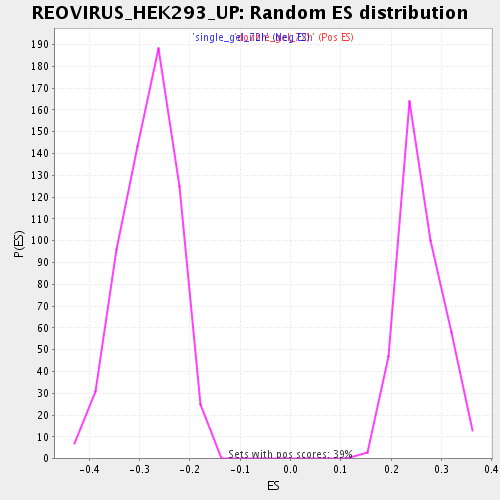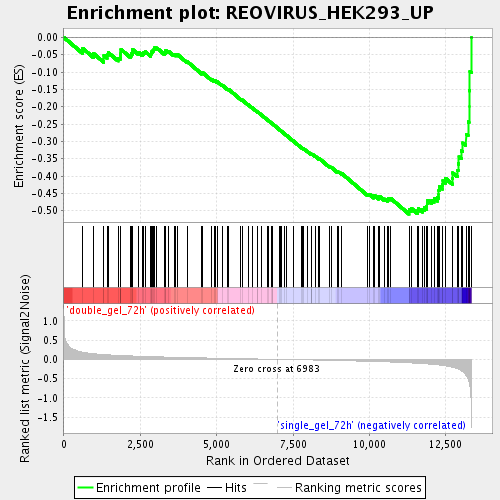
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | REOVIRUS_HEK293_UP |
| Enrichment Score (ES) | -0.50974524 |
| Normalized Enrichment Score (NES) | -1.8043047 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.037203725 |
| FWER p-Value | 0.959 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 613 | 0.184 | -0.0315 | No |
| 2 | PRKAA1 | PRKAA1 Entrez, Source | protein kinase, AMP-activated, alpha 1 catalytic subunit | 967 | 0.148 | -0.0463 | No |
| 3 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 1307 | 0.124 | -0.0619 | No |
| 4 | PIR | PIR Entrez, Source | pirin (iron-binding nuclear protein) | 1308 | 0.124 | -0.0520 | No |
| 5 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1413 | 0.118 | -0.0503 | No |
| 6 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1449 | 0.116 | -0.0436 | No |
| 7 | PREI3 | PREI3 Entrez, Source | preimplantation protein 3 | 1773 | 0.102 | -0.0598 | No |
| 8 | BET1 | BET1 Entrez, Source | BET1 homolog (S. cerevisiae) | 1841 | 0.100 | -0.0569 | No |
| 9 | OAS1 | OAS1 Entrez, Source | 2',5'-oligoadenylate synthetase 1, 40/46kDa | 1850 | 0.099 | -0.0495 | No |
| 10 | GULP1 | GULP1 Entrez, Source | GULP, engulfment adaptor PTB domain containing 1 | 1862 | 0.099 | -0.0423 | No |
| 11 | BNIP1 | BNIP1 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 1 | 1865 | 0.099 | -0.0345 | No |
| 12 | RCHY1 | RCHY1 Entrez, Source | ring finger and CHY zinc finger domain containing 1 | 2191 | 0.088 | -0.0519 | No |
| 13 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 2217 | 0.088 | -0.0468 | No |
| 14 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 2241 | 0.087 | -0.0415 | No |
| 15 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 2245 | 0.086 | -0.0348 | No |
| 16 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 2453 | 0.081 | -0.0439 | No |
| 17 | RBM13 | RBM13 Entrez, Source | RNA binding motif protein 13 | 2581 | 0.077 | -0.0473 | No |
| 18 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 2613 | 0.076 | -0.0435 | No |
| 19 | HIST2H2AA3 | HIST2H2AA3 Entrez, Source | histone cluster 2, H2aa3 | 2660 | 0.075 | -0.0410 | No |
| 20 | CLK1 | CLK1 Entrez, Source | CDC-like kinase 1 | 2848 | 0.070 | -0.0494 | No |
| 21 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 2854 | 0.070 | -0.0442 | No |
| 22 | PITPNB | PITPNB Entrez, Source | phosphatidylinositol transfer protein, beta | 2877 | 0.070 | -0.0402 | No |
| 23 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 2911 | 0.069 | -0.0372 | No |
| 24 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 2944 | 0.068 | -0.0341 | No |
| 25 | SIP1 | SIP1 Entrez, Source | survival of motor neuron protein interacting protein 1 | 2963 | 0.067 | -0.0301 | No |
| 26 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 3036 | 0.066 | -0.0302 | No |
| 27 | TIMM17A | TIMM17A Entrez, Source | translocase of inner mitochondrial membrane 17 homolog A (yeast) | 3292 | 0.060 | -0.0446 | No |
| 28 | SHOX2 | SHOX2 Entrez, Source | short stature homeobox 2 | 3329 | 0.059 | -0.0426 | No |
| 29 | SMAD5 | SMAD5 Entrez, Source | SMAD, mothers against DPP homolog 5 (Drosophila) | 3330 | 0.059 | -0.0378 | No |
| 30 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 3416 | 0.058 | -0.0396 | No |
| 31 | UBE2V2 | UBE2V2 Entrez, Source | ubiquitin-conjugating enzyme E2 variant 2 | 3602 | 0.054 | -0.0492 | No |
| 32 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 3662 | 0.053 | -0.0494 | No |
| 33 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 3719 | 0.052 | -0.0495 | No |
| 34 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 4030 | 0.046 | -0.0693 | No |
| 35 | TAC1 | TAC1 Entrez, Source | tachykinin, precursor 1 (substance K, substance P, neurokinin 1, neurokinin 2, neuromedin L, neurokinin alpha, neuropeptide K, neuropeptide gamma) | 4517 | 0.038 | -0.1030 | No |
| 36 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 4532 | 0.037 | -0.1010 | No |
| 37 | EIF2B5 | EIF2B5 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa | 4827 | 0.032 | -0.1207 | No |
| 38 | CAP2 | CAP2 Entrez, Source | CAP, adenylate cyclase-associated protein, 2 (yeast) | 4920 | 0.030 | -0.1252 | No |
| 39 | WASL | WASL Entrez, Source | Wiskott-Aldrich syndrome-like | 4951 | 0.030 | -0.1251 | No |
| 40 | ARPP-19 | ARPP-19 Entrez, Source | - | 5020 | 0.029 | -0.1279 | No |
| 41 | THUMPD1 | THUMPD1 Entrez, Source | THUMP domain containing 1 | 5182 | 0.026 | -0.1379 | No |
| 42 | CLK3 | CLK3 Entrez, Source | CDC-like kinase 3 | 5359 | 0.024 | -0.1493 | No |
| 43 | MKLN1 | MKLN1 Entrez, Source | muskelin 1, intracellular mediator containing kelch motifs | 5396 | 0.023 | -0.1502 | No |
| 44 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 5794 | 0.017 | -0.1788 | No |
| 45 | NUPL1 | NUPL1 Entrez, Source | nucleoporin like 1 | 5833 | 0.017 | -0.1803 | No |
| 46 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 6026 | 0.014 | -0.1937 | No |
| 47 | SMNDC1 | SMNDC1 Entrez, Source | survival motor neuron domain containing 1 | 6157 | 0.012 | -0.2026 | No |
| 48 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 6331 | 0.009 | -0.2149 | No |
| 49 | PPM1A | PPM1A Entrez, Source | protein phosphatase 1A (formerly 2C), magnesium-dependent, alpha isoform | 6343 | 0.009 | -0.2150 | No |
| 50 | BCL10 | BCL10 Entrez, Source | B-cell CLL/lymphoma 10 | 6471 | 0.007 | -0.2241 | No |
| 51 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 6673 | 0.004 | -0.2389 | No |
| 52 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 6703 | 0.004 | -0.2408 | No |
| 53 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 6801 | 0.003 | -0.2479 | No |
| 54 | RNMT | RNMT Entrez, Source | RNA (guanine-7-) methyltransferase | 6816 | 0.002 | -0.2488 | No |
| 55 | SLC3A2 | SLC3A2 Entrez, Source | solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 | 6817 | 0.002 | -0.2486 | No |
| 56 | MAP2K4 | MAP2K4 Entrez, Source | mitogen-activated protein kinase kinase 4 | 7050 | -0.001 | -0.2661 | No |
| 57 | COPS2 | COPS2 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 2 (Arabidopsis) | 7084 | -0.002 | -0.2684 | No |
| 58 | SNRK | SNRK Entrez, Source | SNF related kinase | 7109 | -0.002 | -0.2701 | No |
| 59 | ZRF1 | ZRF1 Entrez, Source | zuotin related factor 1 | 7203 | -0.003 | -0.2769 | No |
| 60 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 7275 | -0.004 | -0.2819 | No |
| 61 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 7497 | -0.007 | -0.2980 | No |
| 62 | NR1D2 | NR1D2 Entrez, Source | nuclear receptor subfamily 1, group D, member 2 | 7779 | -0.012 | -0.3183 | No |
| 63 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 7803 | -0.012 | -0.3191 | No |
| 64 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 7849 | -0.013 | -0.3215 | No |
| 65 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 7962 | -0.014 | -0.3288 | No |
| 66 | TADA2L | TADA2L Entrez, Source | transcriptional adaptor 2 (ADA2 homolog, yeast)-like | 8099 | -0.016 | -0.3377 | No |
| 67 | SP1 | SP1 Entrez, Source | Sp1 transcription factor | 8115 | -0.016 | -0.3375 | No |
| 68 | OGFR | OGFR Entrez, Source | opioid growth factor receptor | 8117 | -0.016 | -0.3363 | No |
| 69 | IREB2 | IREB2 Entrez, Source | iron-responsive element binding protein 2 | 8229 | -0.018 | -0.3432 | No |
| 70 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 8340 | -0.020 | -0.3499 | No |
| 71 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 8356 | -0.021 | -0.3493 | No |
| 72 | HTATIP | HTATIP Entrez, Source | HIV-1 Tat interacting protein, 60kDa | 8687 | -0.026 | -0.3722 | No |
| 73 | CETN3 | CETN3 Entrez, Source | centrin, EF-hand protein, 3 (CDC31 homolog, yeast) | 8746 | -0.027 | -0.3744 | No |
| 74 | IFITM3 | IFITM3 Entrez, Source | interferon induced transmembrane protein 3 (1-8U) | 8938 | -0.031 | -0.3863 | No |
| 75 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 8998 | -0.032 | -0.3882 | No |
| 76 | YTHDC1 | YTHDC1 Entrez, Source | YTH domain containing 1 | 9099 | -0.034 | -0.3930 | No |
| 77 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 9946 | -0.051 | -0.4528 | No |
| 78 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 9998 | -0.052 | -0.4525 | No |
| 79 | FGF9 | FGF9 Entrez, Source | fibroblast growth factor 9 (glia-activating factor) | 10145 | -0.055 | -0.4591 | No |
| 80 | CAPZA2 | CAPZA2 Entrez, Source | capping protein (actin filament) muscle Z-line, alpha 2 | 10153 | -0.056 | -0.4551 | No |
| 81 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10306 | -0.059 | -0.4618 | No |
| 82 | UBE1C | UBE1C Entrez, Source | ubiquitin-activating enzyme E1C (UBA3 homolog, yeast) | 10319 | -0.060 | -0.4579 | No |
| 83 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 10496 | -0.064 | -0.4661 | No |
| 84 | PRPF4B | PRPF4B Entrez, Source | PRP4 pre-mRNA processing factor 4 homolog B (yeast) | 10586 | -0.066 | -0.4674 | No |
| 85 | PGAM1 | PGAM1 Entrez, Source | phosphoglycerate mutase 1 (brain) | 10612 | -0.067 | -0.4639 | No |
| 86 | SHOC2 | SHOC2 Entrez, Source | soc-2 suppressor of clear homolog (C. elegans) | 10682 | -0.069 | -0.4635 | No |
| 87 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 11294 | -0.090 | -0.5025 | Yes |
| 88 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 11298 | -0.090 | -0.4954 | Yes |
| 89 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 11377 | -0.094 | -0.4938 | Yes |
| 90 | RAB21 | RAB21 Entrez, Source | RAB21, member RAS oncogene family | 11573 | -0.103 | -0.5002 | Yes |
| 91 | STX3 | STX3 Entrez, Source | syntaxin 3 | 11595 | -0.104 | -0.4934 | Yes |
| 92 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 11736 | -0.111 | -0.4951 | Yes |
| 93 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 11801 | -0.114 | -0.4908 | Yes |
| 94 | TRPC1 | TRPC1 Entrez, Source | transient receptor potential cation channel, subfamily C, member 1 | 11873 | -0.118 | -0.4866 | Yes |
| 95 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 11882 | -0.118 | -0.4777 | Yes |
| 96 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 11905 | -0.119 | -0.4698 | Yes |
| 97 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 12036 | -0.127 | -0.4694 | Yes |
| 98 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 12113 | -0.132 | -0.4645 | Yes |
| 99 | TDRD7 | TDRD7 Entrez, Source | tudor domain containing 7 | 12218 | -0.141 | -0.4610 | Yes |
| 100 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 12264 | -0.145 | -0.4527 | Yes |
| 101 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 12269 | -0.146 | -0.4412 | Yes |
| 102 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 12287 | -0.147 | -0.4307 | Yes |
| 103 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 12381 | -0.156 | -0.4251 | Yes |
| 104 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 12403 | -0.158 | -0.4140 | Yes |
| 105 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 12492 | -0.169 | -0.4070 | Yes |
| 106 | MAP3K1 | MAP3K1 Entrez, Source | mitogen-activated protein kinase kinase kinase 1 | 12714 | -0.208 | -0.4069 | Yes |
| 107 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 12725 | -0.209 | -0.3908 | Yes |
| 108 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 12881 | -0.245 | -0.3828 | Yes |
| 109 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 12906 | -0.254 | -0.3641 | Yes |
| 110 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 12929 | -0.262 | -0.3447 | Yes |
| 111 | ZNF292 | ZNF292 Entrez, Source | zinc finger protein 292 | 13006 | -0.293 | -0.3268 | Yes |
| 112 | RRAS2 | RRAS2 Entrez, Source | related RAS viral (r-ras) oncogene homolog 2 | 13055 | -0.322 | -0.3045 | Yes |
| 113 | ABI2 | ABI2 Entrez, Source | abl interactor 2 | 13159 | -0.411 | -0.2791 | Yes |
| 114 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 13229 | -0.509 | -0.2433 | Yes |
| 115 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13260 | -0.571 | -0.1995 | Yes |
| 116 | GNAI1 | GNAI1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | 13262 | -0.580 | -0.1528 | Yes |
| 117 | ADM | ADM Entrez, Source | adrenomedullin | 13287 | -0.697 | -0.0984 | Yes |
| 118 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 13336 | -1.272 | 0.0005 | Yes |