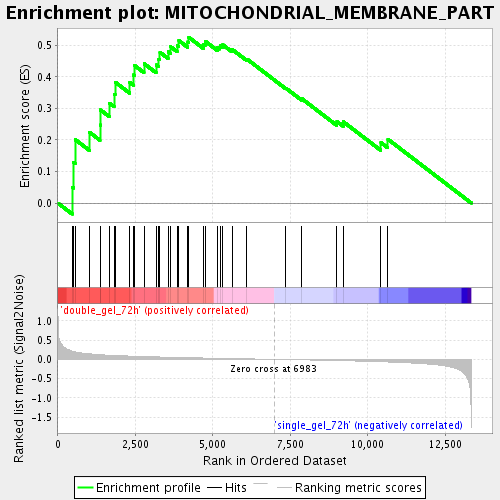
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | MITOCHONDRIAL_MEMBRANE_PART |
| Enrichment Score (ES) | 0.5252808 |
| Normalized Enrichment Score (NES) | 1.6315124 |
| Nominal p-value | 0.018348623 |
| FDR q-value | 0.058501583 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TIMM9 | TIMM9 Entrez, Source | translocase of inner mitochondrial membrane 9 homolog (yeast) | 463 | 0.212 | 0.0504 | Yes |
| 2 | BCS1L | BCS1L Entrez, Source | BCS1-like (yeast) | 504 | 0.201 | 0.1285 | Yes |
| 3 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 561 | 0.191 | 0.2011 | Yes |
| 4 | TIMM10 | TIMM10 Entrez, Source | translocase of inner mitochondrial membrane 10 homolog (yeast) | 1021 | 0.143 | 0.2242 | Yes |
| 5 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 1360 | 0.121 | 0.2476 | Yes |
| 6 | NDUFS7 | NDUFS7 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 7, 20kDa (NADH-coenzyme Q reductase) | 1371 | 0.121 | 0.2954 | Yes |
| 7 | NDUFA1 | NDUFA1 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 1, 7.5kDa | 1663 | 0.106 | 0.3163 | Yes |
| 8 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 1817 | 0.101 | 0.3454 | Yes |
| 9 | SURF1 | SURF1 Entrez, Source | surfeit 1 | 1848 | 0.100 | 0.3832 | Yes |
| 10 | ATP5G2 | ATP5G2 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C2 (subunit 9) | 2318 | 0.085 | 0.3820 | Yes |
| 11 | ATP5C1 | ATP5C1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, gamma polypeptide 1 | 2437 | 0.082 | 0.4060 | Yes |
| 12 | NDUFV1 | NDUFV1 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 1, 51kDa | 2468 | 0.081 | 0.4362 | Yes |
| 13 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2780 | 0.072 | 0.4418 | Yes |
| 14 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 3172 | 0.063 | 0.4377 | Yes |
| 15 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 3253 | 0.061 | 0.4563 | Yes |
| 16 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 3292 | 0.060 | 0.4777 | Yes |
| 17 | TIMM8B | TIMM8B Entrez, Source | translocase of inner mitochondrial membrane 8 homolog B (yeast) | 3559 | 0.055 | 0.4797 | Yes |
| 18 | ATP5E | ATP5E Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit | 3634 | 0.053 | 0.4955 | Yes |
| 19 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 3850 | 0.049 | 0.4991 | Yes |
| 20 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 3903 | 0.048 | 0.5147 | Yes |
| 21 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 4193 | 0.043 | 0.5103 | Yes |
| 22 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 4222 | 0.043 | 0.5253 | Yes |
| 23 | FIS1 | FIS1 Entrez, Source | fission 1 (mitochondrial outer membrane) homolog (S. cerevisiae) | 4704 | 0.034 | 0.5029 | No |
| 24 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 4753 | 0.033 | 0.5128 | No |
| 25 | TIMM13 | TIMM13 Entrez, Source | translocase of inner mitochondrial membrane 13 homolog (yeast) | 5146 | 0.027 | 0.4942 | No |
| 26 | UQCRB | UQCRB Entrez, Source | ubiquinol-cytochrome c reductase binding protein | 5231 | 0.026 | 0.4983 | No |
| 27 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 5323 | 0.024 | 0.5012 | No |
| 28 | OPA1 | OPA1 Entrez, Source | optic atrophy 1 (autosomal dominant) | 5620 | 0.020 | 0.4871 | No |
| 29 | TOMM22 | TOMM22 Entrez, Source | translocase of outer mitochondrial membrane 22 homolog (yeast) | 6098 | 0.013 | 0.4564 | No |
| 30 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 7358 | -0.005 | 0.3639 | No |
| 31 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 7875 | -0.013 | 0.3305 | No |
| 32 | MFN2 | MFN2 Entrez, Source | mitofusin 2 | 9004 | -0.032 | 0.2587 | No |
| 33 | COX6B2 | COX6B2 Entrez, Source | cytochrome c oxidase subunit VIb polypeptide 2 (testis) | 9208 | -0.036 | 0.2579 | No |
| 34 | COX15 | COX15 Entrez, Source | COX15 homolog, cytochrome c oxidase assembly protein (yeast) | 10424 | -0.062 | 0.1916 | No |
| 35 | RHOT2 | RHOT2 Entrez, Source | ras homolog gene family, member T2 | 10650 | -0.068 | 0.2023 | No |