Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | TNFALPHA_ADIP_DN |
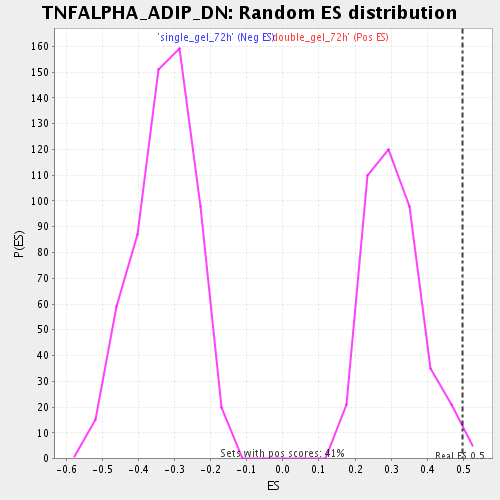
| Enrichment Score (ES) | 0.49875128 |
| Normalized Enrichment Score (NES) | 1.6291535 |
| Nominal p-value | 0.0121951215 |
| FDR q-value | 0.05918308 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 66 | 0.461 | 0.0668 | Yes |
| 2 | FASN | FASN Entrez, Source | fatty acid synthase | 101 | 0.417 | 0.1292 | Yes |
| 3 | PC | PC Entrez, Source | pyruvate carboxylase | 155 | 0.347 | 0.1794 | Yes |
| 4 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.2195 | Yes |
| 5 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 266 | 0.281 | 0.2606 | Yes |
| 6 | RASD1 | RASD1 Entrez, Source | RAS, dexamethasone-induced 1 | 269 | 0.278 | 0.3038 | Yes |
| 7 | TST | TST Entrez, Source | thiosulfate sulfurtransferase (rhodanese) | 388 | 0.235 | 0.3316 | Yes |
| 8 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 419 | 0.224 | 0.3644 | Yes |
| 9 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 583 | 0.188 | 0.3814 | Yes |
| 10 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 609 | 0.185 | 0.4083 | Yes |
| 11 | PEX11A | PEX11A Entrez, Source | peroxisomal biogenesis factor 11A | 631 | 0.182 | 0.4351 | Yes |
| 12 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 662 | 0.178 | 0.4606 | Yes |
| 13 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 800 | 0.163 | 0.4757 | Yes |
| 14 | ALAD | ALAD Entrez, Source | aminolevulinate, delta-, dehydratase | 826 | 0.160 | 0.4988 | Yes |
| 15 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 1494 | 0.114 | 0.4663 | No |
| 16 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 1926 | 0.096 | 0.4489 | No |
| 17 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 2100 | 0.091 | 0.4501 | No |
| 18 | ENTPD5 | ENTPD5 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 5 | 2300 | 0.085 | 0.4484 | No |
| 19 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 2353 | 0.084 | 0.4575 | No |
| 20 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 2453 | 0.081 | 0.4627 | No |
| 21 | DBI | DBI Entrez, Source | diazepam binding inhibitor (GABA receptor modulator, acyl-Coenzyme A binding protein) | 2576 | 0.077 | 0.4656 | No |
| 22 | PLA2G6 | PLA2G6 Entrez, Source | phospholipase A2, group VI (cytosolic, calcium-independent) | 2615 | 0.076 | 0.4745 | No |
| 23 | AQP7 | AQP7 Entrez, Source | aquaporin 7 | 2868 | 0.070 | 0.4665 | No |
| 24 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 3410 | 0.058 | 0.4348 | No |
| 25 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 3592 | 0.054 | 0.4296 | No |
| 26 | TALDO1 | TALDO1 Entrez, Source | transaldolase 1 | 4010 | 0.046 | 0.4054 | No |
| 27 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4299 | 0.041 | 0.3902 | No |
| 28 | DGAT1 | DGAT1 Entrez, Source | diacylglycerol O-acyltransferase homolog 1 (mouse) | 5118 | 0.027 | 0.3330 | No |
| 29 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 5859 | 0.016 | 0.2799 | No |
| 30 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5886 | 0.016 | 0.2804 | No |
| 31 | HIPK3 | HIPK3 Entrez, Source | homeodomain interacting protein kinase 3 | 7248 | -0.004 | 0.1787 | No |
| 32 | CIB2 | CIB2 Entrez, Source | calcium and integrin binding family member 2 | 7716 | -0.011 | 0.1452 | No |
| 33 | LTC4S | LTC4S Entrez, Source | leukotriene C4 synthase | 8127 | -0.017 | 0.1170 | No |
| 34 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 9276 | -0.037 | 0.0365 | No |
| 35 | TPP2 | TPP2 Entrez, Source | tripeptidyl peptidase II | 9613 | -0.044 | 0.0181 | No |
| 36 | SLC2A4 | SLC2A4 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 4 | 9794 | -0.048 | 0.0120 | No |
| 37 | COL15A1 | COL15A1 Entrez, Source | collagen, type XV, alpha 1 | 10011 | -0.052 | 0.0039 | No |
| 38 | ADIPOQ | ADIPOQ Entrez, Source | adiponectin, C1Q and collagen domain containing | 10637 | -0.068 | -0.0325 | No |
| 39 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 11162 | -0.084 | -0.0588 | No |
| 40 | SCARB1 | SCARB1 Entrez, Source | scavenger receptor class B, member 1 | 11498 | -0.099 | -0.0685 | No |
| 41 | FAM62A | FAM62A Entrez, Source | family with sequence similarity 62 (C2 domain containing), member A | 11541 | -0.101 | -0.0559 | No |
| 42 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 12855 | -0.239 | -0.1174 | No |
| 43 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 13321 | -0.988 | 0.0016 | No |