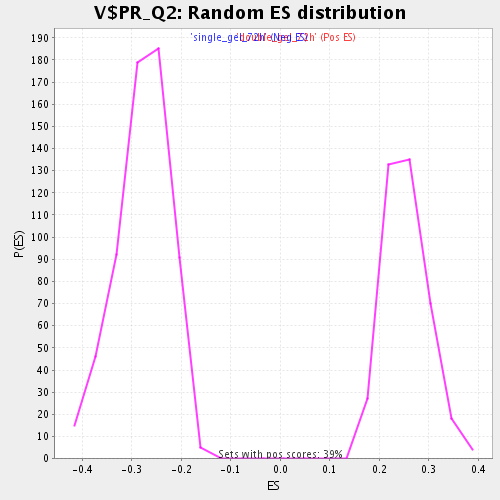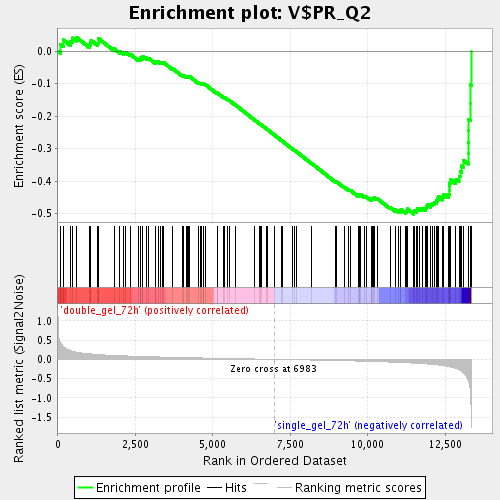
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | V$PR_Q2 |
| Enrichment Score (ES) | -0.50101495 |
| Normalized Enrichment Score (NES) | -1.8021153 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.036899533 |
| FWER p-Value | 0.966 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 83 | 0.433 | 0.0210 | No |
| 2 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 178 | 0.330 | 0.0346 | No |
| 3 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 414 | 0.227 | 0.0311 | No |
| 4 | POLD4 | POLD4 Entrez, Source | polymerase (DNA-directed), delta 4 | 467 | 0.211 | 0.0404 | No |
| 5 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 593 | 0.187 | 0.0427 | No |
| 6 | HFE | HFE Entrez, Source | hemochromatosis | 1027 | 0.142 | 0.0189 | No |
| 7 | MAP2K1IP1 | MAP2K1IP1 Entrez, Source | mitogen-activated protein kinase kinase 1 interacting protein 1 | 1042 | 0.141 | 0.0267 | No |
| 8 | GRM3 | GRM3 Entrez, Source | glutamate receptor, metabotropic 3 | 1066 | 0.139 | 0.0337 | No |
| 9 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 1291 | 0.125 | 0.0246 | No |
| 10 | PAX3 | PAX3 Entrez, Source | paired box gene 3 (Waardenburg syndrome 1) | 1312 | 0.123 | 0.0309 | No |
| 11 | CACNA1G | CACNA1G Entrez, Source | calcium channel, voltage-dependent, alpha 1G subunit | 1313 | 0.123 | 0.0386 | No |
| 12 | OTX1 | OTX1 Entrez, Source | orthodenticle homolog 1 (Drosophila) | 1813 | 0.101 | 0.0072 | No |
| 13 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 1991 | 0.094 | -0.0002 | No |
| 14 | MRPS18B | MRPS18B Entrez, Source | mitochondrial ribosomal protein S18B | 2118 | 0.090 | -0.0041 | No |
| 15 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 2178 | 0.089 | -0.0030 | No |
| 16 | OTP | OTP Entrez, Source | orthopedia homolog (Drosophila) | 2326 | 0.084 | -0.0088 | No |
| 17 | CNTN4 | CNTN4 Entrez, Source | contactin 4 | 2609 | 0.076 | -0.0253 | No |
| 18 | AMY2A | AMY2A Entrez, Source | amylase, alpha 2A; pancreatic | 2658 | 0.075 | -0.0243 | No |
| 19 | ZDHHC22 | ZDHHC22 Entrez, Source | zinc finger, DHHC-type containing 22 | 2675 | 0.074 | -0.0208 | No |
| 20 | TNMD | TNMD Entrez, Source | tenomodulin | 2726 | 0.073 | -0.0200 | No |
| 21 | WNT2 | WNT2 Entrez, Source | wingless-type MMTV integration site family member 2 | 2739 | 0.073 | -0.0163 | No |
| 22 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 2854 | 0.070 | -0.0206 | No |
| 23 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 2936 | 0.068 | -0.0224 | No |
| 24 | TNNI1 | TNNI1 Entrez, Source | troponin I type 1 (skeletal, slow) | 3145 | 0.063 | -0.0341 | No |
| 25 | KCNJ1 | KCNJ1 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 1 | 3163 | 0.063 | -0.0315 | No |
| 26 | SND1 | SND1 Entrez, Source | - | 3230 | 0.062 | -0.0326 | No |
| 27 | KCNB2 | KCNB2 Entrez, Source | potassium voltage-gated channel, Shab-related subfamily, member 2 | 3320 | 0.060 | -0.0356 | No |
| 28 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 3372 | 0.059 | -0.0357 | No |
| 29 | EYA2 | EYA2 Entrez, Source | eyes absent homolog 2 (Drosophila) | 3402 | 0.058 | -0.0343 | No |
| 30 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 3710 | 0.052 | -0.0543 | No |
| 31 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 4035 | 0.046 | -0.0759 | No |
| 32 | IL23A | IL23A Entrez, Source | interleukin 23, alpha subunit p19 | 4043 | 0.046 | -0.0735 | No |
| 33 | SMPX | SMPX Entrez, Source | small muscle protein, X-linked | 4135 | 0.044 | -0.0777 | No |
| 34 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 4177 | 0.043 | -0.0780 | No |
| 35 | GRM2 | GRM2 Entrez, Source | glutamate receptor, metabotropic 2 | 4198 | 0.043 | -0.0769 | No |
| 36 | TMEM24 | TMEM24 Entrez, Source | transmembrane protein 24 | 4242 | 0.042 | -0.0774 | No |
| 37 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 4541 | 0.037 | -0.0976 | No |
| 38 | KCNIP1 | KCNIP1 Entrez, Source | Kv channel interacting protein 1 | 4586 | 0.036 | -0.0987 | No |
| 39 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 4640 | 0.035 | -0.1005 | No |
| 40 | RWDD1 | RWDD1 Entrez, Source | RWD domain containing 1 | 4687 | 0.034 | -0.1018 | No |
| 41 | RHEBL1 | RHEBL1 Entrez, Source | Ras homolog enriched in brain like 1 | 4688 | 0.034 | -0.0996 | No |
| 42 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 4754 | 0.033 | -0.1024 | No |
| 43 | FMO1 | FMO1 Entrez, Source | flavin containing monooxygenase 1 | 5137 | 0.027 | -0.1296 | No |
| 44 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 5141 | 0.027 | -0.1281 | No |
| 45 | CA5B | CA5B Entrez, Source | carbonic anhydrase VB, mitochondrial | 5338 | 0.024 | -0.1415 | No |
| 46 | OBSCN | OBSCN Entrez, Source | obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF | 5377 | 0.024 | -0.1429 | No |
| 47 | RAB3C | RAB3C Entrez, Source | RAB3C, member RAS oncogene family | 5461 | 0.022 | -0.1477 | No |
| 48 | ADCY3 | ADCY3 Entrez, Source | adenylate cyclase 3 | 5521 | 0.021 | -0.1508 | No |
| 49 | KCNC1 | KCNC1 Entrez, Source | potassium voltage-gated channel, Shaw-related subfamily, member 1 | 5723 | 0.018 | -0.1649 | No |
| 50 | AHSG | AHSG Entrez, Source | alpha-2-HS-glycoprotein | 6354 | 0.009 | -0.2120 | No |
| 51 | ZBTB9 | ZBTB9 Entrez, Source | zinc finger and BTB domain containing 9 | 6507 | 0.006 | -0.2231 | No |
| 52 | HAPLN2 | HAPLN2 Entrez, Source | hyaluronan and proteoglycan link protein 2 | 6545 | 0.006 | -0.2255 | No |
| 53 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 6579 | 0.006 | -0.2277 | No |
| 54 | KCNK3 | KCNK3 Entrez, Source | potassium channel, subfamily K, member 3 | 6725 | 0.004 | -0.2384 | No |
| 55 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 6768 | 0.003 | -0.2414 | No |
| 56 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 6998 | -0.000 | -0.2587 | No |
| 57 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7214 | -0.003 | -0.2748 | No |
| 58 | LTBP3 | LTBP3 Entrez, Source | latent transforming growth factor beta binding protein 3 | 7258 | -0.004 | -0.2778 | No |
| 59 | SYP | SYP Entrez, Source | synaptophysin | 7562 | -0.008 | -0.3002 | No |
| 60 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 7647 | -0.010 | -0.3059 | No |
| 61 | PDC | PDC Entrez, Source | phosducin | 7700 | -0.010 | -0.3092 | No |
| 62 | NR2F1 | NR2F1 Entrez, Source | nuclear receptor subfamily 2, group F, member 1 | 8175 | -0.018 | -0.3439 | No |
| 63 | VGF | VGF Entrez, Source | VGF nerve growth factor inducible | 8973 | -0.032 | -0.4022 | No |
| 64 | FILIP1 | FILIP1 Entrez, Source | filamin A interacting protein 1 | 8977 | -0.032 | -0.4004 | No |
| 65 | FGF10 | FGF10 Entrez, Source | fibroblast growth factor 10 | 9238 | -0.037 | -0.4178 | No |
| 66 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 9383 | -0.039 | -0.4262 | No |
| 67 | PRKAG1 | PRKAG1 Entrez, Source | protein kinase, AMP-activated, gamma 1 non-catalytic subunit | 9431 | -0.040 | -0.4272 | No |
| 68 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 9690 | -0.045 | -0.4439 | No |
| 69 | CDH6 | CDH6 Entrez, Source | cadherin 6, type 2, K-cadherin (fetal kidney) | 9721 | -0.046 | -0.4433 | No |
| 70 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 9737 | -0.046 | -0.4415 | No |
| 71 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 9778 | -0.047 | -0.4415 | No |
| 72 | GRAP | GRAP Entrez, Source | GRB2-related adaptor protein | 9889 | -0.050 | -0.4467 | No |
| 73 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 9949 | -0.051 | -0.4480 | No |
| 74 | NCDN | NCDN Entrez, Source | neurochondrin | 10110 | -0.055 | -0.4567 | No |
| 75 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 10111 | -0.055 | -0.4532 | No |
| 76 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 10163 | -0.056 | -0.4536 | No |
| 77 | BRD4 | BRD4 Entrez, Source | bromodomain containing 4 | 10182 | -0.056 | -0.4514 | No |
| 78 | CAPN6 | CAPN6 Entrez, Source | calpain 6 | 10215 | -0.057 | -0.4503 | No |
| 79 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10306 | -0.059 | -0.4533 | No |
| 80 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 10720 | -0.070 | -0.4801 | No |
| 81 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 10885 | -0.075 | -0.4878 | No |
| 82 | BACE2 | BACE2 Entrez, Source | beta-site APP-cleaving enzyme 2 | 10980 | -0.078 | -0.4901 | No |
| 83 | GABRA1 | GABRA1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 1 | 11063 | -0.081 | -0.4912 | No |
| 84 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 11071 | -0.081 | -0.4866 | No |
| 85 | PRG2 | PRG2 Entrez, Source | proteoglycan 2, bone marrow (natural killer cell activator, eosinophil granule major basic protein) | 11222 | -0.087 | -0.4925 | No |
| 86 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 11264 | -0.089 | -0.4901 | No |
| 87 | PACS1 | PACS1 Entrez, Source | phosphofurin acidic cluster sorting protein 1 | 11271 | -0.089 | -0.4849 | No |
| 88 | RERE | RERE Entrez, Source | arginine-glutamic acid dipeptide (RE) repeats | 11485 | -0.099 | -0.4948 | Yes |
| 89 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 11508 | -0.100 | -0.4902 | Yes |
| 90 | SHH | SHH Entrez, Source | sonic hedgehog homolog (Drosophila) | 11581 | -0.103 | -0.4891 | Yes |
| 91 | TCN2 | TCN2 Entrez, Source | transcobalamin II; macrocytic anemia | 11593 | -0.104 | -0.4834 | Yes |
| 92 | RGL2 | RGL2 Entrez, Source | ral guanine nucleotide dissociation stimulator-like 2 | 11681 | -0.108 | -0.4832 | Yes |
| 93 | ITGAL | ITGAL Entrez, Source | integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) | 11774 | -0.112 | -0.4831 | Yes |
| 94 | CNTF | CNTF Entrez, Source | ciliary neurotrophic factor | 11868 | -0.117 | -0.4828 | Yes |
| 95 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11886 | -0.118 | -0.4766 | Yes |
| 96 | SLC15A2 | SLC15A2 Entrez, Source | solute carrier family 15 (H+/peptide transporter), member 2 | 11919 | -0.119 | -0.4715 | Yes |
| 97 | NNAT | NNAT Entrez, Source | neuronatin | 12027 | -0.127 | -0.4717 | Yes |
| 98 | SLC38A6 | SLC38A6 Entrez, Source | solute carrier family 38, member 6 | 12082 | -0.130 | -0.4676 | Yes |
| 99 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 12152 | -0.135 | -0.4643 | Yes |
| 100 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 12212 | -0.140 | -0.4599 | Yes |
| 101 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 12239 | -0.143 | -0.4529 | Yes |
| 102 | CAMLG | CAMLG Entrez, Source | calcium modulating ligand | 12277 | -0.146 | -0.4465 | Yes |
| 103 | RGN | RGN Entrez, Source | regucalcin (senescence marker protein-30) | 12427 | -0.161 | -0.4477 | Yes |
| 104 | TSGA10 | TSGA10 Entrez, Source | testis specific, 10 | 12455 | -0.164 | -0.4394 | Yes |
| 105 | KRT20 | KRT20 Entrez, Source | keratin 20 | 12610 | -0.186 | -0.4393 | Yes |
| 106 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 12628 | -0.189 | -0.4287 | Yes |
| 107 | CACNB3 | CACNB3 Entrez, Source | calcium channel, voltage-dependent, beta 3 subunit | 12633 | -0.191 | -0.4170 | Yes |
| 108 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 12640 | -0.193 | -0.4054 | Yes |
| 109 | SH3BP4 | SH3BP4 Entrez, Source | SH3-domain binding protein 4 | 12682 | -0.201 | -0.3958 | Yes |
| 110 | DTX2 | DTX2 Entrez, Source | deltex homolog 2 (Drosophila) | 12845 | -0.236 | -0.3933 | Yes |
| 111 | CD96 | CD96 Entrez, Source | CD96 molecule | 12946 | -0.269 | -0.3839 | Yes |
| 112 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 12984 | -0.284 | -0.3688 | Yes |
| 113 | RAP2B | RAP2B Entrez, Source | RAP2B, member of RAS oncogene family | 13030 | -0.307 | -0.3530 | Yes |
| 114 | ONECUT1 | ONECUT1 Entrez, Source | one cut domain, family member 1 | 13087 | -0.346 | -0.3354 | Yes |
| 115 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 13242 | -0.532 | -0.3137 | Yes |
| 116 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 13245 | -0.543 | -0.2797 | Yes |
| 117 | NEXN | NEXN Entrez, Source | nexilin (F actin binding protein) | 13256 | -0.568 | -0.2447 | Yes |
| 118 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 13260 | -0.571 | -0.2090 | Yes |
| 119 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13312 | -0.847 | -0.1596 | Yes |
| 120 | LOX | LOX Entrez, Source | lysyl oxidase | 13315 | -0.896 | -0.1034 | Yes |
| 121 | CD24 | CD24 Entrez, Source | CD24 molecule | 13341 | -1.675 | 0.0001 | Yes |