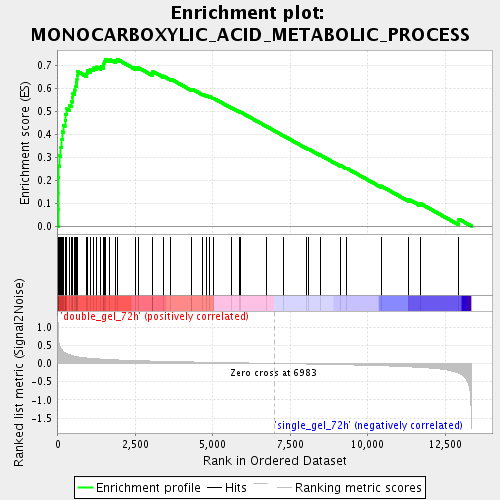
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | MONOCARBOXYLIC_ACID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.72834957 |
| Normalized Enrichment Score (NES) | 2.5386128 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 9 | 0.779 | 0.0728 | Yes |
| 2 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 13 | 0.740 | 0.1423 | Yes |
| 3 | FTCD | FTCD Entrez, Source | formiminotransferase cyclodeaminase | 15 | 0.731 | 0.2113 | Yes |
| 4 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 33 | 0.554 | 0.2623 | Yes |
| 5 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 54 | 0.488 | 0.3068 | Yes |
| 6 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 95 | 0.419 | 0.3433 | Yes |
| 7 | AGXT | AGXT Entrez, Source | alanine-glyoxylate aminotransferase (oxalosis I; hyperoxaluria I; glycolicaciduria; serine-pyruvate aminotransferase) | 128 | 0.384 | 0.3771 | Yes |
| 8 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 140 | 0.368 | 0.4109 | Yes |
| 9 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.4384 | Yes |
| 10 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.4625 | Yes |
| 11 | NR1H4 | NR1H4 Entrez, Source | nuclear receptor subfamily 1, group H, member 4 | 246 | 0.288 | 0.4886 | Yes |
| 12 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.5116 | Yes |
| 13 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 378 | 0.240 | 0.5272 | Yes |
| 14 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 444 | 0.218 | 0.5428 | Yes |
| 15 | ALDH1L1 | ALDH1L1 Entrez, Source | aldehyde dehydrogenase 1 family, member L1 | 465 | 0.211 | 0.5612 | Yes |
| 16 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 482 | 0.207 | 0.5795 | Yes |
| 17 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 536 | 0.195 | 0.5939 | Yes |
| 18 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 553 | 0.192 | 0.6108 | Yes |
| 19 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 597 | 0.186 | 0.6252 | Yes |
| 20 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.6415 | Yes |
| 21 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 619 | 0.184 | 0.6584 | Yes |
| 22 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.6746 | Yes |
| 23 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 933 | 0.150 | 0.6662 | Yes |
| 24 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.6790 | Yes |
| 25 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 1046 | 0.140 | 0.6852 | Yes |
| 26 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1151 | 0.133 | 0.6899 | Yes |
| 27 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 1252 | 0.127 | 0.6943 | Yes |
| 28 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 1387 | 0.119 | 0.6955 | Yes |
| 29 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 1471 | 0.115 | 0.7001 | Yes |
| 30 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1480 | 0.114 | 0.7102 | Yes |
| 31 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.7199 | Yes |
| 32 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.7283 | Yes |
| 33 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.7268 | No |
| 34 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.7226 | No |
| 35 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 1926 | 0.096 | 0.7267 | No |
| 36 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 2496 | 0.080 | 0.6914 | No |
| 37 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | 0.6910 | No |
| 38 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 3046 | 0.066 | 0.6635 | No |
| 39 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.6689 | No |
| 40 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 3063 | 0.065 | 0.6747 | No |
| 41 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3398 | 0.058 | 0.6550 | No |
| 42 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 3642 | 0.053 | 0.6417 | No |
| 43 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.5962 | No |
| 44 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | 0.5981 | No |
| 45 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.5752 | No |
| 46 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.5687 | No |
| 47 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 4887 | 0.031 | 0.5653 | No |
| 48 | ARPP-19 | ARPP-19 Entrez, Source | - | 5020 | 0.029 | 0.5581 | No |
| 49 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 5602 | 0.020 | 0.5163 | No |
| 50 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.4986 | No |
| 51 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.4981 | No |
| 52 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | 0.4343 | No |
| 53 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 7269 | -0.004 | 0.3948 | No |
| 54 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.3386 | No |
| 55 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8089 | -0.016 | 0.3362 | No |
| 56 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 8460 | -0.022 | 0.3104 | No |
| 57 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9114 | -0.034 | 0.2645 | No |
| 58 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 9316 | -0.038 | 0.2530 | No |
| 59 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 10430 | -0.062 | 0.1750 | No |
| 60 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | 0.1165 | No |
| 61 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 11691 | -0.109 | 0.0992 | No |
| 62 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 12939 | -0.266 | 0.0303 | No |

