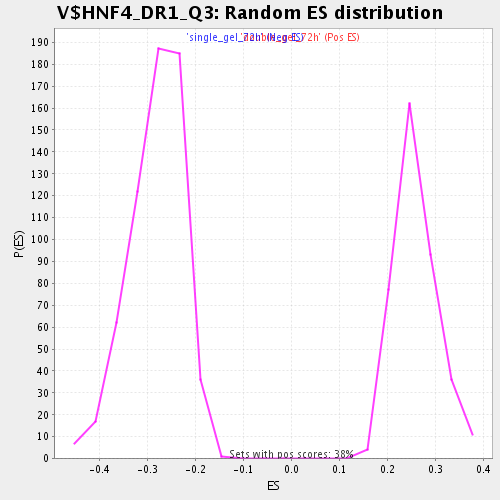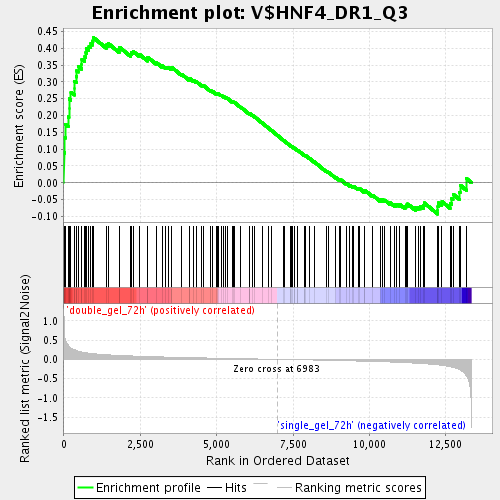
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | V$HNF4_DR1_Q3 |
| Enrichment Score (ES) | 0.43257305 |
| Normalized Enrichment Score (NES) | 1.6754745 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.043059494 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 2 | 1.112 | 0.0903 | Yes |
| 2 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 31 | 0.561 | 0.1338 | Yes |
| 3 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 44 | 0.506 | 0.1740 | Yes |
| 4 | LCAT | LCAT Entrez, Source | lecithin-cholesterol acyltransferase | 142 | 0.365 | 0.1964 | Yes |
| 5 | GUCY2C | GUCY2C Entrez, Source | guanylate cyclase 2C (heat stable enterotoxin receptor) | 167 | 0.341 | 0.2223 | Yes |
| 6 | PRODH2 | PRODH2 Entrez, Source | proline dehydrogenase (oxidase) 2 | 169 | 0.338 | 0.2497 | Yes |
| 7 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 230 | 0.293 | 0.2690 | Yes |
| 8 | FOXA3 | FOXA3 Entrez, Source | forkhead box A3 | 340 | 0.252 | 0.2812 | Yes |
| 9 | F10 | F10 Entrez, Source | coagulation factor X | 342 | 0.251 | 0.3016 | Yes |
| 10 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 406 | 0.229 | 0.3154 | Yes |
| 11 | PEX16 | PEX16 Entrez, Source | peroxisomal biogenesis factor 16 | 421 | 0.224 | 0.3326 | Yes |
| 12 | POLD4 | POLD4 Entrez, Source | polymerase (DNA-directed), delta 4 | 467 | 0.211 | 0.3464 | Yes |
| 13 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 583 | 0.188 | 0.3529 | Yes |
| 14 | ERBB3 | ERBB3 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 3 (avian) | 588 | 0.187 | 0.3679 | Yes |
| 15 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 686 | 0.175 | 0.3748 | Yes |
| 16 | MASP2 | MASP2 Entrez, Source | mannan-binding lectin serine peptidase 2 | 718 | 0.171 | 0.3864 | Yes |
| 17 | INSR | INSR Entrez, Source | insulin receptor | 735 | 0.169 | 0.3989 | Yes |
| 18 | HAGH | HAGH Entrez, Source | hydroxyacylglutathione hydrolase | 819 | 0.161 | 0.4057 | Yes |
| 19 | SLC34A1 | SLC34A1 Entrez, Source | solute carrier family 34 (sodium phosphate), member 1 | 881 | 0.155 | 0.4136 | Yes |
| 20 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 939 | 0.150 | 0.4215 | Yes |
| 21 | ZPBP2 | ZPBP2 Entrez, Source | zona pellucida binding protein 2 | 953 | 0.149 | 0.4326 | Yes |
| 22 | HIBADH | HIBADH Entrez, Source | 3-hydroxyisobutyrate dehydrogenase | 1384 | 0.120 | 0.4098 | No |
| 23 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 1450 | 0.116 | 0.4143 | No |
| 24 | F12 | F12 Entrez, Source | coagulation factor XII (Hageman factor) | 1808 | 0.101 | 0.3955 | No |
| 25 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 1817 | 0.101 | 0.4031 | No |
| 26 | EFEMP1 | EFEMP1 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 1 | 2189 | 0.088 | 0.3822 | No |
| 27 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 2212 | 0.088 | 0.3877 | No |
| 28 | TOMM70A | TOMM70A Entrez, Source | translocase of outer mitochondrial membrane 70 homolog A (S. cerevisiae) | 2268 | 0.086 | 0.3905 | No |
| 29 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 2470 | 0.081 | 0.3819 | No |
| 30 | MRPL2 | MRPL2 Entrez, Source | mitochondrial ribosomal protein L2 | 2738 | 0.073 | 0.3676 | No |
| 31 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 2743 | 0.073 | 0.3732 | No |
| 32 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 3020 | 0.066 | 0.3577 | No |
| 33 | SCN1A | SCN1A Entrez, Source | sodium channel, voltage-gated, type I, alpha | 3219 | 0.062 | 0.3478 | No |
| 34 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 3340 | 0.059 | 0.3435 | No |
| 35 | KCNK10 | KCNK10 Entrez, Source | potassium channel, subfamily K, member 10 | 3406 | 0.058 | 0.3433 | No |
| 36 | LGI4 | LGI4 Entrez, Source | leucine-rich repeat LGI family, member 4 | 3512 | 0.056 | 0.3399 | No |
| 37 | MRPL11 | MRPL11 Entrez, Source | mitochondrial ribosomal protein L11 | 3529 | 0.055 | 0.3432 | No |
| 38 | ENO3 | ENO3 Entrez, Source | enolase 3 (beta, muscle) | 3849 | 0.049 | 0.3231 | No |
| 39 | TTR | TTR Entrez, Source | transthyretin (prealbumin, amyloidosis type I) | 4117 | 0.044 | 0.3065 | No |
| 40 | DAB2IP | DAB2IP Entrez, Source | DAB2 interacting protein | 4122 | 0.044 | 0.3098 | No |
| 41 | TMEM24 | TMEM24 Entrez, Source | transmembrane protein 24 | 4242 | 0.042 | 0.3042 | No |
| 42 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 4337 | 0.040 | 0.3004 | No |
| 43 | KLK6 | KLK6 Entrez, Source | kallikrein 6 (neurosin, zyme) | 4508 | 0.038 | 0.2906 | No |
| 44 | PELO | PELO Entrez, Source | pelota homolog (Drosophila) | 4563 | 0.037 | 0.2895 | No |
| 45 | SCAMP3 | SCAMP3 Entrez, Source | secretory carrier membrane protein 3 | 4809 | 0.032 | 0.2736 | No |
| 46 | FXYD1 | FXYD1 Entrez, Source | FXYD domain containing ion transport regulator 1 (phospholemman) | 4864 | 0.032 | 0.2721 | No |
| 47 | HSD17B8 | HSD17B8 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 8 | 5000 | 0.029 | 0.2643 | No |
| 48 | GATA4 | GATA4 Entrez, Source | GATA binding protein 4 | 5021 | 0.029 | 0.2651 | No |
| 49 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 5059 | 0.028 | 0.2646 | No |
| 50 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 5141 | 0.027 | 0.2606 | No |
| 51 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 5229 | 0.026 | 0.2562 | No |
| 52 | CSF2 | CSF2 Entrez, Source | colony stimulating factor 2 (granulocyte-macrophage) | 5289 | 0.025 | 0.2537 | No |
| 53 | GALK1 | GALK1 Entrez, Source | galactokinase 1 | 5339 | 0.024 | 0.2520 | No |
| 54 | ITGA1 | ITGA1 Entrez, Source | integrin, alpha 1 | 5518 | 0.021 | 0.2403 | No |
| 55 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 5539 | 0.021 | 0.2405 | No |
| 56 | SNAP25 | SNAP25 Entrez, Source | synaptosomal-associated protein, 25kDa | 5580 | 0.021 | 0.2391 | No |
| 57 | CLCN2 | CLCN2 Entrez, Source | chloride channel 2 | 5793 | 0.017 | 0.2245 | No |
| 58 | DNAJA2 | DNAJA2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 2 | 6068 | 0.013 | 0.2049 | No |
| 59 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 6072 | 0.013 | 0.2057 | No |
| 60 | AGPS | AGPS Entrez, Source | alkylglycerone phosphate synthase | 6079 | 0.013 | 0.2063 | No |
| 61 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 6158 | 0.012 | 0.2014 | No |
| 62 | HSPD1 | HSPD1 Entrez, Source | heat shock 60kDa protein 1 (chaperonin) | 6242 | 0.011 | 0.1960 | No |
| 63 | PABPN1 | PABPN1 Entrez, Source | poly(A) binding protein, nuclear 1 | 6488 | 0.007 | 0.1780 | No |
| 64 | ARHGEF12 | ARHGEF12 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 12 | 6688 | 0.004 | 0.1633 | No |
| 65 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 6801 | 0.003 | 0.1550 | No |
| 66 | IHPK2 | IHPK2 Entrez, Source | inositol hexaphosphate kinase 2 | 7190 | -0.003 | 0.1259 | No |
| 67 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7214 | -0.003 | 0.1244 | No |
| 68 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 7404 | -0.006 | 0.1106 | No |
| 69 | GBL | GBL Entrez, Source | - | 7427 | -0.006 | 0.1095 | No |
| 70 | ZBTB12 | ZBTB12 Entrez, Source | zinc finger and BTB domain containing 12 | 7461 | -0.007 | 0.1075 | No |
| 71 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 7482 | -0.007 | 0.1066 | No |
| 72 | SENP2 | SENP2 Entrez, Source | SUMO1/sentrin/SMT3 specific peptidase 2 | 7539 | -0.008 | 0.1030 | No |
| 73 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 7542 | -0.008 | 0.1035 | No |
| 74 | TREX1 | TREX1 Entrez, Source | three prime repair exonuclease 1 | 7643 | -0.010 | 0.0968 | No |
| 75 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 7647 | -0.010 | 0.0973 | No |
| 76 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 7860 | -0.013 | 0.0824 | No |
| 77 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7883 | -0.013 | 0.0818 | No |
| 78 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 7917 | -0.014 | 0.0804 | No |
| 79 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 8036 | -0.015 | 0.0727 | No |
| 80 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 8186 | -0.018 | 0.0629 | No |
| 81 | PCTK1 | PCTK1 Entrez, Source | PCTAIRE protein kinase 1 | 8584 | -0.024 | 0.0349 | No |
| 82 | AP1B1 | AP1B1 Entrez, Source | adaptor-related protein complex 1, beta 1 subunit | 8657 | -0.026 | 0.0315 | No |
| 83 | SYMPK | SYMPK Entrez, Source | symplekin | 8892 | -0.030 | 0.0163 | No |
| 84 | MOBP | MOBP Entrez, Source | myelin-associated oligodendrocyte basic protein | 9009 | -0.032 | 0.0101 | No |
| 85 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 9043 | -0.033 | 0.0103 | No |
| 86 | EIF4EBP2 | EIF4EBP2 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 2 | 9250 | -0.037 | -0.0023 | No |
| 87 | TREH | TREH Entrez, Source | trehalase (brush-border membrane glycoprotein) | 9359 | -0.039 | -0.0073 | No |
| 88 | SNAI1 | SNAI1 Entrez, Source | snail homolog 1 (Drosophila) | 9442 | -0.040 | -0.0102 | No |
| 89 | SMOC1 | SMOC1 Entrez, Source | SPARC related modular calcium binding 1 | 9480 | -0.041 | -0.0096 | No |
| 90 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 9636 | -0.044 | -0.0178 | No |
| 91 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 9664 | -0.045 | -0.0162 | No |
| 92 | BIN3 | BIN3 Entrez, Source | bridging integrator 3 | 9830 | -0.048 | -0.0247 | No |
| 93 | NR2F6 | NR2F6 Entrez, Source | nuclear receptor subfamily 2, group F, member 6 | 9842 | -0.049 | -0.0216 | No |
| 94 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 10111 | -0.055 | -0.0374 | No |
| 95 | DCTN2 | DCTN2 Entrez, Source | dynactin 2 (p50) | 10348 | -0.060 | -0.0504 | No |
| 96 | GTF2I | GTF2I Entrez, Source | general transcription factor II, i | 10410 | -0.062 | -0.0500 | No |
| 97 | PTHR1 | PTHR1 Entrez, Source | parathyroid hormone receptor 1 | 10494 | -0.064 | -0.0510 | No |
| 98 | STMN2 | STMN2 Entrez, Source | stathmin-like 2 | 10679 | -0.069 | -0.0593 | No |
| 99 | ZDHHC3 | ZDHHC3 Entrez, Source | zinc finger, DHHC-type containing 3 | 10824 | -0.073 | -0.0643 | No |
| 100 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 10891 | -0.075 | -0.0632 | No |
| 101 | DCTN1 | DCTN1 Entrez, Source | dynactin 1 (p150, glued homolog, Drosophila) | 10978 | -0.078 | -0.0634 | No |
| 102 | SS18 | SS18 Entrez, Source | synovial sarcoma translocation, chromosome 18 | 11166 | -0.084 | -0.0706 | No |
| 103 | USH1C | USH1C Entrez, Source | Usher syndrome 1C (autosomal recessive, severe) | 11196 | -0.086 | -0.0659 | No |
| 104 | PTMS | PTMS Entrez, Source | parathymosin | 11251 | -0.088 | -0.0628 | No |
| 105 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 11508 | -0.100 | -0.0740 | No |
| 106 | TCN2 | TCN2 Entrez, Source | transcobalamin II; macrocytic anemia | 11593 | -0.104 | -0.0719 | No |
| 107 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 11673 | -0.108 | -0.0692 | No |
| 108 | PFN1 | PFN1 Entrez, Source | profilin 1 | 11762 | -0.112 | -0.0667 | No |
| 109 | ARHGEF7 | ARHGEF7 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 7 | 11790 | -0.113 | -0.0596 | No |
| 110 | ERBB2 | ERBB2 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 12233 | -0.142 | -0.0814 | No |
| 111 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 12239 | -0.143 | -0.0702 | No |
| 112 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 12254 | -0.144 | -0.0595 | No |
| 113 | MARK2 | MARK2 Entrez, Source | MAP/microtubule affinity-regulating kinase 2 | 12371 | -0.155 | -0.0557 | No |
| 114 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 12640 | -0.193 | -0.0603 | No |
| 115 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 12691 | -0.203 | -0.0476 | No |
| 116 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 12735 | -0.210 | -0.0337 | No |
| 117 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12940 | -0.266 | -0.0275 | No |
| 118 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 12969 | -0.279 | -0.0069 | No |
| 119 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 13173 | -0.431 | 0.0128 | No |