Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
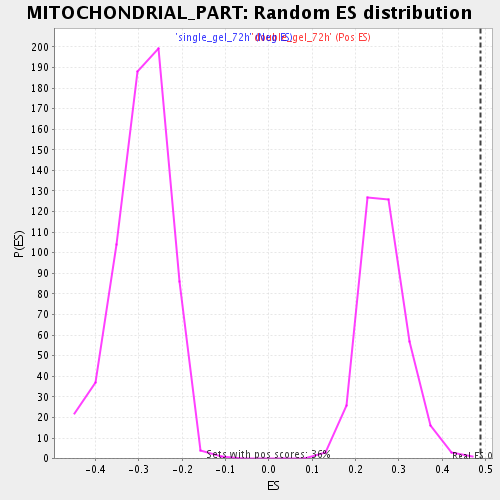
| GeneSet | MITOCHONDRIAL_PART |
| Enrichment Score (ES) | 0.4881155 |
| Normalized Enrichment Score (NES) | 1.8367475 |
| Nominal p-value | 0.0027855153 |
| FDR q-value | 0.011380916 |
| FWER p-Value | 0.759 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.0217 | Yes |
| 2 | TIMM9 | TIMM9 Entrez, Source | translocase of inner mitochondrial membrane 9 homolog (yeast) | 463 | 0.212 | 0.0417 | Yes |
| 3 | BCS1L | BCS1L Entrez, Source | BCS1-like (yeast) | 504 | 0.201 | 0.0704 | Yes |
| 4 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 561 | 0.191 | 0.0963 | Yes |
| 5 | SLC25A15 | SLC25A15 Entrez, Source | solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15 | 595 | 0.186 | 0.1233 | Yes |
| 6 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 651 | 0.179 | 0.1474 | Yes |
| 7 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 662 | 0.178 | 0.1748 | Yes |
| 8 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 746 | 0.168 | 0.1950 | Yes |
| 9 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 854 | 0.157 | 0.2117 | Yes |
| 10 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 868 | 0.156 | 0.2354 | Yes |
| 11 | MRPS15 | MRPS15 Entrez, Source | mitochondrial ribosomal protein S15 | 936 | 0.150 | 0.2540 | Yes |
| 12 | DNAJA3 | DNAJA3 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 3 | 972 | 0.147 | 0.2746 | Yes |
| 13 | TIMM10 | TIMM10 Entrez, Source | translocase of inner mitochondrial membrane 10 homolog (yeast) | 1021 | 0.143 | 0.2936 | Yes |
| 14 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 1360 | 0.121 | 0.2872 | Yes |
| 15 | NDUFS7 | NDUFS7 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 7, 20kDa (NADH-coenzyme Q reductase) | 1371 | 0.121 | 0.3055 | Yes |
| 16 | TFB2M | TFB2M Entrez, Source | transcription factor B2, mitochondrial | 1604 | 0.109 | 0.3051 | Yes |
| 17 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 1652 | 0.107 | 0.3184 | Yes |
| 18 | NDUFA1 | NDUFA1 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 1, 7.5kDa | 1663 | 0.106 | 0.3344 | Yes |
| 19 | MAOB | MAOB Entrez, Source | monoamine oxidase B | 1779 | 0.102 | 0.3418 | Yes |
| 20 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 1817 | 0.101 | 0.3550 | Yes |
| 21 | SURF1 | SURF1 Entrez, Source | surfeit 1 | 1848 | 0.100 | 0.3684 | Yes |
| 22 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 1931 | 0.096 | 0.3774 | Yes |
| 23 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 2100 | 0.091 | 0.3791 | Yes |
| 24 | SLC25A22 | SLC25A22 Entrez, Source | solute carrier family 25 (mitochondrial carrier: glutamate), member 22 | 2140 | 0.090 | 0.3903 | Yes |
| 25 | ATP5G2 | ATP5G2 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C2 (subunit 9) | 2318 | 0.085 | 0.3903 | Yes |
| 26 | ATP5C1 | ATP5C1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, gamma polypeptide 1 | 2437 | 0.082 | 0.3943 | Yes |
| 27 | NDUFV1 | NDUFV1 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 1, 51kDa | 2468 | 0.081 | 0.4048 | Yes |
| 28 | SDHD | SDHD Entrez, Source | succinate dehydrogenase complex, subunit D, integral membrane protein | 2529 | 0.079 | 0.4127 | Yes |
| 29 | ABCB8 | ABCB8 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 8 | 2624 | 0.076 | 0.4175 | Yes |
| 30 | GRPEL1 | GRPEL1 Entrez, Source | GrpE-like 1, mitochondrial (E. coli) | 2691 | 0.074 | 0.4242 | Yes |
| 31 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2780 | 0.072 | 0.4289 | Yes |
| 32 | SLC25A1 | SLC25A1 Entrez, Source | solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 | 2912 | 0.069 | 0.4298 | Yes |
| 33 | PHB2 | PHB2 Entrez, Source | prohibitin 2 | 3135 | 0.064 | 0.4231 | Yes |
| 34 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 3172 | 0.063 | 0.4303 | Yes |
| 35 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 3253 | 0.061 | 0.4339 | Yes |
| 36 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 3292 | 0.060 | 0.4406 | Yes |
| 37 | ABCB6 | ABCB6 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 6 | 3356 | 0.059 | 0.4451 | Yes |
| 38 | TIMM8B | TIMM8B Entrez, Source | translocase of inner mitochondrial membrane 8 homolog B (yeast) | 3559 | 0.055 | 0.4385 | Yes |
| 39 | ATP5E | ATP5E Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit | 3634 | 0.053 | 0.4413 | Yes |
| 40 | CENTA2 | CENTA2 Entrez, Source | centaurin, alpha 2 | 3650 | 0.053 | 0.4485 | Yes |
| 41 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 3842 | 0.049 | 0.4419 | Yes |
| 42 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 3850 | 0.049 | 0.4491 | Yes |
| 43 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 3903 | 0.048 | 0.4528 | Yes |
| 44 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 3912 | 0.048 | 0.4598 | Yes |
| 45 | RAF1 | RAF1 Entrez, Source | v-raf-1 murine leukemia viral oncogene homolog 1 | 3941 | 0.048 | 0.4652 | Yes |
| 46 | SLC25A11 | SLC25A11 Entrez, Source | solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 | 4171 | 0.043 | 0.4548 | Yes |
| 47 | CS | CS Entrez, Source | citrate synthase | 4175 | 0.043 | 0.4614 | Yes |
| 48 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 4193 | 0.043 | 0.4669 | Yes |
| 49 | ATP5J | ATP5J Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 | 4194 | 0.043 | 0.4737 | Yes |
| 50 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 4222 | 0.043 | 0.4784 | Yes |
| 51 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.4791 | Yes |
| 52 | PHB | PHB Entrez, Source | prohibitin | 4309 | 0.041 | 0.4849 | Yes |
| 53 | MRPL23 | MRPL23 Entrez, Source | mitochondrial ribosomal protein L23 | 4352 | 0.040 | 0.4881 | Yes |
| 54 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.4693 | No |
| 55 | FIS1 | FIS1 Entrez, Source | fission 1 (mitochondrial outer membrane) homolog (S. cerevisiae) | 4704 | 0.034 | 0.4726 | No |
| 56 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 4753 | 0.033 | 0.4742 | No |
| 57 | MRPL41 | MRPL41 Entrez, Source | mitochondrial ribosomal protein L41 | 4921 | 0.030 | 0.4664 | No |
| 58 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 4997 | 0.029 | 0.4654 | No |
| 59 | TIMM13 | TIMM13 Entrez, Source | translocase of inner mitochondrial membrane 13 homolog (yeast) | 5146 | 0.027 | 0.4585 | No |
| 60 | UQCRB | UQCRB Entrez, Source | ubiquinol-cytochrome c reductase binding protein | 5231 | 0.026 | 0.4562 | No |
| 61 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 5252 | 0.025 | 0.4587 | No |
| 62 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 5323 | 0.024 | 0.4573 | No |
| 63 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 5336 | 0.024 | 0.4602 | No |
| 64 | ALAS2 | ALAS2 Entrez, Source | aminolevulinate, delta-, synthase 2 (sideroblastic/hypochromic anemia) | 5606 | 0.020 | 0.4431 | No |
| 65 | OPA1 | OPA1 Entrez, Source | optic atrophy 1 (autosomal dominant) | 5620 | 0.020 | 0.4452 | No |
| 66 | IMMT | IMMT Entrez, Source | inner membrane protein, mitochondrial (mitofilin) | 5652 | 0.020 | 0.4460 | No |
| 67 | VDAC2 | VDAC2 Entrez, Source | voltage-dependent anion channel 2 | 5987 | 0.014 | 0.4230 | No |
| 68 | TOMM22 | TOMM22 Entrez, Source | translocase of outer mitochondrial membrane 22 homolog (yeast) | 6098 | 0.013 | 0.4168 | No |
| 69 | MTX2 | MTX2 Entrez, Source | metaxin 2 | 6206 | 0.011 | 0.4105 | No |
| 70 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 6707 | 0.004 | 0.3733 | No |
| 71 | PMPCA | PMPCA Entrez, Source | peptidase (mitochondrial processing) alpha | 7146 | -0.002 | 0.3407 | No |
| 72 | VDAC1 | VDAC1 Entrez, Source | voltage-dependent anion channel 1 | 7242 | -0.004 | 0.3341 | No |
| 73 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 7358 | -0.005 | 0.3262 | No |
| 74 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 7875 | -0.013 | 0.2894 | No |
| 75 | TIMM44 | TIMM44 Entrez, Source | translocase of inner mitochondrial membrane 44 homolog (yeast) | 8155 | -0.017 | 0.2711 | No |
| 76 | UCP3 | UCP3 Entrez, Source | uncoupling protein 3 (mitochondrial, proton carrier) | 8164 | -0.017 | 0.2732 | No |
| 77 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 8887 | -0.030 | 0.2234 | No |
| 78 | MFN2 | MFN2 Entrez, Source | mitofusin 2 | 9004 | -0.032 | 0.2198 | No |
| 79 | COX6B2 | COX6B2 Entrez, Source | cytochrome c oxidase subunit VIb polypeptide 2 (testis) | 9208 | -0.036 | 0.2101 | No |
| 80 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 9244 | -0.037 | 0.2133 | No |
| 81 | VDAC3 | VDAC3 Entrez, Source | voltage-dependent anion channel 3 | 9603 | -0.043 | 0.1931 | No |
| 82 | MRPS18A | MRPS18A Entrez, Source | mitochondrial ribosomal protein S18A | 9675 | -0.045 | 0.1949 | No |
| 83 | BAX | BAX Entrez, Source | BCL2-associated X protein | 9773 | -0.047 | 0.1950 | No |
| 84 | CASP7 | CASP7 Entrez, Source | caspase 7, apoptosis-related cysteine peptidase | 10338 | -0.060 | 0.1619 | No |
| 85 | COX15 | COX15 Entrez, Source | COX15 homolog, cytochrome c oxidase assembly protein (yeast) | 10424 | -0.062 | 0.1653 | No |
| 86 | RHOT2 | RHOT2 Entrez, Source | ras homolog gene family, member T2 | 10650 | -0.068 | 0.1591 | No |
| 87 | SLC25A3 | SLC25A3 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | 11129 | -0.083 | 0.1362 | No |
| 88 | HTRA2 | HTRA2 Entrez, Source | HtrA serine peptidase 2 | 11309 | -0.091 | 0.1370 | No |
| 89 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 11580 | -0.103 | 0.1329 | No |