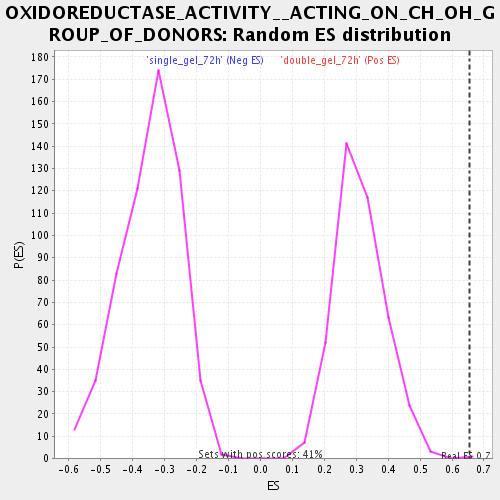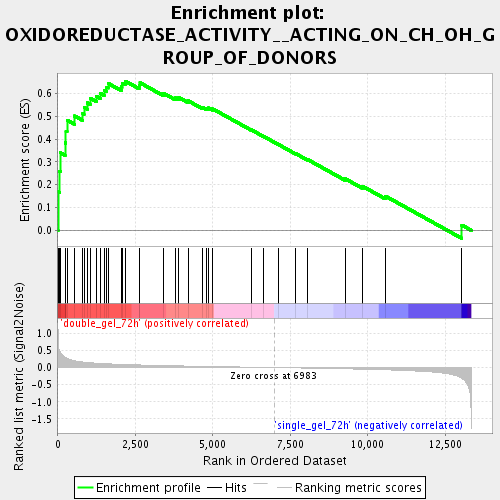
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | OXIDOREDUCTASE_ACTIVITY__ACTING_ON_CH_OH_GROUP_OF_DONORS |
| Enrichment Score (ES) | 0.65350693 |
| Normalized Enrichment Score (NES) | 2.0931115 |
| Nominal p-value | 0.0024509805 |
| FDR q-value | 6.183967E-4 |
| FWER p-Value | 0.024 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 6 | 0.908 | 0.1702 | Yes |
| 2 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 53 | 0.495 | 0.2597 | Yes |
| 3 | HSD17B7 | HSD17B7 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 7 | 78 | 0.438 | 0.3403 | Yes |
| 4 | MIOX | MIOX Entrez, Source | myo-inositol oxygenase | 234 | 0.291 | 0.3834 | Yes |
| 5 | AKR7A2 | AKR7A2 Entrez, Source | aldo-keto reductase family 7, member A2 (aflatoxin aldehyde reductase) | 260 | 0.283 | 0.4347 | Yes |
| 6 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 296 | 0.267 | 0.4823 | Yes |
| 7 | HAO2 | HAO2 Entrez, Source | hydroxyacid oxidase 2 (long chain) | 536 | 0.195 | 0.5010 | Yes |
| 8 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.5118 | Yes |
| 9 | AKR7A3 | AKR7A3 Entrez, Source | aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase) | 845 | 0.158 | 0.5382 | Yes |
| 10 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 951 | 0.149 | 0.5583 | Yes |
| 11 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 1044 | 0.141 | 0.5778 | Yes |
| 12 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 1246 | 0.127 | 0.5866 | Yes |
| 13 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 1370 | 0.121 | 0.6000 | Yes |
| 14 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1494 | 0.114 | 0.6121 | Yes |
| 15 | BDH1 | BDH1 Entrez, Source | 3-hydroxybutyrate dehydrogenase, type 1 | 1574 | 0.110 | 0.6268 | Yes |
| 16 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1622 | 0.108 | 0.6435 | Yes |
| 17 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 2053 | 0.092 | 0.6285 | Yes |
| 18 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 2069 | 0.092 | 0.6446 | Yes |
| 19 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2174 | 0.089 | 0.6535 | Yes |
| 20 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 2625 | 0.076 | 0.6339 | No |
| 21 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 2648 | 0.075 | 0.6463 | No |
| 22 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 3410 | 0.058 | 0.6000 | No |
| 23 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 3800 | 0.050 | 0.5801 | No |
| 24 | IMPDH2 | IMPDH2 Entrez, Source | IMP (inosine monophosphate) dehydrogenase 2 | 3898 | 0.048 | 0.5820 | No |
| 25 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 4199 | 0.043 | 0.5675 | No |
| 26 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 4675 | 0.035 | 0.5383 | No |
| 27 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 4803 | 0.032 | 0.5348 | No |
| 28 | DHRS9 | DHRS9 Entrez, Source | dehydrogenase/reductase (SDR family) member 9 | 4865 | 0.031 | 0.5361 | No |
| 29 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 4997 | 0.029 | 0.5318 | No |
| 30 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 6237 | 0.011 | 0.4406 | No |
| 31 | LDHC | LDHC Entrez, Source | lactate dehydrogenase C | 6640 | 0.005 | 0.4113 | No |
| 32 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 7127 | -0.002 | 0.3752 | No |
| 33 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 7676 | -0.010 | 0.3359 | No |
| 34 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 8061 | -0.016 | 0.3100 | No |
| 35 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 9268 | -0.037 | 0.2263 | No |
| 36 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 9824 | -0.048 | 0.1937 | No |
| 37 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 10569 | -0.066 | 0.1502 | No |
| 38 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 13034 | -0.309 | 0.0231 | No |