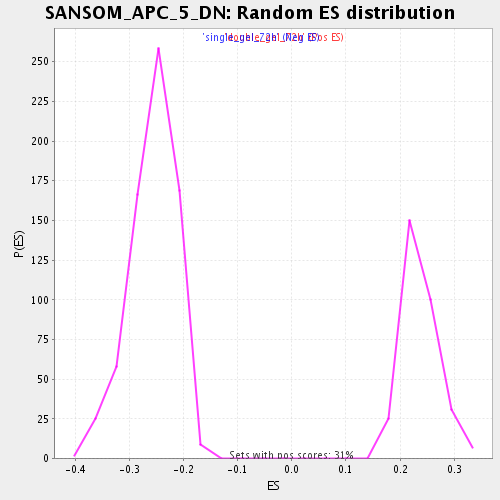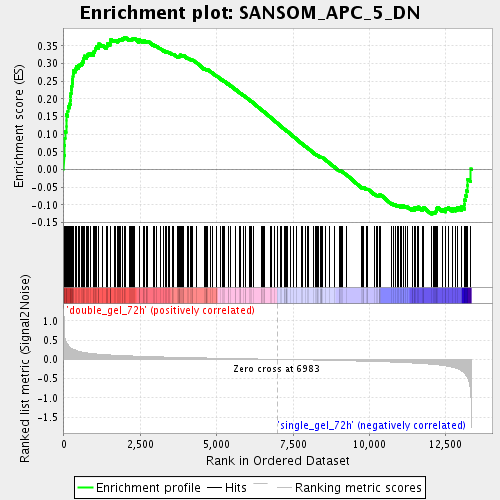
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | SANSOM_APC_5_DN |
| Enrichment Score (ES) | 0.37457597 |
| Normalized Enrichment Score (NES) | 1.5850903 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07401436 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 2 | 1.112 | 0.0408 | Yes |
| 2 | AADAC | AADAC Entrez, Source | arylacetamide deacetylase (esterase) | 11 | 0.758 | 0.0681 | Yes |
| 3 | ABP1 | ABP1 Entrez, Source | amiloride binding protein 1 (amine oxidase (copper-containing)) | 24 | 0.594 | 0.0891 | Yes |
| 4 | USP2 | USP2 Entrez, Source | ubiquitin specific peptidase 2 | 37 | 0.535 | 0.1079 | Yes |
| 5 | RDH7 | 70 | 0.454 | 0.1221 | Yes | ||
| 6 | CD302 | CD302 Entrez, Source | CD302 molecule | 72 | 0.452 | 0.1387 | Yes |
| 7 | NR1I3 | NR1I3 Entrez, Source | nuclear receptor subfamily 1, group I, member 3 | 73 | 0.449 | 0.1552 | Yes |
| 8 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 129 | 0.383 | 0.1652 | Yes |
| 9 | XDH | XDH Entrez, Source | xanthine dehydrogenase | 134 | 0.375 | 0.1787 | Yes |
| 10 | FAHD1 | FAHD1 Entrez, Source | fumarylacetoacetate hydrolase domain containing 1 | 194 | 0.312 | 0.1857 | Yes |
| 11 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 207 | 0.307 | 0.1961 | Yes |
| 12 | MTTP | MTTP Entrez, Source | microsomal triglyceride transfer protein | 218 | 0.301 | 0.2064 | Yes |
| 13 | RBP4 | RBP4 Entrez, Source | retinol binding protein 4, plasma | 229 | 0.294 | 0.2164 | Yes |
| 14 | SLC2A5 | SLC2A5 Entrez, Source | solute carrier family 2 (facilitated glucose/fructose transporter), member 5 | 238 | 0.290 | 0.2265 | Yes |
| 15 | PRLR | PRLR Entrez, Source | prolactin receptor | 257 | 0.285 | 0.2356 | Yes |
| 16 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 266 | 0.281 | 0.2454 | Yes |
| 17 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 282 | 0.274 | 0.2543 | Yes |
| 18 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 296 | 0.267 | 0.2632 | Yes |
| 19 | AKR1B7 | 299 | 0.266 | 0.2728 | Yes | ||
| 20 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 322 | 0.259 | 0.2807 | Yes |
| 21 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 383 | 0.238 | 0.2849 | Yes |
| 22 | ITGAE | ITGAE Entrez, Source | integrin, alpha E (antigen CD103, human mucosal lymphocyte antigen 1; alpha polypeptide) | 410 | 0.228 | 0.2913 | Yes |
| 23 | GSTK1 | GSTK1 Entrez, Source | glutathione S-transferase kappa 1 | 471 | 0.210 | 0.2944 | Yes |
| 24 | LTBP4 | LTBP4 Entrez, Source | latent transforming growth factor beta binding protein 4 | 523 | 0.197 | 0.2978 | Yes |
| 25 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 583 | 0.188 | 0.3002 | Yes |
| 26 | ST8SIA4 | ST8SIA4 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 | 611 | 0.184 | 0.3049 | Yes |
| 27 | MYH14 | MYH14 Entrez, Source | myosin, heavy chain 14 | 634 | 0.182 | 0.3100 | Yes |
| 28 | MERTK | MERTK Entrez, Source | c-mer proto-oncogene tyrosine kinase | 647 | 0.180 | 0.3157 | Yes |
| 29 | TCEA3 | TCEA3 Entrez, Source | transcription elongation factor A (SII), 3 | 657 | 0.179 | 0.3216 | Yes |
| 30 | TYROBP | TYROBP Entrez, Source | TYRO protein tyrosine kinase binding protein | 747 | 0.168 | 0.3210 | Yes |
| 31 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 778 | 0.165 | 0.3247 | Yes |
| 32 | HAGH | HAGH Entrez, Source | hydroxyacylglutathione hydrolase | 819 | 0.161 | 0.3276 | Yes |
| 33 | SRXN1 | SRXN1 Entrez, Source | sulfiredoxin 1 homolog (S. cerevisiae) | 870 | 0.156 | 0.3295 | Yes |
| 34 | SAR1B | SAR1B Entrez, Source | SAR1 gene homolog B (S. cerevisiae) | 968 | 0.148 | 0.3275 | Yes |
| 35 | GSTT1 | GSTT1 Entrez, Source | glutathione S-transferase theta 1 | 970 | 0.147 | 0.3329 | Yes |
| 36 | ALDOB | ALDOB Entrez, Source | aldolase B, fructose-bisphosphate | 1007 | 0.144 | 0.3355 | Yes |
| 37 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 1017 | 0.143 | 0.3401 | Yes |
| 38 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 1041 | 0.141 | 0.3435 | Yes |
| 39 | CD79B | CD79B Entrez, Source | CD79b molecule, immunoglobulin-associated beta | 1051 | 0.140 | 0.3480 | Yes |
| 40 | MCFD2 | MCFD2 Entrez, Source | multiple coagulation factor deficiency 2 | 1127 | 0.135 | 0.3472 | Yes |
| 41 | SLC5A1 | SLC5A1 Entrez, Source | solute carrier family 5 (sodium/glucose cotransporter), member 1 | 1139 | 0.134 | 0.3513 | Yes |
| 42 | CTSW | CTSW Entrez, Source | cathepsin W (lymphopain) | 1146 | 0.134 | 0.3558 | Yes |
| 43 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 1252 | 0.127 | 0.3524 | Yes |
| 44 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 1393 | 0.119 | 0.3461 | Yes |
| 45 | NAPSA | NAPSA Entrez, Source | napsin A aspartic peptidase | 1408 | 0.118 | 0.3494 | Yes |
| 46 | ENTPD2 | ENTPD2 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 2 | 1414 | 0.118 | 0.3534 | Yes |
| 47 | SEC14L2 | SEC14L2 Entrez, Source | SEC14-like 2 (S. cerevisiae) | 1434 | 0.117 | 0.3562 | Yes |
| 48 | CMA1 | CMA1 Entrez, Source | chymase 1, mast cell | 1510 | 0.113 | 0.3547 | Yes |
| 49 | EDN2 | EDN2 Entrez, Source | endothelin 2 | 1518 | 0.112 | 0.3583 | Yes |
| 50 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 1520 | 0.112 | 0.3623 | Yes |
| 51 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1526 | 0.112 | 0.3661 | Yes |
| 52 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 1535 | 0.111 | 0.3696 | Yes |
| 53 | VAV1 | VAV1 Entrez, Source | vav 1 oncogene | 1642 | 0.107 | 0.3654 | Yes |
| 54 | GALM | GALM Entrez, Source | galactose mutarotase (aldose 1-epimerase) | 1687 | 0.105 | 0.3659 | Yes |
| 55 | ATP6V0E2 | ATP6V0E2 Entrez, Source | ATPase, H+ transporting V0 subunit e2 | 1758 | 0.103 | 0.3644 | Yes |
| 56 | SLC35B1 | SLC35B1 Entrez, Source | solute carrier family 35, member B1 | 1780 | 0.102 | 0.3665 | Yes |
| 57 | GIP | GIP Entrez, Source | gastric inhibitory polypeptide | 1806 | 0.101 | 0.3683 | Yes |
| 58 | KEAP1 | KEAP1 Entrez, Source | kelch-like ECH-associated protein 1 | 1858 | 0.099 | 0.3681 | Yes |
| 59 | CCL21B | 1908 | 0.097 | 0.3679 | Yes | ||
| 60 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 1923 | 0.097 | 0.3704 | Yes |
| 61 | PCSK5 | PCSK5 Entrez, Source | proprotein convertase subtilisin/kexin type 5 | 1968 | 0.095 | 0.3705 | Yes |
| 62 | LYL1 | LYL1 Entrez, Source | lymphoblastic leukemia derived sequence 1 | 1977 | 0.095 | 0.3734 | Yes |
| 63 | TCTA | TCTA Entrez, Source | T-cell leukemia translocation altered gene | 2008 | 0.094 | 0.3746 | Yes |
| 64 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 2131 | 0.090 | 0.3686 | No |
| 65 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 2179 | 0.089 | 0.3683 | No |
| 66 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 2212 | 0.088 | 0.3691 | No |
| 67 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 2243 | 0.086 | 0.3700 | No |
| 68 | MGAT3 | MGAT3 Entrez, Source | mannosyl (beta-1,4-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase | 2272 | 0.086 | 0.3710 | No |
| 69 | ENTPD5 | ENTPD5 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 5 | 2300 | 0.085 | 0.3720 | No |
| 70 | SLC2A2 | SLC2A2 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 2 | 2459 | 0.081 | 0.3630 | No |
| 71 | CD53 | CD53 Entrez, Source | CD53 molecule | 2467 | 0.081 | 0.3654 | No |
| 72 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 2470 | 0.081 | 0.3682 | No |
| 73 | ADIPOR2 | ADIPOR2 Entrez, Source | adiponectin receptor 2 | 2598 | 0.076 | 0.3613 | No |
| 74 | ACP5 | ACP5 Entrez, Source | acid phosphatase 5, tartrate resistant | 2601 | 0.076 | 0.3640 | No |
| 75 | ACE | ACE Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 1 | 2622 | 0.076 | 0.3653 | No |
| 76 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 2693 | 0.074 | 0.3626 | No |
| 77 | SLC35C2 | SLC35C2 Entrez, Source | solute carrier family 35, member C2 | 2714 | 0.073 | 0.3638 | No |
| 78 | CD5 | CD5 Entrez, Source | CD5 molecule | 2750 | 0.073 | 0.3638 | No |
| 79 | NALP6 | NALP6 Entrez, Source | NACHT, leucine rich repeat and PYD containing 6 | 2918 | 0.068 | 0.3536 | No |
| 80 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 2974 | 0.067 | 0.3519 | No |
| 81 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 3036 | 0.066 | 0.3496 | No |
| 82 | PDE9A | PDE9A Entrez, Source | phosphodiesterase 9A | 3146 | 0.063 | 0.3436 | No |
| 83 | KCNJ11 | KCNJ11 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 11 | 3266 | 0.061 | 0.3368 | No |
| 84 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 3337 | 0.059 | 0.3336 | No |
| 85 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 3372 | 0.059 | 0.3332 | No |
| 86 | WBP2 | WBP2 Entrez, Source | WW domain binding protein 2 | 3407 | 0.058 | 0.3327 | No |
| 87 | ABCC3 | ABCC3 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 3 | 3450 | 0.057 | 0.3316 | No |
| 88 | MAPRE3 | MAPRE3 Entrez, Source | microtubule-associated protein, RP/EB family, member 3 | 3542 | 0.055 | 0.3267 | No |
| 89 | SLC5A11 | SLC5A11 Entrez, Source | solute carrier family 5 (sodium/glucose cotransporter), member 11 | 3564 | 0.055 | 0.3271 | No |
| 90 | DHRS1 | DHRS1 Entrez, Source | dehydrogenase/reductase (SDR family) member 1 | 3594 | 0.054 | 0.3269 | No |
| 91 | GGTLA1 | GGTLA1 Entrez, Source | gamma-glutamyltransferase-like activity 1 | 3715 | 0.052 | 0.3196 | No |
| 92 | CENPJ | CENPJ Entrez, Source | centromere protein J | 3740 | 0.051 | 0.3197 | No |
| 93 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 3743 | 0.051 | 0.3214 | No |
| 94 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 3781 | 0.050 | 0.3204 | No |
| 95 | CD2 | CD2 Entrez, Source | CD2 molecule | 3794 | 0.050 | 0.3213 | No |
| 96 | DDR2 | DDR2 Entrez, Source | discoidin domain receptor family, member 2 | 3797 | 0.050 | 0.3230 | No |
| 97 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 3800 | 0.050 | 0.3247 | No |
| 98 | CD6 | CD6 Entrez, Source | CD6 molecule | 3856 | 0.049 | 0.3223 | No |
| 99 | SLC22A18 | SLC22A18 Entrez, Source | solute carrier family 22 (organic cation transporter), member 18 | 3870 | 0.049 | 0.3231 | No |
| 100 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 3905 | 0.048 | 0.3223 | No |
| 101 | CYP2C40 | 3910 | 0.048 | 0.3238 | No | ||
| 102 | TPH1 | TPH1 Entrez, Source | tryptophan hydroxylase 1 (tryptophan 5-monooxygenase) | 4036 | 0.046 | 0.3159 | No |
| 103 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 4076 | 0.045 | 0.3146 | No |
| 104 | UPP1 | UPP1 Entrez, Source | uridine phosphorylase 1 | 4127 | 0.044 | 0.3124 | No |
| 105 | CD8A | CD8A Entrez, Source | CD8a molecule | 4174 | 0.043 | 0.3105 | No |
| 106 | RFXANK | RFXANK Entrez, Source | regulatory factor X-associated ankyrin-containing protein | 4200 | 0.043 | 0.3102 | No |
| 107 | CYP27B1 | CYP27B1 Entrez, Source | cytochrome P450, family 27, subfamily B, polypeptide 1 | 4218 | 0.043 | 0.3104 | No |
| 108 | FLT4 | FLT4 Entrez, Source | fms-related tyrosine kinase 4 | 4331 | 0.041 | 0.3034 | No |
| 109 | GPSM3 | GPSM3 Entrez, Source | G-protein signalling modulator 3 (AGS3-like, C. elegans) | 4585 | 0.036 | 0.2854 | No |
| 110 | ARSA | ARSA Entrez, Source | arylsulfatase A | 4628 | 0.036 | 0.2835 | No |
| 111 | BTD | BTD Entrez, Source | biotinidase | 4657 | 0.035 | 0.2826 | No |
| 112 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 4666 | 0.035 | 0.2833 | No |
| 113 | CNTNAP1 | CNTNAP1 Entrez, Source | contactin associated protein 1 | 4667 | 0.035 | 0.2846 | No |
| 114 | SST | SST Entrez, Source | somatostatin | 4712 | 0.034 | 0.2825 | No |
| 115 | SEPT4 | SEPT4 Entrez, Source | septin 4 | 4795 | 0.033 | 0.2774 | No |
| 116 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 4861 | 0.032 | 0.2736 | No |
| 117 | PRPSAP1 | PRPSAP1 Entrez, Source | phosphoribosyl pyrophosphate synthetase-associated protein 1 | 4992 | 0.029 | 0.2648 | No |
| 118 | TMPRSS8 | 5003 | 0.029 | 0.2651 | No | ||
| 119 | DGAT1 | DGAT1 Entrez, Source | diacylglycerol O-acyltransferase homolog 1 (mouse) | 5118 | 0.027 | 0.2574 | No |
| 120 | IL2RB | IL2RB Entrez, Source | interleukin 2 receptor, beta | 5178 | 0.027 | 0.2539 | No |
| 121 | DNASE1 | DNASE1 Entrez, Source | deoxyribonuclease I | 5226 | 0.026 | 0.2513 | No |
| 122 | CCK | CCK Entrez, Source | cholecystokinin | 5255 | 0.025 | 0.2500 | No |
| 123 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 5373 | 0.024 | 0.2420 | No |
| 124 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 5375 | 0.024 | 0.2428 | No |
| 125 | GDI1 | GDI1 Entrez, Source | GDP dissociation inhibitor 1 | 5466 | 0.022 | 0.2367 | No |
| 126 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 5608 | 0.020 | 0.2267 | No |
| 127 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 5630 | 0.020 | 0.2258 | No |
| 128 | SIRPA | SIRPA Entrez, Source | signal-regulatory protein alpha | 5733 | 0.018 | 0.2187 | No |
| 129 | CD3G | CD3G Entrez, Source | CD3g molecule, gamma (CD3-TCR complex) | 5771 | 0.018 | 0.2166 | No |
| 130 | CLCN2 | CLCN2 Entrez, Source | chloride channel 2 | 5793 | 0.017 | 0.2156 | No |
| 131 | PAX8 | PAX8 Entrez, Source | paired box gene 8 | 5866 | 0.016 | 0.2107 | No |
| 132 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 5937 | 0.015 | 0.2059 | No |
| 133 | CAPN9 | CAPN9 Entrez, Source | calpain 9 | 5952 | 0.015 | 0.2054 | No |
| 134 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 6063 | 0.013 | 0.1975 | No |
| 135 | GZMA | GZMA Entrez, Source | granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) | 6121 | 0.012 | 0.1936 | No |
| 136 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 6123 | 0.012 | 0.1940 | No |
| 137 | INPP5D | INPP5D Entrez, Source | inositol polyphosphate-5-phosphatase, 145kDa | 6188 | 0.011 | 0.1895 | No |
| 138 | GSTM1 | GSTM1 Entrez, Source | glutathione S-transferase M1 | 6456 | 0.007 | 0.1694 | No |
| 139 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 6461 | 0.007 | 0.1693 | No |
| 140 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 6489 | 0.007 | 0.1675 | No |
| 141 | RBP2 | RBP2 Entrez, Source | retinol binding protein 2, cellular | 6542 | 0.006 | 0.1638 | No |
| 142 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6567 | 0.006 | 0.1621 | No |
| 143 | ARG2 | ARG2 Entrez, Source | arginase, type II | 6580 | 0.006 | 0.1614 | No |
| 144 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 6764 | 0.003 | 0.1476 | No |
| 145 | IL12RB2 | IL12RB2 Entrez, Source | interleukin 12 receptor, beta 2 | 6792 | 0.003 | 0.1456 | No |
| 146 | IFNA1 | IFNA1 Entrez, Source | interferon, alpha 1 | 6891 | 0.001 | 0.1382 | No |
| 147 | SPHK2 | SPHK2 Entrez, Source | sphingosine kinase 2 | 6991 | -0.000 | 0.1306 | No |
| 148 | NUDT5 | NUDT5 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 5 | 7094 | -0.002 | 0.1229 | No |
| 149 | MEP1B | MEP1B Entrez, Source | meprin A, beta | 7105 | -0.002 | 0.1222 | No |
| 150 | ICOS | ICOS Entrez, Source | inducible T-cell co-stimulator | 7211 | -0.003 | 0.1143 | No |
| 151 | TEP1 | TEP1 Entrez, Source | telomerase-associated protein 1 | 7217 | -0.003 | 0.1141 | No |
| 152 | CD52 | CD52 Entrez, Source | CD52 molecule | 7261 | -0.004 | 0.1109 | No |
| 153 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 7275 | -0.004 | 0.1101 | No |
| 154 | SOD3 | SOD3 Entrez, Source | superoxide dismutase 3, extracellular | 7289 | -0.004 | 0.1093 | No |
| 155 | LTB4R | LTB4R Entrez, Source | leukotriene B4 receptor | 7299 | -0.005 | 0.1087 | No |
| 156 | SDF4 | SDF4 Entrez, Source | stromal cell derived factor 4 | 7312 | -0.005 | 0.1080 | No |
| 157 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 7404 | -0.006 | 0.1013 | No |
| 158 | TMEM50B | TMEM50B Entrez, Source | transmembrane protein 50B | 7522 | -0.008 | 0.0926 | No |
| 159 | GNAZ | GNAZ Entrez, Source | guanine nucleotide binding protein (G protein), alpha z polypeptide | 7605 | -0.009 | 0.0867 | No |
| 160 | CD28 | CD28 Entrez, Source | CD28 molecule | 7763 | -0.011 | 0.0751 | No |
| 161 | ADAM19 | ADAM19 Entrez, Source | ADAM metallopeptidase domain 19 (meltrin beta) | 7823 | -0.012 | 0.0711 | No |
| 162 | NUMB | NUMB Entrez, Source | numb homolog (Drosophila) | 7901 | -0.014 | 0.0657 | No |
| 163 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 7982 | -0.015 | 0.0602 | No |
| 164 | CYB5R3 | CYB5R3 Entrez, Source | cytochrome b5 reductase 3 | 8007 | -0.015 | 0.0589 | No |
| 165 | DAB1 | DAB1 Entrez, Source | disabled homolog 1 (Drosophila) | 8172 | -0.017 | 0.0470 | No |
| 166 | PPAP2A | PPAP2A Entrez, Source | phosphatidic acid phosphatase type 2A | 8226 | -0.018 | 0.0436 | No |
| 167 | ITGB2 | ITGB2 Entrez, Source | integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) | 8281 | -0.020 | 0.0402 | No |
| 168 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 8284 | -0.020 | 0.0408 | No |
| 169 | SLC7A8 | SLC7A8 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 8 | 8292 | -0.020 | 0.0410 | No |
| 170 | CD48 | CD48 Entrez, Source | CD48 molecule | 8344 | -0.021 | 0.0379 | No |
| 171 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 8384 | -0.021 | 0.0357 | No |
| 172 | DDC | DDC Entrez, Source | dopa decarboxylase (aromatic L-amino acid decarboxylase) | 8401 | -0.021 | 0.0352 | No |
| 173 | GPR12 | GPR12 Entrez, Source | G protein-coupled receptor 12 | 8425 | -0.022 | 0.0343 | No |
| 174 | DPEP1 | DPEP1 Entrez, Source | dipeptidase 1 (renal) | 8438 | -0.022 | 0.0342 | No |
| 175 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 8440 | -0.022 | 0.0349 | No |
| 176 | ABCB9 | ABCB9 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 9 | 8454 | -0.022 | 0.0347 | No |
| 177 | GNA11 | GNA11 Entrez, Source | guanine nucleotide binding protein (G protein), alpha 11 (Gq class) | 8559 | -0.024 | 0.0277 | No |
| 178 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 8704 | -0.026 | 0.0177 | No |
| 179 | MCPT2 | 8849 | -0.029 | 0.0077 | No | ||
| 180 | GALNT10 | GALNT10 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 10 (GalNAc-T10) | 9008 | -0.032 | -0.0031 | No |
| 181 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 9012 | -0.032 | -0.0022 | No |
| 182 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 9062 | -0.033 | -0.0047 | No |
| 183 | TRP53INP2 | 9070 | -0.033 | -0.0040 | No | ||
| 184 | CHRNG | CHRNG Entrez, Source | cholinergic receptor, nicotinic, gamma | 9124 | -0.035 | -0.0067 | No |
| 185 | ACY3 | ACY3 Entrez, Source | aspartoacylase (aminocyclase) 3 | 9233 | -0.036 | -0.0137 | No |
| 186 | REG3A | REG3A Entrez, Source | regenerating islet-derived 3 alpha | 9746 | -0.047 | -0.0510 | No |
| 187 | SMPD2 | SMPD2 Entrez, Source | sphingomyelin phosphodiesterase 2, neutral membrane (neutral sphingomyelinase) | 9756 | -0.047 | -0.0500 | No |
| 188 | BCAT2 | BCAT2 Entrez, Source | branched chain aminotransferase 2, mitochondrial | 9815 | -0.048 | -0.0526 | No |
| 189 | SLC7A9 | SLC7A9 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 | 9816 | -0.048 | -0.0509 | No |
| 190 | SOX18 | SOX18 Entrez, Source | SRY (sex determining region Y)-box 18 | 9896 | -0.050 | -0.0550 | No |
| 191 | SYNPO | SYNPO Entrez, Source | synaptopodin | 9931 | -0.051 | -0.0558 | No |
| 192 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 9949 | -0.051 | -0.0552 | No |
| 193 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 10161 | -0.056 | -0.0692 | No |
| 194 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 10237 | -0.058 | -0.0728 | No |
| 195 | CD82 | CD82 Entrez, Source | CD82 molecule | 10257 | -0.058 | -0.0722 | No |
| 196 | MCPT1 | 10317 | -0.060 | -0.0745 | No | ||
| 197 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 10323 | -0.060 | -0.0726 | No |
| 198 | CASP7 | CASP7 Entrez, Source | caspase 7, apoptosis-related cysteine peptidase | 10338 | -0.060 | -0.0715 | No |
| 199 | PSMB8 | PSMB8 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) | 10371 | -0.061 | -0.0717 | No |
| 200 | CDKN2B | CDKN2B Entrez, Source | cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) | 10726 | -0.071 | -0.0961 | No |
| 201 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 10788 | -0.072 | -0.0981 | No |
| 202 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 10842 | -0.074 | -0.0995 | No |
| 203 | TRG@ | TRG@ Entrez, Source | T cell receptor gamma locus | 10920 | -0.076 | -0.1025 | No |
| 204 | CDKN2D | CDKN2D Entrez, Source | cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) | 10949 | -0.077 | -0.1018 | No |
| 205 | REG1 | 11021 | -0.079 | -0.1043 | No | ||
| 206 | GCG | GCG Entrez, Source | glucagon | 11033 | -0.080 | -0.1023 | No |
| 207 | GRINA | GRINA Entrez, Source | glutamate receptor, ionotropic, N-methyl D-asparate-associated protein 1 (glutamate binding) | 11108 | -0.082 | -0.1049 | No |
| 208 | MUCDHL | MUCDHL Entrez, Source | mucin and cadherin-like | 11118 | -0.083 | -0.1025 | No |
| 209 | NPDC1 | NPDC1 Entrez, Source | neural proliferation, differentiation and control, 1 | 11192 | -0.085 | -0.1049 | No |
| 210 | IL18 | IL18 Entrez, Source | interleukin 18 (interferon-gamma-inducing factor) | 11246 | -0.088 | -0.1058 | No |
| 211 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 11394 | -0.095 | -0.1135 | No |
| 212 | ST3GAL4 | ST3GAL4 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 4 | 11401 | -0.095 | -0.1105 | No |
| 213 | CASP6 | CASP6 Entrez, Source | caspase 6, apoptosis-related cysteine peptidase | 11468 | -0.098 | -0.1119 | No |
| 214 | CD3D | CD3D Entrez, Source | CD3d molecule, delta (CD3-TCR complex) | 11471 | -0.098 | -0.1084 | No |
| 215 | MYT1L | MYT1L Entrez, Source | myelin transcription factor 1-like | 11504 | -0.100 | -0.1072 | No |
| 216 | SIRT2 | SIRT2 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 2 (S. cerevisiae) | 11576 | -0.103 | -0.1088 | No |
| 217 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 11588 | -0.104 | -0.1058 | No |
| 218 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 11736 | -0.111 | -0.1130 | No |
| 219 | GSTA4 | GSTA4 Entrez, Source | glutathione S-transferase A4 | 11757 | -0.112 | -0.1104 | No |
| 220 | ITGAL | ITGAL Entrez, Source | integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) | 11774 | -0.112 | -0.1075 | No |
| 221 | ABCC2 | ABCC2 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 2 | 12020 | -0.126 | -0.1215 | No |
| 222 | SLC22A4 | SLC22A4 Entrez, Source | solute carrier family 22 (organic cation transporter), member 4 | 12079 | -0.130 | -0.1212 | No |
| 223 | RGS10 | RGS10 Entrez, Source | regulator of G-protein signalling 10 | 12133 | -0.133 | -0.1203 | No |
| 224 | MTF2 | MTF2 Entrez, Source | metal response element binding transcription factor 2 | 12173 | -0.137 | -0.1183 | No |
| 225 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 12191 | -0.138 | -0.1145 | No |
| 226 | SLC27A4 | SLC27A4 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 4 | 12200 | -0.139 | -0.1099 | No |
| 227 | SLC4A4 | SLC4A4 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 4 | 12225 | -0.142 | -0.1065 | No |
| 228 | LSP1 | LSP1 Entrez, Source | lymphocyte-specific protein 1 | 12383 | -0.156 | -0.1128 | No |
| 229 | PCDHA4 | PCDHA4 Entrez, Source | protocadherin alpha 4 | 12482 | -0.168 | -0.1141 | No |
| 230 | STK10 | STK10 Entrez, Source | serine/threonine kinase 10 | 12500 | -0.171 | -0.1091 | No |
| 231 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 12570 | -0.180 | -0.1077 | No |
| 232 | MAP3K1 | MAP3K1 Entrez, Source | mitogen-activated protein kinase kinase kinase 1 | 12714 | -0.208 | -0.1110 | No |
| 233 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 12802 | -0.225 | -0.1093 | No |
| 234 | JAK2 | JAK2 Entrez, Source | Janus kinase 2 (a protein tyrosine kinase) | 12879 | -0.245 | -0.1061 | No |
| 235 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 13012 | -0.297 | -0.1052 | No |
| 236 | ADORA2B | ADORA2B Entrez, Source | adenosine A2b receptor | 13103 | -0.358 | -0.0989 | No |
| 237 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 13112 | -0.366 | -0.0860 | No |
| 238 | CDR2 | CDR2 Entrez, Source | cerebellar degeneration-related protein 2, 62kDa | 13150 | -0.398 | -0.0742 | No |
| 239 | FABP7 | FABP7 Entrez, Source | fatty acid binding protein 7, brain | 13182 | -0.442 | -0.0603 | No |
| 240 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.0447 | No |
| 241 | NBL1 | NBL1 Entrez, Source | neuroblastoma, suppression of tumorigenicity 1 | 13217 | -0.493 | -0.0269 | No |
| 242 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 13321 | -0.988 | 0.0016 | No |