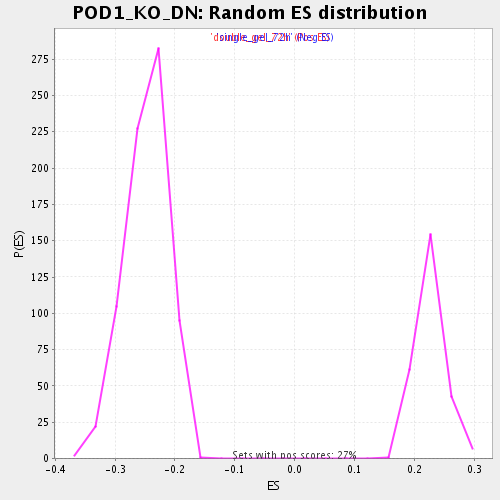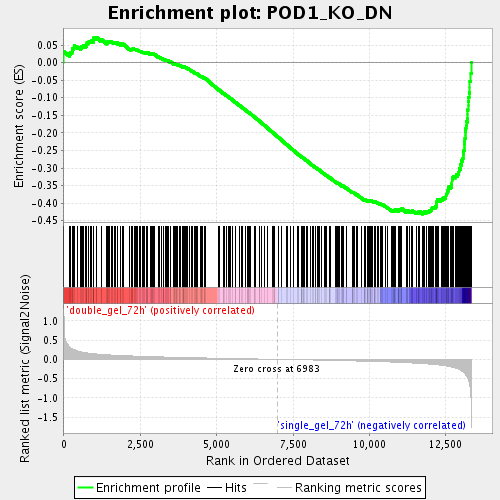
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | POD1_KO_DN |
| Enrichment Score (ES) | -0.43120918 |
| Normalized Enrichment Score (NES) | -1.7439097 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.048659828 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CLEC14A | CLEC14A Entrez, Source | C-type lectin domain family 14, member A | 1 | 1.262 | 0.0315 | No |
| 2 | SLCO2A1 | SLCO2A1 Entrez, Source | solute carrier organic anion transporter family, member 2A1 | 181 | 0.327 | 0.0259 | No |
| 3 | SYCP3 | SYCP3 Entrez, Source | synaptonemal complex protein 3 | 225 | 0.296 | 0.0300 | No |
| 4 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 266 | 0.281 | 0.0340 | No |
| 5 | MAFB | MAFB Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian) | 285 | 0.273 | 0.0394 | No |
| 6 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 329 | 0.257 | 0.0425 | No |
| 7 | LIFR | LIFR Entrez, Source | leukemia inhibitory factor receptor alpha | 339 | 0.253 | 0.0482 | No |
| 8 | CTSS | CTSS Entrez, Source | cathepsin S | 456 | 0.214 | 0.0446 | No |
| 9 | CD97 | CD97 Entrez, Source | CD97 molecule | 554 | 0.192 | 0.0419 | No |
| 10 | RAPGEF5 | RAPGEF5 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 5 | 575 | 0.188 | 0.0451 | No |
| 11 | ST8SIA4 | ST8SIA4 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 | 611 | 0.184 | 0.0470 | No |
| 12 | MERTK | MERTK Entrez, Source | c-mer proto-oncogene tyrosine kinase | 647 | 0.180 | 0.0488 | No |
| 13 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 717 | 0.171 | 0.0478 | No |
| 14 | SH2B3 | SH2B3 Entrez, Source | SH2B adaptor protein 3 | 751 | 0.167 | 0.0495 | No |
| 15 | NT5E | NT5E Entrez, Source | 5'-nucleotidase, ecto (CD73) | 752 | 0.167 | 0.0536 | No |
| 16 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 754 | 0.167 | 0.0577 | No |
| 17 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 817 | 0.161 | 0.0570 | No |
| 18 | HAGH | HAGH Entrez, Source | hydroxyacylglutathione hydrolase | 819 | 0.161 | 0.0609 | No |
| 19 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 884 | 0.154 | 0.0599 | No |
| 20 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 901 | 0.153 | 0.0624 | No |
| 21 | SH3BP5 | SH3BP5 Entrez, Source | SH3-domain binding protein 5 (BTK-associated) | 959 | 0.148 | 0.0618 | No |
| 22 | LYZ | LYZ Entrez, Source | lysozyme (renal amyloidosis) | 961 | 0.148 | 0.0654 | No |
| 23 | PRKAA1 | PRKAA1 Entrez, Source | protein kinase, AMP-activated, alpha 1 catalytic subunit | 967 | 0.148 | 0.0687 | No |
| 24 | SLC33A1 | SLC33A1 Entrez, Source | solute carrier family 33 (acetyl-CoA transporter), member 1 | 981 | 0.147 | 0.0714 | No |
| 25 | THBD | THBD Entrez, Source | thrombomodulin | 1055 | 0.140 | 0.0693 | No |
| 26 | GPR116 | GPR116 Entrez, Source | G protein-coupled receptor 116 | 1063 | 0.139 | 0.0722 | No |
| 27 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 1216 | 0.129 | 0.0637 | No |
| 28 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 1245 | 0.127 | 0.0648 | No |
| 29 | SGPP1 | SGPP1 Entrez, Source | sphingosine-1-phosphate phosphatase 1 | 1402 | 0.119 | 0.0557 | No |
| 30 | SCN7A | SCN7A Entrez, Source | sodium channel, voltage-gated, type VII, alpha | 1423 | 0.117 | 0.0571 | No |
| 31 | SEC11L3 | SEC11L3 Entrez, Source | SEC11-like 3 (S. cerevisiae) | 1436 | 0.116 | 0.0591 | No |
| 32 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1449 | 0.116 | 0.0611 | No |
| 33 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 1483 | 0.114 | 0.0614 | No |
| 34 | TOP1MT | TOP1MT Entrez, Source | topoisomerase (DNA) I, mitochondrial | 1545 | 0.111 | 0.0595 | No |
| 35 | NOSTRIN | NOSTRIN Entrez, Source | nitric oxide synthase trafficker | 1595 | 0.109 | 0.0584 | No |
| 36 | MARK1 | MARK1 Entrez, Source | MAP/microtubule affinity-regulating kinase 1 | 1653 | 0.107 | 0.0567 | No |
| 37 | GABRA4 | GABRA4 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 4 | 1698 | 0.105 | 0.0560 | No |
| 38 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 1747 | 0.103 | 0.0548 | No |
| 39 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 1748 | 0.103 | 0.0574 | No |
| 40 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 1862 | 0.099 | 0.0512 | No |
| 41 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 1863 | 0.099 | 0.0537 | No |
| 42 | EMCN | EMCN Entrez, Source | endomucin | 1920 | 0.097 | 0.0518 | No |
| 43 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 1935 | 0.096 | 0.0531 | No |
| 44 | TCP11L2 | TCP11L2 Entrez, Source | t-complex 11 (mouse) like 2 | 2152 | 0.090 | 0.0387 | No |
| 45 | LPHN3 | LPHN3 Entrez, Source | latrophilin 3 | 2198 | 0.088 | 0.0375 | No |
| 46 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 2217 | 0.088 | 0.0383 | No |
| 47 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 2241 | 0.087 | 0.0387 | No |
| 48 | FOXC2 | FOXC2 Entrez, Source | forkhead box C2 (MFH-1, mesenchyme forkhead 1) | 2258 | 0.086 | 0.0396 | No |
| 49 | WWP1 | WWP1 Entrez, Source | WW domain containing E3 ubiquitin protein ligase 1 | 2306 | 0.085 | 0.0381 | No |
| 50 | IRS1 | IRS1 Entrez, Source | insulin receptor substrate 1 | 2329 | 0.084 | 0.0385 | No |
| 51 | SLK | SLK Entrez, Source | STE20-like kinase (yeast) | 2387 | 0.083 | 0.0362 | No |
| 52 | PTPRO | PTPRO Entrez, Source | protein tyrosine phosphatase, receptor type, O | 2417 | 0.082 | 0.0360 | No |
| 53 | SLC9A3R2 | SLC9A3R2 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 2 | 2485 | 0.080 | 0.0329 | No |
| 54 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 2513 | 0.079 | 0.0328 | No |
| 55 | OPTN | OPTN Entrez, Source | optineurin | 2577 | 0.077 | 0.0299 | No |
| 56 | DPP4 | DPP4 Entrez, Source | dipeptidyl-peptidase 4 (CD26, adenosine deaminase complexing protein 2) | 2611 | 0.076 | 0.0293 | No |
| 57 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 2628 | 0.076 | 0.0299 | No |
| 58 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 2693 | 0.074 | 0.0268 | No |
| 59 | HOMER1 | HOMER1 Entrez, Source | homer homolog 1 (Drosophila) | 2707 | 0.073 | 0.0277 | No |
| 60 | GLRX2 | GLRX2 Entrez, Source | glutaredoxin 2 | 2722 | 0.073 | 0.0284 | No |
| 61 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 2743 | 0.073 | 0.0287 | No |
| 62 | TMEM106B | TMEM106B Entrez, Source | transmembrane protein 106B | 2828 | 0.071 | 0.0240 | No |
| 63 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 2854 | 0.070 | 0.0238 | No |
| 64 | RAPGEF3 | RAPGEF3 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 3 | 2860 | 0.070 | 0.0252 | No |
| 65 | ODZ2 | ODZ2 Entrez, Source | odz, odd Oz/ten-m homolog 2 (Drosophila) | 2889 | 0.069 | 0.0248 | No |
| 66 | ANKRD15 | ANKRD15 Entrez, Source | ankyrin repeat domain 15 | 2894 | 0.069 | 0.0262 | No |
| 67 | AGTR1 | AGTR1 Entrez, Source | angiotensin II receptor, type 1 | 2934 | 0.068 | 0.0249 | No |
| 68 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 2974 | 0.067 | 0.0236 | No |
| 69 | PTPN12 | PTPN12 Entrez, Source | protein tyrosine phosphatase, non-receptor type 12 | 3086 | 0.065 | 0.0167 | No |
| 70 | SACM1L | SACM1L Entrez, Source | SAC1 suppressor of actin mutations 1-like (yeast) | 3120 | 0.064 | 0.0157 | No |
| 71 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 3178 | 0.063 | 0.0129 | No |
| 72 | PDLIM2 | PDLIM2 Entrez, Source | PDZ and LIM domain 2 (mystique) | 3251 | 0.061 | 0.0089 | No |
| 73 | TEX2 | TEX2 Entrez, Source | testis expressed sequence 2 | 3256 | 0.061 | 0.0101 | No |
| 74 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 3274 | 0.061 | 0.0103 | No |
| 75 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 3337 | 0.059 | 0.0070 | No |
| 76 | ARNT | ARNT Entrez, Source | aryl hydrocarbon receptor nuclear translocator | 3368 | 0.059 | 0.0062 | No |
| 77 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 3374 | 0.059 | 0.0073 | No |
| 78 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 3416 | 0.058 | 0.0056 | No |
| 79 | EHD3 | EHD3 Entrez, Source | EH-domain containing 3 | 3481 | 0.056 | 0.0021 | No |
| 80 | CALCRL | CALCRL Entrez, Source | calcitonin receptor-like | 3492 | 0.056 | 0.0027 | No |
| 81 | ARAF | ARAF Entrez, Source | v-raf murine sarcoma 3611 viral oncogene homolog | 3591 | 0.054 | -0.0035 | No |
| 82 | UBE2V2 | UBE2V2 Entrez, Source | ubiquitin-conjugating enzyme E2 variant 2 | 3602 | 0.054 | -0.0029 | No |
| 83 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 3612 | 0.053 | -0.0023 | No |
| 84 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 3665 | 0.053 | -0.0050 | No |
| 85 | IL13RA1 | IL13RA1 Entrez, Source | interleukin 13 receptor, alpha 1 | 3674 | 0.052 | -0.0043 | No |
| 86 | NOTCH4 | NOTCH4 Entrez, Source | Notch homolog 4 (Drosophila) | 3695 | 0.052 | -0.0045 | No |
| 87 | IL6ST | IL6ST Entrez, Source | interleukin 6 signal transducer (gp130, oncostatin M receptor) | 3732 | 0.051 | -0.0060 | No |
| 88 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 3767 | 0.050 | -0.0073 | No |
| 89 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 3792 | 0.050 | -0.0079 | No |
| 90 | EBAG9 | EBAG9 Entrez, Source | estrogen receptor binding site associated, antigen, 9 | 3819 | 0.049 | -0.0087 | No |
| 91 | ACBD5 | ACBD5 Entrez, Source | acyl-Coenzyme A binding domain containing 5 | 3889 | 0.049 | -0.0128 | No |
| 92 | MTM1 | MTM1 Entrez, Source | myotubularin 1 | 3901 | 0.048 | -0.0124 | No |
| 93 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 3905 | 0.048 | -0.0115 | No |
| 94 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 3912 | 0.048 | -0.0107 | No |
| 95 | ATP10A | ATP10A Entrez, Source | ATPase, Class V, type 10A | 3936 | 0.048 | -0.0113 | No |
| 96 | CXCR7 | CXCR7 Entrez, Source | chemokine (C-X-C motif) receptor 7 | 3981 | 0.047 | -0.0135 | No |
| 97 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 3999 | 0.046 | -0.0136 | No |
| 98 | CTBS | CTBS Entrez, Source | chitobiase, di-N-acetyl- | 4059 | 0.045 | -0.0171 | No |
| 99 | SLC38A4 | SLC38A4 Entrez, Source | solute carrier family 38, member 4 | 4109 | 0.044 | -0.0197 | No |
| 100 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 4177 | 0.043 | -0.0238 | No |
| 101 | ARRB1 | ARRB1 Entrez, Source | arrestin, beta 1 | 4212 | 0.043 | -0.0253 | No |
| 102 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 4260 | 0.042 | -0.0279 | No |
| 103 | TLR4 | TLR4 Entrez, Source | toll-like receptor 4 | 4308 | 0.041 | -0.0305 | No |
| 104 | GJA3 | GJA3 Entrez, Source | gap junction protein, alpha 3, 46kDa (connexin 46) | 4332 | 0.041 | -0.0312 | No |
| 105 | PDE4DIP | PDE4DIP Entrez, Source | phosphodiesterase 4D interacting protein (myomegalin) | 4349 | 0.040 | -0.0315 | No |
| 106 | COL4A4 | COL4A4 Entrez, Source | collagen, type IV, alpha 4 | 4358 | 0.040 | -0.0311 | No |
| 107 | LYSMD3 | LYSMD3 Entrez, Source | LysM, putative peptidoglycan-binding, domain containing 3 | 4484 | 0.038 | -0.0397 | No |
| 108 | VPS54 | VPS54 Entrez, Source | vacuolar protein sorting 54 (S. cerevisiae) | 4489 | 0.038 | -0.0391 | No |
| 109 | PRKCH | PRKCH Entrez, Source | protein kinase C, eta | 4516 | 0.038 | -0.0402 | No |
| 110 | PFKFB3 | PFKFB3 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 | 4523 | 0.037 | -0.0397 | No |
| 111 | GPSM3 | GPSM3 Entrez, Source | G-protein signalling modulator 3 (AGS3-like, C. elegans) | 4585 | 0.036 | -0.0435 | No |
| 112 | BACE1 | BACE1 Entrez, Source | beta-site APP-cleaving enzyme 1 | 4611 | 0.036 | -0.0445 | No |
| 113 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 4640 | 0.035 | -0.0458 | No |
| 114 | SLC17A5 | SLC17A5 Entrez, Source | solute carrier family 17 (anion/sugar transporter), member 5 | 5072 | 0.028 | -0.0782 | No |
| 115 | ZFR | ZFR Entrez, Source | zinc finger RNA binding protein | 5075 | 0.028 | -0.0777 | No |
| 116 | EBF1 | EBF1 Entrez, Source | early B-cell factor 1 | 5224 | 0.026 | -0.0884 | No |
| 117 | INPP4A | INPP4A Entrez, Source | inositol polyphosphate-4-phosphatase, type I, 107kDa | 5225 | 0.026 | -0.0878 | No |
| 118 | IL2RG | IL2RG Entrez, Source | interleukin 2 receptor, gamma (severe combined immunodeficiency) | 5243 | 0.025 | -0.0884 | No |
| 119 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 5312 | 0.024 | -0.0931 | No |
| 120 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 5375 | 0.024 | -0.0972 | No |
| 121 | TMEM30A | TMEM30A Entrez, Source | transmembrane protein 30A | 5421 | 0.023 | -0.1001 | No |
| 122 | EFNB1 | EFNB1 Entrez, Source | ephrin-B1 | 5440 | 0.023 | -0.1009 | No |
| 123 | PGGT1B | PGGT1B Entrez, Source | protein geranylgeranyltransferase type I, beta subunit | 5519 | 0.021 | -0.1064 | No |
| 124 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 5608 | 0.020 | -0.1127 | No |
| 125 | UGP2 | UGP2 Entrez, Source | UDP-glucose pyrophosphorylase 2 | 5626 | 0.020 | -0.1135 | No |
| 126 | SIRPA | SIRPA Entrez, Source | signal-regulatory protein alpha | 5733 | 0.018 | -0.1212 | No |
| 127 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 5745 | 0.018 | -0.1216 | No |
| 128 | SON | SON Entrez, Source | SON DNA binding protein | 5809 | 0.017 | -0.1260 | No |
| 129 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 5853 | 0.017 | -0.1289 | No |
| 130 | ART3 | ART3 Entrez, Source | ADP-ribosyltransferase 3 | 5951 | 0.015 | -0.1360 | No |
| 131 | SPOP | SPOP Entrez, Source | speckle-type POZ protein | 6006 | 0.014 | -0.1398 | No |
| 132 | CENTB2 | CENTB2 Entrez, Source | centaurin, beta 2 | 6012 | 0.014 | -0.1398 | No |
| 133 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 6036 | 0.014 | -0.1412 | No |
| 134 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 6039 | 0.014 | -0.1411 | No |
| 135 | CTTNBP2 | CTTNBP2 Entrez, Source | cortactin binding protein 2 | 6087 | 0.013 | -0.1443 | No |
| 136 | RSN | RSN Entrez, Source | restin (Reed-Steinberg cell-expressed intermediate filament-associated protein) | 6102 | 0.013 | -0.1451 | No |
| 137 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 6237 | 0.011 | -0.1551 | No |
| 138 | PROS1 | PROS1 Entrez, Source | protein S (alpha) | 6244 | 0.011 | -0.1553 | No |
| 139 | PARVA | PARVA Entrez, Source | parvin, alpha | 6285 | 0.010 | -0.1582 | No |
| 140 | RBBP6 | RBBP6 Entrez, Source | retinoblastoma binding protein 6 | 6400 | 0.008 | -0.1667 | No |
| 141 | TSPAN5 | TSPAN5 Entrez, Source | tetraspanin 5 | 6410 | 0.008 | -0.1672 | No |
| 142 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 6476 | 0.007 | -0.1721 | No |
| 143 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6567 | 0.006 | -0.1789 | No |
| 144 | METRNL | METRNL Entrez, Source | meteorin, glial cell differentiation regulator-like | 6664 | 0.004 | -0.1861 | No |
| 145 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 6814 | 0.002 | -0.1975 | No |
| 146 | FUT11 | FUT11 Entrez, Source | fucosyltransferase 11 (alpha (1,3) fucosyltransferase) | 6854 | 0.002 | -0.2005 | No |
| 147 | RFK | RFK Entrez, Source | riboflavin kinase | 6901 | 0.001 | -0.2040 | No |
| 148 | PHLPP | PHLPP Entrez, Source | PH domain and leucine rich repeat protein phosphatase | 6904 | 0.001 | -0.2041 | No |
| 149 | EPN2 | EPN2 Entrez, Source | epsin 2 | 7010 | -0.000 | -0.2122 | No |
| 150 | GIMAP6 | GIMAP6 Entrez, Source | GTPase, IMAP family member 6 | 7025 | -0.001 | -0.2132 | No |
| 151 | SNRK | SNRK Entrez, Source | SNF related kinase | 7109 | -0.002 | -0.2196 | No |
| 152 | USP33 | USP33 Entrez, Source | ubiquitin specific peptidase 33 | 7270 | -0.004 | -0.2318 | No |
| 153 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 7306 | -0.005 | -0.2344 | No |
| 154 | FLT1 | FLT1 Entrez, Source | fms-related tyrosine kinase 1 (vascular endothelial growth factor/vascular permeability factor receptor) | 7411 | -0.006 | -0.2422 | No |
| 155 | GIMAP1 | GIMAP1 Entrez, Source | GTPase, IMAP family member 1 | 7520 | -0.008 | -0.2503 | No |
| 156 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 7659 | -0.010 | -0.2607 | No |
| 157 | TAOK1 | TAOK1 Entrez, Source | TAO kinase 1 | 7692 | -0.010 | -0.2629 | No |
| 158 | NID1 | NID1 Entrez, Source | nidogen 1 | 7761 | -0.011 | -0.2678 | No |
| 159 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 7785 | -0.012 | -0.2693 | No |
| 160 | SERINC1 | SERINC1 Entrez, Source | serine incorporator 1 | 7788 | -0.012 | -0.2692 | No |
| 161 | PLD1 | PLD1 Entrez, Source | phospholipase D1, phosphatidylcholine-specific | 7809 | -0.012 | -0.2704 | No |
| 162 | PTPRV | PTPRV Entrez, Source | protein tyrosine phosphatase, receptor type, V (pseudogene) | 7826 | -0.013 | -0.2713 | No |
| 163 | DDN | DDN Entrez, Source | dendrin | 7856 | -0.013 | -0.2732 | No |
| 164 | LRRC33 | LRRC33 Entrez, Source | leucine rich repeat containing 33 | 7861 | -0.013 | -0.2732 | No |
| 165 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 7925 | -0.014 | -0.2777 | No |
| 166 | MPP3 | MPP3 Entrez, Source | membrane protein, palmitoylated 3 (MAGUK p55 subfamily member 3) | 7959 | -0.014 | -0.2799 | No |
| 167 | ATP7A | ATP7A Entrez, Source | ATPase, Cu++ transporting, alpha polypeptide (Menkes syndrome) | 8085 | -0.016 | -0.2891 | No |
| 168 | MYO1B | MYO1B Entrez, Source | myosin IB | 8141 | -0.017 | -0.2929 | No |
| 169 | SYBL1 | SYBL1 Entrez, Source | synaptobrevin-like 1 | 8181 | -0.018 | -0.2955 | No |
| 170 | PPAP2A | PPAP2A Entrez, Source | phosphatidic acid phosphatase type 2A | 8226 | -0.018 | -0.2984 | No |
| 171 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 8231 | -0.019 | -0.2982 | No |
| 172 | SH2D4A | SH2D4A Entrez, Source | SH2 domain containing 4A | 8283 | -0.020 | -0.3017 | No |
| 173 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 8308 | -0.020 | -0.3030 | No |
| 174 | SEMA3G | SEMA3G Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3G | 8324 | -0.020 | -0.3037 | No |
| 175 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 8356 | -0.021 | -0.3055 | No |
| 176 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 8440 | -0.022 | -0.3114 | No |
| 177 | ABI3 | ABI3 Entrez, Source | ABI gene family, member 3 | 8535 | -0.024 | -0.3180 | No |
| 178 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 8561 | -0.024 | -0.3193 | No |
| 179 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 8589 | -0.025 | -0.3208 | No |
| 180 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 8676 | -0.026 | -0.3267 | No |
| 181 | TMEM33 | TMEM33 Entrez, Source | transmembrane protein 33 | 8702 | -0.026 | -0.3280 | No |
| 182 | CSNK1G3 | CSNK1G3 Entrez, Source | casein kinase 1, gamma 3 | 8705 | -0.026 | -0.3275 | No |
| 183 | WDFY1 | WDFY1 Entrez, Source | WD repeat and FYVE domain containing 1 | 8732 | -0.027 | -0.3288 | No |
| 184 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 8901 | -0.030 | -0.3410 | No |
| 185 | TSPAN12 | TSPAN12 Entrez, Source | tetraspanin 12 | 8909 | -0.030 | -0.3408 | No |
| 186 | GRM7 | GRM7 Entrez, Source | glutamate receptor, metabotropic 7 | 8939 | -0.031 | -0.3422 | No |
| 187 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 8958 | -0.031 | -0.3428 | No |
| 188 | LMBRD1 | LMBRD1 Entrez, Source | LMBR1 domain containing 1 | 8992 | -0.032 | -0.3446 | No |
| 189 | GALNT10 | GALNT10 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 10 (GalNAc-T10) | 9008 | -0.032 | -0.3449 | No |
| 190 | CRSP3 | CRSP3 Entrez, Source | cofactor required for Sp1 transcriptional activation, subunit 3, 130kDa | 9098 | -0.034 | -0.3509 | No |
| 191 | ZHX1 | ZHX1 Entrez, Source | zinc fingers and homeoboxes 1 | 9119 | -0.034 | -0.3516 | No |
| 192 | PARP4 | PARP4 Entrez, Source | poly (ADP-ribose) polymerase family, member 4 | 9126 | -0.035 | -0.3512 | No |
| 193 | POLB | POLB Entrez, Source | polymerase (DNA directed), beta | 9135 | -0.035 | -0.3509 | No |
| 194 | CLCA2 | CLCA2 Entrez, Source | chloride channel, calcium activated, family member 2 | 9159 | -0.035 | -0.3518 | No |
| 195 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 9244 | -0.037 | -0.3574 | No |
| 196 | HIP1 | HIP1 Entrez, Source | huntingtin interacting protein 1 | 9248 | -0.037 | -0.3567 | No |
| 197 | CRK | CRK Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian) | 9430 | -0.040 | -0.3696 | No |
| 198 | FABP4 | FABP4 Entrez, Source | fatty acid binding protein 4, adipocyte | 9450 | -0.041 | -0.3700 | No |
| 199 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 9454 | -0.041 | -0.3693 | No |
| 200 | YPEL5 | YPEL5 Entrez, Source | yippee-like 5 (Drosophila) | 9461 | -0.041 | -0.3687 | No |
| 201 | MORF4L1 | MORF4L1 Entrez, Source | mortality factor 4 like 1 | 9516 | -0.042 | -0.3718 | No |
| 202 | CDC73 | CDC73 Entrez, Source | cell division cycle 73, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 9588 | -0.043 | -0.3762 | No |
| 203 | YIPF4 | YIPF4 Entrez, Source | Yip1 domain family, member 4 | 9608 | -0.044 | -0.3765 | No |
| 204 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 9737 | -0.046 | -0.3852 | No |
| 205 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 9831 | -0.048 | -0.3912 | No |
| 206 | GMDS | GMDS Entrez, Source | GDP-mannose 4,6-dehydratase | 9847 | -0.049 | -0.3911 | No |
| 207 | ELTD1 | ELTD1 Entrez, Source | EGF, latrophilin and seven transmembrane domain containing 1 | 9849 | -0.049 | -0.3900 | No |
| 208 | KLHL5 | KLHL5 Entrez, Source | kelch-like 5 (Drosophila) | 9873 | -0.049 | -0.3905 | No |
| 209 | SYNPO | SYNPO Entrez, Source | synaptopodin | 9931 | -0.051 | -0.3936 | No |
| 210 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 9940 | -0.051 | -0.3930 | No |
| 211 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 9946 | -0.051 | -0.3921 | No |
| 212 | MAGI2 | MAGI2 Entrez, Source | membrane associated guanylate kinase, WW and PDZ domain containing 2 | 9967 | -0.051 | -0.3923 | No |
| 213 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 10008 | -0.052 | -0.3941 | No |
| 214 | DSCR1L1 | DSCR1L1 Entrez, Source | Down syndrome critical region gene 1-like 1 | 10021 | -0.053 | -0.3937 | No |
| 215 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 10034 | -0.053 | -0.3933 | No |
| 216 | LYPLA1 | LYPLA1 Entrez, Source | lysophospholipase I | 10038 | -0.053 | -0.3922 | No |
| 217 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 10068 | -0.053 | -0.3931 | No |
| 218 | TMBIM1 | TMBIM1 Entrez, Source | transmembrane BAX inhibitor motif containing 1 | 10109 | -0.055 | -0.3948 | No |
| 219 | UMOD | UMOD Entrez, Source | uromodulin (uromucoid, Tamm-Horsfall glycoprotein) | 10156 | -0.056 | -0.3970 | No |
| 220 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 10161 | -0.056 | -0.3959 | No |
| 221 | NCOR1 | NCOR1 Entrez, Source | nuclear receptor co-repressor 1 | 10165 | -0.056 | -0.3947 | No |
| 222 | SAMD8 | SAMD8 Entrez, Source | sterile alpha motif domain containing 8 | 10201 | -0.057 | -0.3960 | No |
| 223 | PLCL1 | PLCL1 Entrez, Source | phospholipase C-like 1 | 10259 | -0.058 | -0.3989 | No |
| 224 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10306 | -0.059 | -0.4010 | No |
| 225 | LEPREL1 | LEPREL1 Entrez, Source | leprecan-like 1 | 10355 | -0.060 | -0.4032 | No |
| 226 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 10358 | -0.060 | -0.4018 | No |
| 227 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 10405 | -0.062 | -0.4038 | No |
| 228 | PDE2A | PDE2A Entrez, Source | phosphodiesterase 2A, cGMP-stimulated | 10423 | -0.062 | -0.4036 | No |
| 229 | STXBP3 | STXBP3 Entrez, Source | syntaxin binding protein 3 | 10515 | -0.064 | -0.4089 | No |
| 230 | HIVEP2 | HIVEP2 Entrez, Source | human immunodeficiency virus type I enhancer binding protein 2 | 10603 | -0.067 | -0.4140 | No |
| 231 | COLEC12 | COLEC12 Entrez, Source | collectin sub-family member 12 | 10716 | -0.070 | -0.4208 | No |
| 232 | SULF2 | SULF2 Entrez, Source | sulfatase 2 | 10768 | -0.072 | -0.4230 | No |
| 233 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 10772 | -0.072 | -0.4214 | No |
| 234 | JAK1 | JAK1 Entrez, Source | Janus kinase 1 (a protein tyrosine kinase) | 10774 | -0.072 | -0.4197 | No |
| 235 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 10814 | -0.073 | -0.4209 | No |
| 236 | MAPT | MAPT Entrez, Source | microtubule-associated protein tau | 10830 | -0.073 | -0.4202 | No |
| 237 | PDE4B | PDE4B Entrez, Source | phosphodiesterase 4B, cAMP-specific (phosphodiesterase E4 dunce homolog, Drosophila) | 10834 | -0.073 | -0.4186 | No |
| 238 | LNPEP | LNPEP Entrez, Source | leucyl/cystinyl aminopeptidase | 10866 | -0.074 | -0.4191 | No |
| 239 | TMOD3 | TMOD3 Entrez, Source | tropomodulin 3 (ubiquitous) | 10944 | -0.077 | -0.4231 | No |
| 240 | SH3GLB1 | SH3GLB1 Entrez, Source | SH3-domain GRB2-like endophilin B1 | 10950 | -0.077 | -0.4216 | No |
| 241 | RIPK5 | RIPK5 Entrez, Source | receptor interacting protein kinase 5 | 10954 | -0.077 | -0.4199 | No |
| 242 | SOS2 | SOS2 Entrez, Source | son of sevenless homolog 2 (Drosophila) | 10962 | -0.077 | -0.4185 | No |
| 243 | SLC12A1 | SLC12A1 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 1 | 10968 | -0.078 | -0.4169 | No |
| 244 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 11002 | -0.079 | -0.4175 | No |
| 245 | CYP20A1 | CYP20A1 Entrez, Source | cytochrome P450, family 20, subfamily A, polypeptide 1 | 11047 | -0.080 | -0.4189 | No |
| 246 | TNFRSF11B | TNFRSF11B Entrez, Source | tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin) | 11055 | -0.080 | -0.4174 | No |
| 247 | LYST | LYST Entrez, Source | lysosomal trafficking regulator | 11059 | -0.080 | -0.4156 | No |
| 248 | CYP4B1 | CYP4B1 Entrez, Source | cytochrome P450, family 4, subfamily B, polypeptide 1 | 11195 | -0.086 | -0.4238 | No |
| 249 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 11218 | -0.087 | -0.4234 | No |
| 250 | ROCK1 | ROCK1 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 1 | 11231 | -0.087 | -0.4221 | No |
| 251 | CAST | CAST Entrez, Source | calpastatin | 11239 | -0.087 | -0.4205 | No |
| 252 | DEGS1 | DEGS1 Entrez, Source | degenerative spermatocyte homolog 1, lipid desaturase (Drosophila) | 11324 | -0.092 | -0.4246 | No |
| 253 | WDR22 | WDR22 Entrez, Source | WD repeat domain 22 | 11366 | -0.093 | -0.4255 | No |
| 254 | SPEG | SPEG Entrez, Source | SPEG complex locus | 11368 | -0.093 | -0.4232 | No |
| 255 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 11394 | -0.095 | -0.4228 | No |
| 256 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 11396 | -0.095 | -0.4205 | No |
| 257 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 11531 | -0.101 | -0.4283 | No |
| 258 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 11551 | -0.102 | -0.4272 | No |
| 259 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 11588 | -0.104 | -0.4273 | No |
| 260 | SDPR | SDPR Entrez, Source | serum deprivation response (phosphatidylserine binding protein) | 11603 | -0.105 | -0.4258 | No |
| 261 | CAMK2D | CAMK2D Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II delta | 11618 | -0.105 | -0.4243 | No |
| 262 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 11644 | -0.106 | -0.4235 | No |
| 263 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 11745 | -0.111 | -0.4284 | Yes |
| 264 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 11772 | -0.112 | -0.4276 | Yes |
| 265 | ITGA9 | ITGA9 Entrez, Source | integrin, alpha 9 | 11796 | -0.113 | -0.4266 | Yes |
| 266 | STRN | STRN Entrez, Source | striatin, calmodulin binding protein | 11807 | -0.114 | -0.4245 | Yes |
| 267 | SERINC3 | SERINC3 Entrez, Source | serine incorporator 3 | 11857 | -0.117 | -0.4253 | Yes |
| 268 | TRPC1 | TRPC1 Entrez, Source | transient receptor potential cation channel, subfamily C, member 1 | 11873 | -0.118 | -0.4235 | Yes |
| 269 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 11916 | -0.119 | -0.4238 | Yes |
| 270 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 11950 | -0.122 | -0.4233 | Yes |
| 271 | EXOC8 | EXOC8 Entrez, Source | exocyst complex component 8 | 11965 | -0.123 | -0.4213 | Yes |
| 272 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 11990 | -0.124 | -0.4200 | Yes |
| 273 | CBARA1 | CBARA1 Entrez, Source | calcium binding atopy-related autoantigen 1 | 12016 | -0.126 | -0.4188 | Yes |
| 274 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 12036 | -0.127 | -0.4171 | Yes |
| 275 | ANXA4 | ANXA4 Entrez, Source | annexin A4 | 12040 | -0.127 | -0.4141 | Yes |
| 276 | ABCA5 | ABCA5 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 5 | 12067 | -0.129 | -0.4129 | Yes |
| 277 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 12099 | -0.132 | -0.4120 | Yes |
| 278 | RAP1B | RAP1B Entrez, Source | RAP1B, member of RAS oncogene family | 12148 | -0.134 | -0.4123 | Yes |
| 279 | HTRA1 | HTRA1 Entrez, Source | HtrA serine peptidase 1 | 12167 | -0.136 | -0.4103 | Yes |
| 280 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 12179 | -0.137 | -0.4077 | Yes |
| 281 | PBX2 | PBX2 Entrez, Source | pre-B-cell leukemia transcription factor 2 | 12188 | -0.138 | -0.4049 | Yes |
| 282 | GYG1 | GYG1 Entrez, Source | glycogenin 1 | 12194 | -0.139 | -0.4018 | Yes |
| 283 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 12198 | -0.139 | -0.3986 | Yes |
| 284 | LASP1 | LASP1 Entrez, Source | LIM and SH3 protein 1 | 12204 | -0.140 | -0.3954 | Yes |
| 285 | PLSCR4 | PLSCR4 Entrez, Source | phospholipid scramblase 4 | 12219 | -0.141 | -0.3930 | Yes |
| 286 | PIP5K2B | PIP5K2B Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type II, beta | 12221 | -0.142 | -0.3895 | Yes |
| 287 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 12262 | -0.145 | -0.3890 | Yes |
| 288 | CALM2 | CALM2 Entrez, Source | calmodulin 2 (phosphorylase kinase, delta) | 12340 | -0.152 | -0.3911 | Yes |
| 289 | PDE10A | PDE10A Entrez, Source | phosphodiesterase 10A | 12363 | -0.154 | -0.3889 | Yes |
| 290 | GSN | GSN Entrez, Source | gelsolin (amyloidosis, Finnish type) | 12398 | -0.157 | -0.3876 | Yes |
| 291 | GPC1 | GPC1 Entrez, Source | glypican 1 | 12419 | -0.160 | -0.3851 | Yes |
| 292 | PTPRJ | PTPRJ Entrez, Source | protein tyrosine phosphatase, receptor type, J | 12463 | -0.165 | -0.3843 | Yes |
| 293 | PPP1R2 | PPP1R2 Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 2 | 12496 | -0.170 | -0.3825 | Yes |
| 294 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 12504 | -0.172 | -0.3788 | Yes |
| 295 | EHD4 | EHD4 Entrez, Source | EH-domain containing 4 | 12508 | -0.172 | -0.3747 | Yes |
| 296 | PIK3CA | PIK3CA Entrez, Source | phosphoinositide-3-kinase, catalytic, alpha polypeptide | 12536 | -0.176 | -0.3724 | Yes |
| 297 | DYNC1LI2 | DYNC1LI2 Entrez, Source | dynein, cytoplasmic 1, light intermediate chain 2 | 12539 | -0.177 | -0.3681 | Yes |
| 298 | BICD2 | BICD2 Entrez, Source | bicaudal D homolog 2 (Drosophila) | 12558 | -0.179 | -0.3650 | Yes |
| 299 | PLEKHC1 | PLEKHC1 Entrez, Source | pleckstrin homology domain containing, family C (with FERM domain) member 1 | 12574 | -0.181 | -0.3616 | Yes |
| 300 | TGFBR2 | TGFBR2 Entrez, Source | transforming growth factor, beta receptor II (70/80kDa) | 12582 | -0.181 | -0.3576 | Yes |
| 301 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 12594 | -0.184 | -0.3539 | Yes |
| 302 | DSP | DSP Entrez, Source | desmoplakin | 12648 | -0.194 | -0.3531 | Yes |
| 303 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 12691 | -0.203 | -0.3512 | Yes |
| 304 | CAV2 | CAV2 Entrez, Source | caveolin 2 | 12694 | -0.203 | -0.3463 | Yes |
| 305 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 12699 | -0.205 | -0.3415 | Yes |
| 306 | CD47 | CD47 Entrez, Source | CD47 molecule | 12700 | -0.205 | -0.3364 | Yes |
| 307 | LZTFL1 | LZTFL1 Entrez, Source | leucine zipper transcription factor-like 1 | 12702 | -0.205 | -0.3313 | Yes |
| 308 | MYO1E | MYO1E Entrez, Source | myosin IE | 12726 | -0.209 | -0.3279 | Yes |
| 309 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12735 | -0.210 | -0.3232 | Yes |
| 310 | RALB | RALB Entrez, Source | v-ral simian leukemia viral oncogene homolog B (ras related; GTP binding protein) | 12806 | -0.226 | -0.3229 | Yes |
| 311 | PAMCI | PAMCI Entrez, Source | peptidylglycine alpha-amidating monooxygenase COOH-terminal interactor | 12835 | -0.233 | -0.3193 | Yes |
| 312 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 12889 | -0.247 | -0.3172 | Yes |
| 313 | MYADM | MYADM Entrez, Source | myeloid-associated differentiation marker | 12919 | -0.258 | -0.3129 | Yes |
| 314 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12930 | -0.262 | -0.3072 | Yes |
| 315 | FRK | FRK Entrez, Source | fyn-related kinase | 12950 | -0.272 | -0.3018 | Yes |
| 316 | ADAMTS5 | ADAMTS5 Entrez, Source | ADAM metallopeptidase with thrombospondin type 1 motif, 5 (aggrecanase-2) | 12977 | -0.282 | -0.2967 | Yes |
| 317 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 12978 | -0.283 | -0.2897 | Yes |
| 318 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 13011 | -0.297 | -0.2847 | Yes |
| 319 | MYO1C | MYO1C Entrez, Source | myosin IC | 13017 | -0.300 | -0.2776 | Yes |
| 320 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13055 | -0.322 | -0.2724 | Yes |
| 321 | CLIC5 | CLIC5 Entrez, Source | chloride intracellular channel 5 | 13065 | -0.327 | -0.2649 | Yes |
| 322 | TSPAN2 | TSPAN2 Entrez, Source | tetraspanin 2 | 13068 | -0.331 | -0.2568 | Yes |
| 323 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 13088 | -0.346 | -0.2496 | Yes |
| 324 | TRAK2 | TRAK2 Entrez, Source | trafficking protein, kinesin binding 2 | 13108 | -0.363 | -0.2420 | Yes |
| 325 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 13110 | -0.363 | -0.2330 | Yes |
| 326 | AVPR1A | AVPR1A Entrez, Source | arginine vasopressin receptor 1A | 13117 | -0.366 | -0.2242 | Yes |
| 327 | TACC1 | TACC1 Entrez, Source | transforming, acidic coiled-coil containing protein 1 | 13123 | -0.377 | -0.2152 | Yes |
| 328 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 13141 | -0.392 | -0.2067 | Yes |
| 329 | OSBPL5 | OSBPL5 Entrez, Source | oxysterol binding protein-like 5 | 13143 | -0.394 | -0.1969 | Yes |
| 330 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 13153 | -0.404 | -0.1875 | Yes |
| 331 | TSPAN8 | TSPAN8 Entrez, Source | tetraspanin 8 | 13161 | -0.417 | -0.1776 | Yes |
| 332 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 13180 | -0.438 | -0.1680 | Yes |
| 333 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 13198 | -0.463 | -0.1578 | Yes |
| 334 | LMO7 | LMO7 Entrez, Source | LIM domain 7 | 13209 | -0.477 | -0.1466 | Yes |
| 335 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.1347 | Yes |
| 336 | SERINC5 | SERINC5 Entrez, Source | serine incorporator 5 | 13225 | -0.503 | -0.1231 | Yes |
| 337 | VIM | VIM Entrez, Source | vimentin | 13227 | -0.506 | -0.1105 | Yes |
| 338 | NES | NES Entrez, Source | nestin | 13241 | -0.529 | -0.0983 | Yes |
| 339 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13260 | -0.571 | -0.0853 | Yes |
| 340 | AHNAK | AHNAK Entrez, Source | AHNAK nucleoprotein (desmoyokin) | 13286 | -0.695 | -0.0699 | Yes |
| 341 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | -0.0524 | Yes |
| 342 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 13321 | -0.988 | -0.0303 | Yes |
| 343 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | Yes |