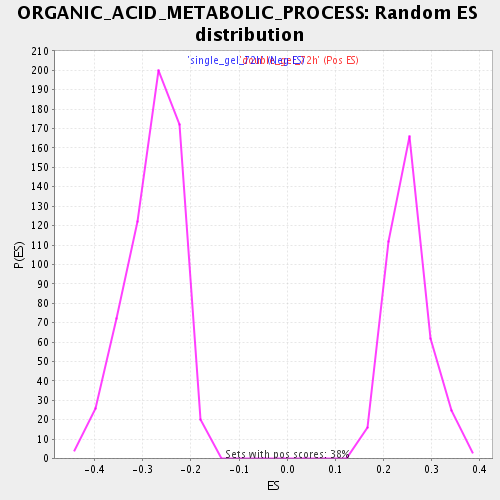
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | ORGANIC_ACID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.63714945 |
| Normalized Enrichment Score (NES) | 2.5360432 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 9 | 0.779 | 0.0355 | Yes |
| 2 | BBOX1 | BBOX1 Entrez, Source | butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 | 13 | 0.740 | 0.0696 | Yes |
| 3 | FTCD | FTCD Entrez, Source | formiminotransferase cyclodeaminase | 15 | 0.731 | 0.1035 | Yes |
| 4 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 33 | 0.554 | 0.1280 | Yes |
| 5 | HACL1 | HACL1 Entrez, Source | 2-hydroxyacyl-CoA lyase 1 | 54 | 0.488 | 0.1492 | Yes |
| 6 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 58 | 0.484 | 0.1714 | Yes |
| 7 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 95 | 0.419 | 0.1882 | Yes |
| 8 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 110 | 0.403 | 0.2058 | Yes |
| 9 | HGD | HGD Entrez, Source | homogentisate 1,2-dioxygenase (homogentisate oxidase) | 126 | 0.385 | 0.2226 | Yes |
| 10 | AGXT | AGXT Entrez, Source | alanine-glyoxylate aminotransferase (oxalosis I; hyperoxaluria I; glycolicaciduria; serine-pyruvate aminotransferase) | 128 | 0.384 | 0.2403 | Yes |
| 11 | GCK | GCK Entrez, Source | glucokinase (hexokinase 4, maturity onset diabetes of the young 2) | 140 | 0.368 | 0.2566 | Yes |
| 12 | ARG1 | ARG1 Entrez, Source | arginase, liver | 145 | 0.358 | 0.2729 | Yes |
| 13 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 182 | 0.324 | 0.2852 | Yes |
| 14 | ASL | ASL Entrez, Source | argininosuccinate lyase | 183 | 0.323 | 0.3003 | Yes |
| 15 | ACLY | ACLY Entrez, Source | ATP citrate lyase | 212 | 0.305 | 0.3123 | Yes |
| 16 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 219 | 0.299 | 0.3258 | Yes |
| 17 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.3386 | Yes |
| 18 | AS3MT | AS3MT Entrez, Source | arsenic (+3 oxidation state) methyltransferase | 242 | 0.289 | 0.3512 | Yes |
| 19 | NR1H4 | NR1H4 Entrez, Source | nuclear receptor subfamily 1, group H, member 4 | 246 | 0.288 | 0.3644 | Yes |
| 20 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 284 | 0.273 | 0.3742 | Yes |
| 21 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 295 | 0.267 | 0.3859 | Yes |
| 22 | MAT1A | MAT1A Entrez, Source | methionine adenosyltransferase I, alpha | 315 | 0.262 | 0.3966 | Yes |
| 23 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 320 | 0.260 | 0.4084 | Yes |
| 24 | GLS2 | GLS2 Entrez, Source | glutaminase 2 (liver, mitochondrial) | 341 | 0.252 | 0.4186 | Yes |
| 25 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 378 | 0.240 | 0.4270 | Yes |
| 26 | TST | TST Entrez, Source | thiosulfate sulfurtransferase (rhodanese) | 388 | 0.235 | 0.4372 | Yes |
| 27 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 428 | 0.222 | 0.4446 | Yes |
| 28 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 444 | 0.218 | 0.4536 | Yes |
| 29 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 445 | 0.217 | 0.4637 | Yes |
| 30 | SCLY | SCLY Entrez, Source | selenocysteine lyase | 454 | 0.214 | 0.4730 | Yes |
| 31 | ALDH1L1 | ALDH1L1 Entrez, Source | aldehyde dehydrogenase 1 family, member L1 | 465 | 0.211 | 0.4821 | Yes |
| 32 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 482 | 0.207 | 0.4905 | Yes |
| 33 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 536 | 0.195 | 0.4955 | Yes |
| 34 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 553 | 0.192 | 0.5032 | Yes |
| 35 | SLC25A15 | SLC25A15 Entrez, Source | solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15 | 595 | 0.186 | 0.5088 | Yes |
| 36 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 597 | 0.186 | 0.5174 | Yes |
| 37 | GNE | GNE Entrez, Source | glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase | 599 | 0.186 | 0.5259 | Yes |
| 38 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 612 | 0.184 | 0.5336 | Yes |
| 39 | ACOT12 | ACOT12 Entrez, Source | acyl-CoA thioesterase 12 | 619 | 0.184 | 0.5417 | Yes |
| 40 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 628 | 0.182 | 0.5496 | Yes |
| 41 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 633 | 0.182 | 0.5577 | Yes |
| 42 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 651 | 0.179 | 0.5647 | Yes |
| 43 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 662 | 0.178 | 0.5723 | Yes |
| 44 | SLC7A2 | SLC7A2 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 | 776 | 0.165 | 0.5713 | Yes |
| 45 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.5772 | Yes |
| 46 | GCSH | GCSH Entrez, Source | glycine cleavage system protein H (aminomethyl carrier) | 816 | 0.161 | 0.5835 | Yes |
| 47 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 823 | 0.160 | 0.5905 | Yes |
| 48 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 868 | 0.156 | 0.5944 | Yes |
| 49 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 933 | 0.150 | 0.5965 | Yes |
| 50 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.6022 | Yes |
| 51 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 984 | 0.146 | 0.6065 | Yes |
| 52 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 1046 | 0.140 | 0.6084 | Yes |
| 53 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 1151 | 0.133 | 0.6068 | Yes |
| 54 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 1252 | 0.127 | 0.6051 | Yes |
| 55 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 1387 | 0.119 | 0.6005 | Yes |
| 56 | AMT | AMT Entrez, Source | aminomethyltransferase | 1397 | 0.119 | 0.6053 | Yes |
| 57 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 1471 | 0.115 | 0.6051 | Yes |
| 58 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1480 | 0.114 | 0.6098 | Yes |
| 59 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1494 | 0.114 | 0.6141 | Yes |
| 60 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1495 | 0.114 | 0.6194 | Yes |
| 61 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 1524 | 0.112 | 0.6225 | Yes |
| 62 | HDC | HDC Entrez, Source | histidine decarboxylase | 1527 | 0.112 | 0.6275 | Yes |
| 63 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 1560 | 0.110 | 0.6302 | Yes |
| 64 | ASRGL1 | ASRGL1 Entrez, Source | asparaginase like 1 | 1612 | 0.108 | 0.6314 | Yes |
| 65 | NANP | NANP Entrez, Source | N-acetylneuraminic acid phosphatase | 1648 | 0.107 | 0.6337 | Yes |
| 66 | MLYCD | MLYCD Entrez, Source | malonyl-CoA decarboxylase | 1678 | 0.106 | 0.6364 | Yes |
| 67 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 1733 | 0.104 | 0.6371 | Yes |
| 68 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 1840 | 0.100 | 0.6338 | No |
| 69 | AKR1D1 | AKR1D1 Entrez, Source | aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase) | 1859 | 0.099 | 0.6370 | No |
| 70 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 1926 | 0.096 | 0.6365 | No |
| 71 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 2100 | 0.091 | 0.6276 | No |
| 72 | ADI1 | ADI1 Entrez, Source | acireductone dioxygenase 1 | 2159 | 0.089 | 0.6274 | No |
| 73 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 2259 | 0.086 | 0.6239 | No |
| 74 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 2407 | 0.082 | 0.6166 | No |
| 75 | CPT1B | CPT1B Entrez, Source | carnitine palmitoyltransferase 1B (muscle) | 2496 | 0.080 | 0.6136 | No |
| 76 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | 0.6095 | No |
| 77 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 2648 | 0.075 | 0.6093 | No |
| 78 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 3046 | 0.066 | 0.5823 | No |
| 79 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.5845 | No |
| 80 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 3063 | 0.065 | 0.5872 | No |
| 81 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 3136 | 0.064 | 0.5847 | No |
| 82 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 3398 | 0.058 | 0.5676 | No |
| 83 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 3517 | 0.056 | 0.5613 | No |
| 84 | ACOT2 | ACOT2 Entrez, Source | acyl-CoA thioesterase 2 | 3642 | 0.053 | 0.5544 | No |
| 85 | GGTLA1 | GGTLA1 Entrez, Source | gamma-glutamyltransferase-like activity 1 | 3715 | 0.052 | 0.5513 | No |
| 86 | MPST | MPST Entrez, Source | mercaptopyruvate sulfurtransferase | 3883 | 0.049 | 0.5409 | No |
| 87 | GSS | GSS Entrez, Source | glutathione synthetase | 4011 | 0.046 | 0.5335 | No |
| 88 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 4028 | 0.046 | 0.5344 | No |
| 89 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 4162 | 0.044 | 0.5263 | No |
| 90 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 4275 | 0.042 | 0.5198 | No |
| 91 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.5200 | No |
| 92 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 4326 | 0.041 | 0.5199 | No |
| 93 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 4383 | 0.040 | 0.5175 | No |
| 94 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 4470 | 0.038 | 0.5128 | No |
| 95 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.4989 | No |
| 96 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.4908 | No |
| 97 | PTGS1 | PTGS1 Entrez, Source | prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) | 4887 | 0.031 | 0.4860 | No |
| 98 | ARPP-19 | ARPP-19 Entrez, Source | - | 5020 | 0.029 | 0.4773 | No |
| 99 | GLUD1 | GLUD1 Entrez, Source | glutamate dehydrogenase 1 | 5087 | 0.028 | 0.4736 | No |
| 100 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 5602 | 0.020 | 0.4357 | No |
| 101 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.4170 | No |
| 102 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.4158 | No |
| 103 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 5916 | 0.015 | 0.4143 | No |
| 104 | SCYE1 | SCYE1 Entrez, Source | small inducible cytokine subfamily E, member 1 (endothelial monocyte-activating) | 6040 | 0.014 | 0.4057 | No |
| 105 | DARS | DARS Entrez, Source | aspartyl-tRNA synthetase | 6478 | 0.007 | 0.3729 | No |
| 106 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 6489 | 0.007 | 0.3724 | No |
| 107 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 6738 | 0.003 | 0.3538 | No |
| 108 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 7266 | -0.004 | 0.3141 | No |
| 109 | GBA2 | GBA2 Entrez, Source | glucosidase, beta (bile acid) 2 | 7269 | -0.004 | 0.3141 | No |
| 110 | SLC3A1 | SLC3A1 Entrez, Source | 1 | 7374 | -0.006 | 0.3065 | No |
| 111 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.2572 | No |
| 112 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 8089 | -0.016 | 0.2540 | No |
| 113 | SLC7A8 | SLC7A8 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 8 | 8292 | -0.020 | 0.2396 | No |
| 114 | ADIPOR1 | ADIPOR1 Entrez, Source | adiponectin receptor 1 | 8460 | -0.022 | 0.2280 | No |
| 115 | SDS | SDS Entrez, Source | serine dehydratase | 8683 | -0.026 | 0.2124 | No |
| 116 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 9012 | -0.032 | 0.1891 | No |
| 117 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 9114 | -0.034 | 0.1831 | No |
| 118 | FARS2 | FARS2 Entrez, Source | phenylalanine-tRNA synthetase 2 (mitochondrial) | 9209 | -0.036 | 0.1776 | No |
| 119 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 9316 | -0.038 | 0.1714 | No |
| 120 | SLC7A9 | SLC7A9 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 | 9816 | -0.048 | 0.1358 | No |
| 121 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 10430 | -0.062 | 0.0923 | No |
| 122 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 10797 | -0.072 | 0.0680 | No |
| 123 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 10801 | -0.073 | 0.0711 | No |
| 124 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | 0.0359 | No |
| 125 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 11580 | -0.103 | 0.0214 | No |
| 126 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 11691 | -0.109 | 0.0181 | No |
| 127 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 12626 | -0.189 | -0.0439 | No |
| 128 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 12900 | -0.253 | -0.0528 | No |
| 129 | LYPLA3 | LYPLA3 Entrez, Source | lysophospholipase 3 (lysosomal phospholipase A2) | 12939 | -0.266 | -0.0433 | No |
| 130 | SLC6A6 | SLC6A6 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, taurine), member 6 | 13097 | -0.354 | -0.0387 | No |
| 131 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 13118 | -0.367 | -0.0232 | No |
| 132 | SMS | SMS Entrez, Source | spermine synthase | 13163 | -0.420 | -0.0070 | No |
| 133 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 13181 | -0.442 | 0.0122 | No |