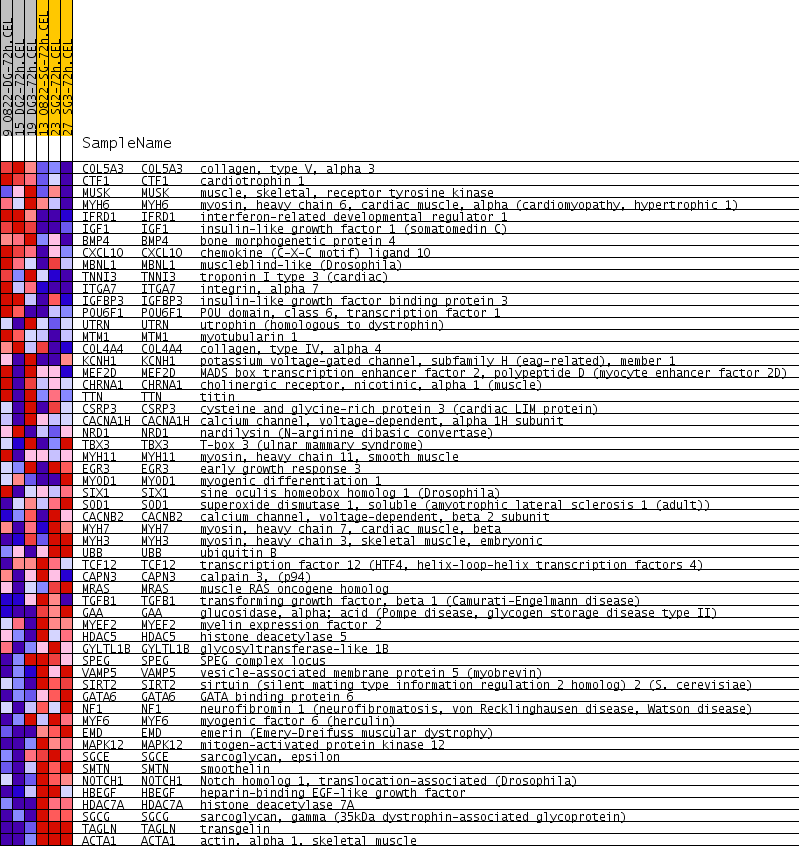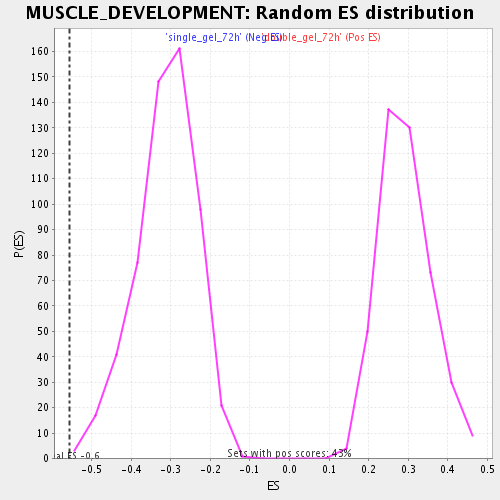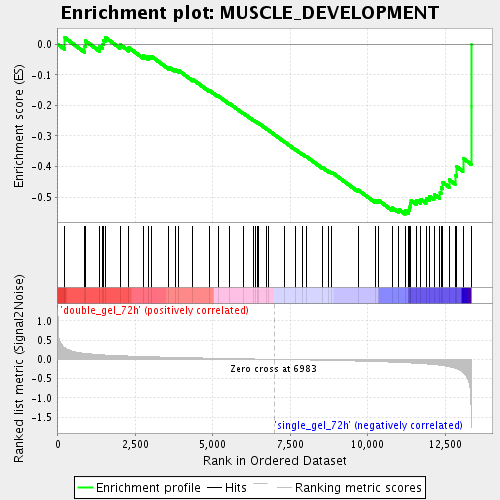
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | MUSCLE_DEVELOPMENT |
| Enrichment Score (ES) | -0.55580914 |
| Normalized Enrichment Score (NES) | -1.7748973 |
| Nominal p-value | 0.0035273368 |
| FDR q-value | 0.041740406 |
| FWER p-Value | 0.995 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL5A3 | COL5A3 Entrez, Source | collagen, type V, alpha 3 | 224 | 0.297 | 0.0213 | No |
| 2 | CTF1 | CTF1 Entrez, Source | cardiotrophin 1 | 869 | 0.156 | -0.0072 | No |
| 3 | MUSK | MUSK Entrez, Source | muscle, skeletal, receptor tyrosine kinase | 887 | 0.154 | 0.0113 | No |
| 4 | MYH6 | MYH6 Entrez, Source | myosin, heavy chain 6, cardiac muscle, alpha (cardiomyopathy, hypertrophic 1) | 1352 | 0.121 | -0.0080 | No |
| 5 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1449 | 0.116 | -0.0004 | No |
| 6 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 1471 | 0.115 | 0.0127 | No |
| 7 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1526 | 0.112 | 0.0230 | No |
| 8 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 2004 | 0.094 | -0.0008 | No |
| 9 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2291 | 0.085 | -0.0114 | No |
| 10 | TNNI3 | TNNI3 Entrez, Source | troponin I type 3 (cardiac) | 2763 | 0.072 | -0.0375 | No |
| 11 | ITGA7 | ITGA7 Entrez, Source | integrin, alpha 7 | 2929 | 0.068 | -0.0412 | No |
| 12 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 3018 | 0.066 | -0.0393 | No |
| 13 | POU6F1 | POU6F1 Entrez, Source | POU domain, class 6, transcription factor 1 | 3583 | 0.054 | -0.0748 | No |
| 14 | UTRN | UTRN Entrez, Source | utrophin (homologous to dystrophin) | 3787 | 0.050 | -0.0837 | No |
| 15 | MTM1 | MTM1 Entrez, Source | myotubularin 1 | 3901 | 0.048 | -0.0860 | No |
| 16 | COL4A4 | COL4A4 Entrez, Source | collagen, type IV, alpha 4 | 4358 | 0.040 | -0.1152 | No |
| 17 | KCNH1 | KCNH1 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 1 | 4901 | 0.031 | -0.1520 | No |
| 18 | MEF2D | MEF2D Entrez, Source | MADS box transcription enhancer factor 2, polypeptide D (myocyte enhancer factor 2D) | 5169 | 0.027 | -0.1687 | No |
| 19 | CHRNA1 | CHRNA1 Entrez, Source | cholinergic receptor, nicotinic, alpha 1 (muscle) | 5541 | 0.021 | -0.1939 | No |
| 20 | TTN | TTN Entrez, Source | titin | 5994 | 0.014 | -0.2261 | No |
| 21 | CSRP3 | CSRP3 Entrez, Source | cysteine and glycine-rich protein 3 (cardiac LIM protein) | 6325 | 0.009 | -0.2497 | No |
| 22 | CACNA1H | CACNA1H Entrez, Source | calcium channel, voltage-dependent, alpha 1H subunit | 6388 | 0.008 | -0.2534 | No |
| 23 | NRD1 | NRD1 Entrez, Source | nardilysin (N-arginine dibasic convertase) | 6449 | 0.007 | -0.2569 | No |
| 24 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 6476 | 0.007 | -0.2580 | No |
| 25 | MYH11 | MYH11 Entrez, Source | myosin, heavy chain 11, smooth muscle | 6721 | 0.004 | -0.2760 | No |
| 26 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 6801 | 0.003 | -0.2816 | No |
| 27 | MYOD1 | MYOD1 Entrez, Source | myogenic differentiation 1 | 7324 | -0.005 | -0.3202 | No |
| 28 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 7671 | -0.010 | -0.3450 | No |
| 29 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7883 | -0.013 | -0.3591 | No |
| 30 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 8013 | -0.015 | -0.3669 | No |
| 31 | MYH7 | MYH7 Entrez, Source | myosin, heavy chain 7, cardiac muscle, beta | 8532 | -0.024 | -0.4029 | No |
| 32 | MYH3 | MYH3 Entrez, Source | myosin, heavy chain 3, skeletal muscle, embryonic | 8743 | -0.027 | -0.4152 | No |
| 33 | UBB | UBB Entrez, Source | ubiquitin B | 8839 | -0.029 | -0.4186 | No |
| 34 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 9690 | -0.045 | -0.4768 | No |
| 35 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 10237 | -0.058 | -0.5105 | No |
| 36 | MRAS | MRAS Entrez, Source | muscle RAS oncogene homolog | 10349 | -0.060 | -0.5111 | No |
| 37 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 10788 | -0.072 | -0.5348 | No |
| 38 | GAA | GAA Entrez, Source | glucosidase, alpha; acid (Pompe disease, glycogen storage disease type II) | 11007 | -0.079 | -0.5411 | No |
| 39 | MYEF2 | MYEF2 Entrez, Source | myelin expression factor 2 | 11204 | -0.086 | -0.5448 | Yes |
| 40 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 11330 | -0.092 | -0.5424 | Yes |
| 41 | GYLTL1B | GYLTL1B Entrez, Source | glycosyltransferase-like 1B | 11358 | -0.093 | -0.5325 | Yes |
| 42 | SPEG | SPEG Entrez, Source | SPEG complex locus | 11368 | -0.093 | -0.5211 | Yes |
| 43 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 11394 | -0.095 | -0.5109 | Yes |
| 44 | SIRT2 | SIRT2 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 2 (S. cerevisiae) | 11576 | -0.103 | -0.5113 | Yes |
| 45 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 11713 | -0.110 | -0.5074 | Yes |
| 46 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 11882 | -0.118 | -0.5049 | Yes |
| 47 | MYF6 | MYF6 Entrez, Source | myogenic factor 6 (herculin) | 11986 | -0.124 | -0.4968 | Yes |
| 48 | EMD | EMD Entrez, Source | emerin (Emery-Dreifuss muscular dystrophy) | 12141 | -0.133 | -0.4912 | Yes |
| 49 | MAPK12 | MAPK12 Entrez, Source | mitogen-activated protein kinase 12 | 12330 | -0.151 | -0.4860 | Yes |
| 50 | SGCE | SGCE Entrez, Source | sarcoglycan, epsilon | 12368 | -0.155 | -0.4690 | Yes |
| 51 | SMTN | SMTN Entrez, Source | smoothelin | 12423 | -0.161 | -0.4523 | Yes |
| 52 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 12628 | -0.189 | -0.4434 | Yes |
| 53 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 12819 | -0.229 | -0.4283 | Yes |
| 54 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 12873 | -0.243 | -0.4011 | Yes |
| 55 | SGCG | SGCG Entrez, Source | sarcoglycan, gamma (35kDa dystrophin-associated glycoprotein) | 13075 | -0.336 | -0.3730 | Yes |
| 56 | TAGLN | TAGLN Entrez, Source | transgelin | 13339 | -1.475 | -0.2035 | Yes |
| 57 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 13340 | -1.586 | 0.0002 | Yes |