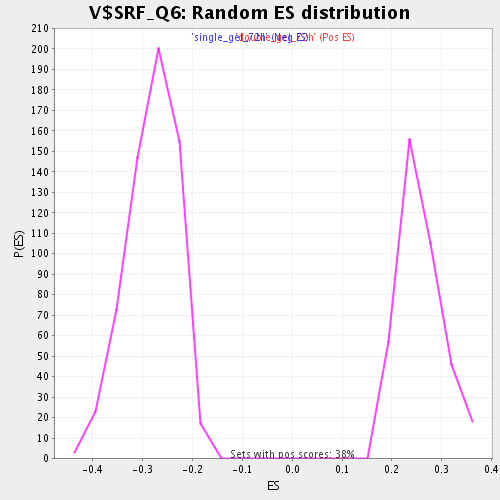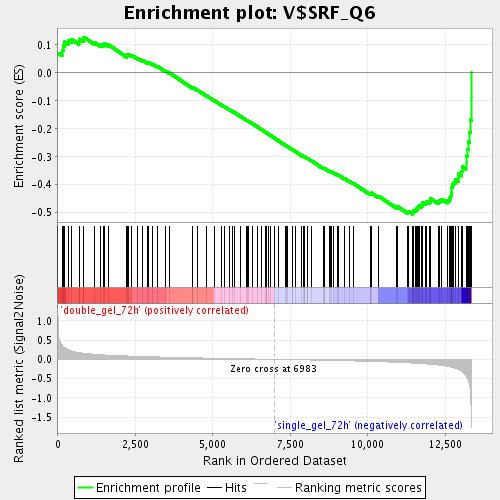
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_72h_versus_single_gel_72h.class.cls #double_gel_72h_versus_single_gel_72h_repos |
| Phenotype | class.cls#double_gel_72h_versus_single_gel_72h_repos |
| Upregulated in class | single_gel_72h |
| GeneSet | V$SRF_Q6 |
| Enrichment Score (ES) | -0.5048628 |
| Normalized Enrichment Score (NES) | -1.8061131 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.036702953 |
| FWER p-Value | 0.953 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 0 | 1.284 | 0.0712 | No |
| 2 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 135 | 0.374 | 0.0818 | No |
| 3 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 180 | 0.328 | 0.0966 | No |
| 4 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 214 | 0.304 | 0.1109 | No |
| 5 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 329 | 0.257 | 0.1166 | No |
| 6 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 437 | 0.219 | 0.1206 | No |
| 7 | SRD5A2 | SRD5A2 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 2 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 2) | 681 | 0.176 | 0.1120 | No |
| 8 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 686 | 0.175 | 0.1213 | No |
| 9 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 829 | 0.160 | 0.1195 | No |
| 10 | CABP1 | CABP1 Entrez, Source | calcium binding protein 1 (calbrain) | 839 | 0.159 | 0.1276 | No |
| 11 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 1182 | 0.131 | 0.1090 | No |
| 12 | GNG8 | GNG8 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 8 | 1368 | 0.121 | 0.1016 | No |
| 13 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 1471 | 0.115 | 0.1003 | No |
| 14 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 1487 | 0.114 | 0.1055 | No |
| 15 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 1619 | 0.108 | 0.1015 | No |
| 16 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 2225 | 0.087 | 0.0606 | No |
| 17 | FOXE3 | FOXE3 Entrez, Source | forkhead box E3 | 2231 | 0.087 | 0.0650 | No |
| 18 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2291 | 0.085 | 0.0653 | No |
| 19 | KCNMB1 | KCNMB1 Entrez, Source | potassium large conductance calcium-activated channel, subfamily M, beta member 1 | 2370 | 0.083 | 0.0640 | No |
| 20 | PDLIM3 | PDLIM3 Entrez, Source | PDZ and LIM domain 3 | 2579 | 0.077 | 0.0526 | No |
| 21 | TNMD | TNMD Entrez, Source | tenomodulin | 2726 | 0.073 | 0.0455 | No |
| 22 | CSNK1E | CSNK1E Entrez, Source | casein kinase 1, epsilon | 2902 | 0.069 | 0.0361 | No |
| 23 | ITGA7 | ITGA7 Entrez, Source | integrin, alpha 7 | 2929 | 0.068 | 0.0379 | No |
| 24 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3058 | 0.065 | 0.0319 | No |
| 25 | NPPA | NPPA Entrez, Source | natriuretic peptide precursor A | 3206 | 0.062 | 0.0242 | No |
| 26 | FGF8 | FGF8 Entrez, Source | fibroblast growth factor 8 (androgen-induced) | 3478 | 0.056 | 0.0068 | No |
| 27 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 3592 | 0.054 | 0.0012 | No |
| 28 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 4346 | 0.040 | -0.0535 | No |
| 29 | PDLIM4 | PDLIM4 Entrez, Source | PDZ and LIM domain 4 | 4357 | 0.040 | -0.0520 | No |
| 30 | JPH2 | JPH2 Entrez, Source | junctophilin 2 | 4515 | 0.038 | -0.0618 | No |
| 31 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 4806 | 0.032 | -0.0820 | No |
| 32 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 5059 | 0.028 | -0.0995 | No |
| 33 | PADI2 | PADI2 Entrez, Source | peptidyl arginine deiminase, type II | 5275 | 0.025 | -0.1144 | No |
| 34 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 5375 | 0.024 | -0.1206 | No |
| 35 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 5523 | 0.021 | -0.1305 | No |
| 36 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 5630 | 0.020 | -0.1374 | No |
| 37 | PRELP | PRELP Entrez, Source | proline/arginine-rich end leucine-rich repeat protein | 5708 | 0.019 | -0.1422 | No |
| 38 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 5896 | 0.016 | -0.1555 | No |
| 39 | HOXA1 | HOXA1 Entrez, Source | homeobox A1 | 6083 | 0.013 | -0.1688 | No |
| 40 | CTDSP1 | CTDSP1 Entrez, Source | CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 1 | 6119 | 0.013 | -0.1708 | No |
| 41 | TRDN | TRDN Entrez, Source | triadin | 6145 | 0.012 | -0.1720 | No |
| 42 | FGFRL1 | FGFRL1 Entrez, Source | fibroblast growth factor receptor-like 1 | 6270 | 0.010 | -0.1808 | No |
| 43 | PLCB3 | PLCB3 Entrez, Source | phospholipase C, beta 3 (phosphatidylinositol-specific) | 6441 | 0.007 | -0.1933 | No |
| 44 | SCN3B | SCN3B Entrez, Source | sodium channel, voltage-gated, type III, beta | 6584 | 0.005 | -0.2037 | No |
| 45 | NKX6-1 | NKX6-1 Entrez, Source | NK6 transcription factor related, locus 1 (Drosophila) | 6704 | 0.004 | -0.2125 | No |
| 46 | MYH11 | MYH11 Entrez, Source | myosin, heavy chain 11, smooth muscle | 6721 | 0.004 | -0.2135 | No |
| 47 | KCNK3 | KCNK3 Entrez, Source | potassium channel, subfamily K, member 3 | 6725 | 0.004 | -0.2136 | No |
| 48 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 6801 | 0.003 | -0.2191 | No |
| 49 | CACNA1B | CACNA1B Entrez, Source | calcium channel, voltage-dependent, L type, alpha 1B subunit | 6851 | 0.002 | -0.2227 | No |
| 50 | RHOJ | RHOJ Entrez, Source | ras homolog gene family, member J | 7002 | -0.000 | -0.2340 | No |
| 51 | MAP1A | MAP1A Entrez, Source | microtubule-associated protein 1A | 7116 | -0.002 | -0.2425 | No |
| 52 | TNNC1 | TNNC1 Entrez, Source | troponin C type 1 (slow) | 7348 | -0.005 | -0.2596 | No |
| 53 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 7382 | -0.006 | -0.2618 | No |
| 54 | IL17B | IL17B Entrez, Source | interleukin 17B | 7417 | -0.006 | -0.2641 | No |
| 55 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 7556 | -0.008 | -0.2740 | No |
| 56 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 7577 | -0.009 | -0.2751 | No |
| 57 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 7676 | -0.010 | -0.2819 | No |
| 58 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 7871 | -0.013 | -0.2959 | No |
| 59 | DGKG | DGKG Entrez, Source | diacylglycerol kinase, gamma 90kDa | 7914 | -0.014 | -0.2983 | No |
| 60 | PRKAB2 | PRKAB2 Entrez, Source | protein kinase, AMP-activated, beta 2 non-catalytic subunit | 7945 | -0.014 | -0.2998 | No |
| 61 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 7948 | -0.014 | -0.2991 | No |
| 62 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 8051 | -0.016 | -0.3060 | No |
| 63 | MATR3 | MATR3 Entrez, Source | matrin 3 | 8066 | -0.016 | -0.3062 | No |
| 64 | NR2F1 | NR2F1 Entrez, Source | nuclear receptor subfamily 2, group F, member 1 | 8175 | -0.018 | -0.3134 | No |
| 65 | CKM | CKM Entrez, Source | creatine kinase, muscle | 8557 | -0.024 | -0.3409 | No |
| 66 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 8587 | -0.025 | -0.3417 | No |
| 67 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 8589 | -0.025 | -0.3404 | No |
| 68 | MRGPRF | MRGPRF Entrez, Source | MAS-related GPR, member F | 8754 | -0.027 | -0.3513 | No |
| 69 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 8798 | -0.028 | -0.3530 | No |
| 70 | TES | TES Entrez, Source | testis derived transcript (3 LIM domains) | 8843 | -0.029 | -0.3547 | No |
| 71 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 8904 | -0.030 | -0.3576 | No |
| 72 | GPR20 | GPR20 Entrez, Source | G protein-coupled receptor 20 | 9010 | -0.032 | -0.3637 | No |
| 73 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 9062 | -0.033 | -0.3657 | No |
| 74 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 9252 | -0.037 | -0.3780 | No |
| 75 | SLC4A3 | SLC4A3 Entrez, Source | solute carrier family 4, anion exchanger, member 3 | 9406 | -0.040 | -0.3874 | No |
| 76 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 9531 | -0.042 | -0.3944 | No |
| 77 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 10088 | -0.054 | -0.4335 | No |
| 78 | DHH | DHH Entrez, Source | desert hedgehog homolog (Drosophila) | 10100 | -0.054 | -0.4313 | No |
| 79 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 10111 | -0.055 | -0.4291 | No |
| 80 | KCNA1 | KCNA1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic ataxia with myokymia) | 10336 | -0.060 | -0.4427 | No |
| 81 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 10358 | -0.060 | -0.4409 | No |
| 82 | ATP1A2 | ATP1A2 Entrez, Source | ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide | 10925 | -0.076 | -0.4795 | No |
| 83 | RIMS1 | RIMS1 Entrez, Source | regulating synaptic membrane exocytosis 1 | 10964 | -0.077 | -0.4781 | No |
| 84 | MUS81 | MUS81 Entrez, Source | MUS81 endonuclease homolog (S. cerevisiae) | 11274 | -0.089 | -0.4965 | No |
| 85 | AQP2 | AQP2 Entrez, Source | aquaporin 2 (collecting duct) | 11323 | -0.092 | -0.4951 | No |
| 86 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 11453 | -0.098 | -0.4995 | Yes |
| 87 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 11475 | -0.099 | -0.4956 | Yes |
| 88 | RERE | RERE Entrez, Source | arginine-glutamic acid dipeptide (RE) repeats | 11485 | -0.099 | -0.4908 | Yes |
| 89 | ACTB | ACTB Entrez, Source | actin, beta | 11552 | -0.102 | -0.4901 | Yes |
| 90 | RASD2 | RASD2 Entrez, Source | RASD family, member 2 | 11574 | -0.103 | -0.4860 | Yes |
| 91 | SDPR | SDPR Entrez, Source | serum deprivation response (phosphatidylserine binding protein) | 11603 | -0.105 | -0.4824 | Yes |
| 92 | DIXDC1 | DIXDC1 Entrez, Source | DIX domain containing 1 | 11625 | -0.106 | -0.4781 | Yes |
| 93 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 11658 | -0.107 | -0.4746 | Yes |
| 94 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 11727 | -0.110 | -0.4736 | Yes |
| 95 | PFN1 | PFN1 Entrez, Source | profilin 1 | 11762 | -0.112 | -0.4700 | Yes |
| 96 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 11764 | -0.112 | -0.4638 | Yes |
| 97 | CNTF | CNTF Entrez, Source | ciliary neurotrophic factor | 11868 | -0.117 | -0.4651 | Yes |
| 98 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 11886 | -0.118 | -0.4599 | Yes |
| 99 | TPM3 | TPM3 Entrez, Source | tropomyosin 3 | 11979 | -0.123 | -0.4600 | Yes |
| 100 | ACTR3 | ACTR3 Entrez, Source | ARP3 actin-related protein 3 homolog (yeast) | 12015 | -0.126 | -0.4557 | Yes |
| 101 | NNAT | NNAT Entrez, Source | neuronatin | 12027 | -0.127 | -0.4495 | Yes |
| 102 | GRK6 | GRK6 Entrez, Source | G protein-coupled receptor kinase 6 | 12273 | -0.146 | -0.4599 | Yes |
| 103 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 12313 | -0.149 | -0.4546 | Yes |
| 104 | LSP1 | LSP1 Entrez, Source | lymphocyte-specific protein 1 | 12383 | -0.156 | -0.4512 | Yes |
| 105 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 12570 | -0.180 | -0.4552 | Yes |
| 106 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 12640 | -0.193 | -0.4498 | Yes |
| 107 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 12657 | -0.196 | -0.4402 | Yes |
| 108 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 12690 | -0.203 | -0.4313 | Yes |
| 109 | PLN | PLN Entrez, Source | phospholamban | 12709 | -0.206 | -0.4213 | Yes |
| 110 | SLC25A4 | SLC25A4 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 | 12710 | -0.207 | -0.4098 | Yes |
| 111 | MYO1E | MYO1E Entrez, Source | myosin IE | 12726 | -0.209 | -0.3994 | Yes |
| 112 | CFL1 | CFL1 Entrez, Source | cofilin 1 (non-muscle) | 12779 | -0.218 | -0.3912 | Yes |
| 113 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 12828 | -0.232 | -0.3820 | Yes |
| 114 | MYADM | MYADM Entrez, Source | myeloid-associated differentiation marker | 12919 | -0.258 | -0.3745 | Yes |
| 115 | RAB27A | RAB27A Entrez, Source | RAB27A, member RAS oncogene family | 12927 | -0.261 | -0.3606 | Yes |
| 116 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 13035 | -0.310 | -0.3515 | Yes |
| 117 | FST | FST Entrez, Source | follistatin | 13045 | -0.318 | -0.3346 | Yes |
| 118 | CNN1 | CNN1 Entrez, Source | calponin 1, basic, smooth muscle | 13171 | -0.429 | -0.3203 | Yes |
| 119 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 13173 | -0.431 | -0.2965 | Yes |
| 120 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 13211 | -0.481 | -0.2726 | Yes |
| 121 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 13242 | -0.532 | -0.2455 | Yes |
| 122 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 13279 | -0.655 | -0.2119 | Yes |
| 123 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 13312 | -0.847 | -0.1674 | Yes |
| 124 | TAGLN | TAGLN Entrez, Source | transgelin | 13339 | -1.475 | -0.0877 | Yes |
| 125 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 13340 | -1.586 | 0.0002 | Yes |