Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
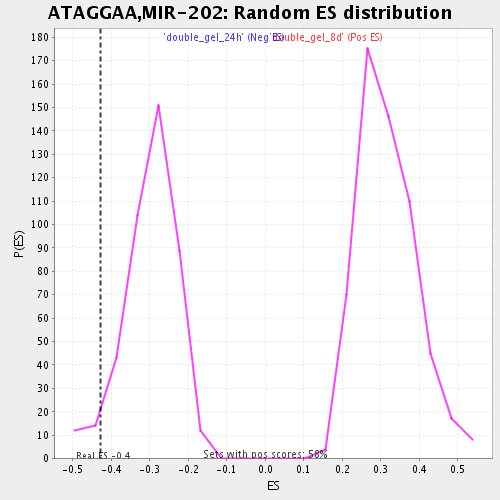
| GeneSet | ATAGGAA,MIR-202 |
| Enrichment Score (ES) | -0.4273553 |
| Normalized Enrichment Score (NES) | -1.4325495 |
| Nominal p-value | 0.049411766 |
| FDR q-value | 0.26552963 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BICD2 | BICD2 Entrez, Source | bicaudal D homolog 2 (Drosophila) | 1794 | 0.117 | -0.1107 | No |
| 2 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 2048 | 0.106 | -0.1080 | No |
| 3 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 2116 | 0.103 | -0.0917 | No |
| 4 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 2584 | 0.084 | -0.1095 | No |
| 5 | KHDRBS2 | KHDRBS2 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 2 | 2589 | 0.084 | -0.0925 | No |
| 6 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 2595 | 0.084 | -0.0756 | No |
| 7 | ROBO2 | ROBO2 Entrez, Source | roundabout, axon guidance receptor, homolog 2 (Drosophila) | 2881 | 0.075 | -0.0816 | No |
| 8 | SNAP91 | SNAP91 Entrez, Source | synaptosomal-associated protein, 91kDa homolog (mouse) | 3006 | 0.071 | -0.0762 | No |
| 9 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 3329 | 0.062 | -0.0877 | No |
| 10 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 3399 | 0.060 | -0.0804 | No |
| 11 | EI24 | EI24 Entrez, Source | etoposide induced 2.4 mRNA | 3410 | 0.060 | -0.0687 | No |
| 12 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 3460 | 0.059 | -0.0603 | No |
| 13 | ENAH | ENAH Entrez, Source | enabled homolog (Drosophila) | 3523 | 0.057 | -0.0532 | No |
| 14 | BAG4 | BAG4 Entrez, Source | BCL2-associated athanogene 4 | 4100 | 0.042 | -0.0878 | No |
| 15 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 4187 | 0.040 | -0.0860 | No |
| 16 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 4288 | 0.038 | -0.0857 | No |
| 17 | ARSB | ARSB Entrez, Source | arylsulfatase B | 4419 | 0.035 | -0.0882 | No |
| 18 | SNX16 | SNX16 Entrez, Source | sorting nexin 16 | 4434 | 0.035 | -0.0820 | No |
| 19 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 4845 | 0.027 | -0.1072 | No |
| 20 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 4920 | 0.026 | -0.1075 | No |
| 21 | STOX2 | STOX2 Entrez, Source | storkhead box 2 | 5167 | 0.021 | -0.1217 | No |
| 22 | SCN2B | SCN2B Entrez, Source | sodium channel, voltage-gated, type II, beta | 5632 | 0.012 | -0.1540 | No |
| 23 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 5997 | 0.005 | -0.1804 | No |
| 24 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 6711 | -0.007 | -0.2325 | No |
| 25 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 7401 | -0.020 | -0.2802 | No |
| 26 | KRAS | KRAS Entrez, Source | v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog | 7921 | -0.030 | -0.3131 | No |
| 27 | ZFAND3 | ZFAND3 Entrez, Source | zinc finger, AN1-type domain 3 | 7940 | -0.030 | -0.3083 | No |
| 28 | ATP2B4 | ATP2B4 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 4 | 9524 | -0.063 | -0.4142 | Yes |
| 29 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9594 | -0.065 | -0.4060 | Yes |
| 30 | APPBP2 | APPBP2 Entrez, Source | amyloid beta precursor protein (cytoplasmic tail) binding protein 2 | 9765 | -0.070 | -0.4044 | Yes |
| 31 | DNAJC10 | DNAJC10 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 10 | 9780 | -0.070 | -0.3909 | Yes |
| 32 | TTC13 | TTC13 Entrez, Source | tetratricopeptide repeat domain 13 | 9935 | -0.075 | -0.3871 | Yes |
| 33 | LBR | LBR Entrez, Source | lamin B receptor | 10137 | -0.080 | -0.3857 | Yes |
| 34 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 10508 | -0.090 | -0.3948 | Yes |
| 35 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 10664 | -0.096 | -0.3867 | Yes |
| 36 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 11041 | -0.109 | -0.3923 | Yes |
| 37 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 11078 | -0.111 | -0.3721 | Yes |
| 38 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 11114 | -0.113 | -0.3514 | Yes |
| 39 | TGFBR2 | TGFBR2 Entrez, Source | transforming growth factor, beta receptor II (70/80kDa) | 11338 | -0.122 | -0.3430 | Yes |
| 40 | HLF | HLF Entrez, Source | hepatic leukemia factor | 11411 | -0.125 | -0.3226 | Yes |
| 41 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 11435 | -0.126 | -0.2982 | Yes |
| 42 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 11550 | -0.131 | -0.2796 | Yes |
| 43 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 11793 | -0.144 | -0.2681 | Yes |
| 44 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 11925 | -0.152 | -0.2466 | Yes |
| 45 | USP15 | USP15 Entrez, Source | ubiquitin specific peptidase 15 | 12366 | -0.185 | -0.2414 | Yes |
| 46 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12761 | -0.235 | -0.2226 | Yes |
| 47 | CD28 | CD28 Entrez, Source | CD28 molecule | 12792 | -0.240 | -0.1753 | Yes |
| 48 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 12877 | -0.261 | -0.1277 | Yes |
| 49 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 12901 | -0.267 | -0.0743 | Yes |
| 50 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 13268 | -0.519 | 0.0056 | Yes |