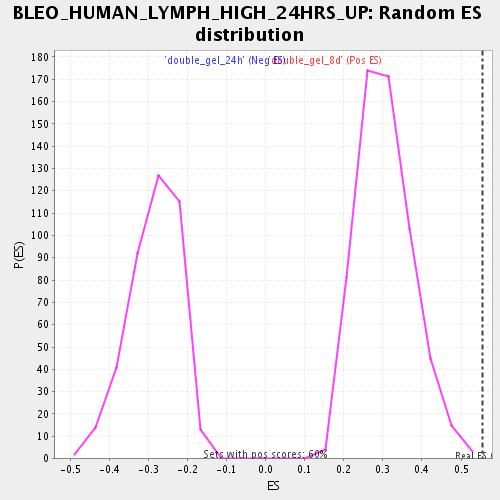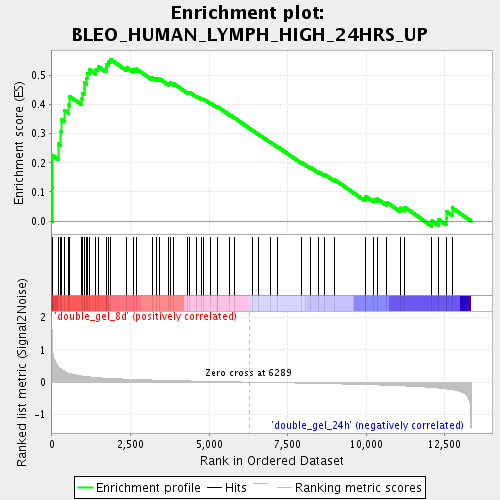
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | BLEO_HUMAN_LYMPH_HIGH_24HRS_UP |
| Enrichment Score (ES) | 0.5559734 |
| Normalized Enrichment Score (NES) | 1.8214296 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.023989206 |
| FWER p-Value | 0.882 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 21 | 1.039 | 0.1157 | Yes |
| 2 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 27 | 0.963 | 0.2242 | Yes |
| 3 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 205 | 0.482 | 0.2653 | Yes |
| 4 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 279 | 0.415 | 0.3067 | Yes |
| 5 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 315 | 0.388 | 0.3478 | Yes |
| 6 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 405 | 0.336 | 0.3791 | Yes |
| 7 | CD40 | CD40 Entrez, Source | CD40 molecule, TNF receptor superfamily member 5 | 539 | 0.273 | 0.3999 | Yes |
| 8 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 565 | 0.265 | 0.4280 | Yes |
| 9 | CRYZ | CRYZ Entrez, Source | crystallin, zeta (quinone reductase) | 951 | 0.188 | 0.4202 | Yes |
| 10 | LPXN | LPXN Entrez, Source | leupaxin | 979 | 0.185 | 0.4391 | Yes |
| 11 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 1025 | 0.179 | 0.4559 | Yes |
| 12 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 1031 | 0.178 | 0.4757 | Yes |
| 13 | ANXA4 | ANXA4 Entrez, Source | annexin A4 | 1109 | 0.170 | 0.4891 | Yes |
| 14 | PKIG | PKIG Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor gamma | 1121 | 0.169 | 0.5073 | Yes |
| 15 | SH3BP5 | SH3BP5 Entrez, Source | SH3-domain binding protein 5 (BTK-associated) | 1184 | 0.162 | 0.5210 | Yes |
| 16 | CTSH | CTSH Entrez, Source | cathepsin H | 1401 | 0.143 | 0.5209 | Yes |
| 17 | PLD3 | PLD3 Entrez, Source | phospholipase D family, member 3 | 1469 | 0.137 | 0.5313 | Yes |
| 18 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 1723 | 0.121 | 0.5260 | Yes |
| 19 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 1730 | 0.121 | 0.5392 | Yes |
| 20 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 1801 | 0.117 | 0.5472 | Yes |
| 21 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 1856 | 0.114 | 0.5560 | Yes |
| 22 | SMTN | SMTN Entrez, Source | smoothelin | 2386 | 0.092 | 0.5266 | No |
| 23 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 2595 | 0.084 | 0.5204 | No |
| 24 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 2686 | 0.081 | 0.5227 | No |
| 25 | PRKAB1 | PRKAB1 Entrez, Source | protein kinase, AMP-activated, beta 1 non-catalytic subunit | 3189 | 0.066 | 0.4924 | No |
| 26 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 3329 | 0.062 | 0.4889 | No |
| 27 | EI24 | EI24 Entrez, Source | etoposide induced 2.4 mRNA | 3410 | 0.060 | 0.4897 | No |
| 28 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 3715 | 0.052 | 0.4726 | No |
| 29 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 3759 | 0.051 | 0.4752 | No |
| 30 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 3872 | 0.048 | 0.4722 | No |
| 31 | CAPN1 | CAPN1 Entrez, Source | calpain 1, (mu/I) large subunit | 4315 | 0.037 | 0.4431 | No |
| 32 | NFKB2 | NFKB2 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | 4395 | 0.036 | 0.4412 | No |
| 33 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 4614 | 0.032 | 0.4284 | No |
| 34 | PRKAR1A | PRKAR1A Entrez, Source | protein kinase, cAMP-dependent, regulatory, type I, alpha (tissue specific extinguisher 1) | 4750 | 0.029 | 0.4215 | No |
| 35 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 4827 | 0.027 | 0.4189 | No |
| 36 | RPS6KA1 | RPS6KA1 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 1 | 5059 | 0.023 | 0.4041 | No |
| 37 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 5264 | 0.019 | 0.3909 | No |
| 38 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 5270 | 0.019 | 0.3927 | No |
| 39 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 5658 | 0.012 | 0.3649 | No |
| 40 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 5827 | 0.009 | 0.3532 | No |
| 41 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 6393 | -0.002 | 0.3109 | No |
| 42 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 6591 | -0.005 | 0.2966 | No |
| 43 | PLEK | PLEK Entrez, Source | pleckstrin | 6970 | -0.012 | 0.2695 | No |
| 44 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 7183 | -0.015 | 0.2553 | No |
| 45 | MRPL49 | MRPL49 Entrez, Source | mitochondrial ribosomal protein L49 | 7943 | -0.030 | 0.2015 | No |
| 46 | LTA | LTA Entrez, Source | lymphotoxin alpha (TNF superfamily, member 1) | 8246 | -0.036 | 0.1829 | No |
| 47 | TNFSF9 | TNFSF9 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 9 | 8499 | -0.041 | 0.1686 | No |
| 48 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8685 | -0.045 | 0.1598 | No |
| 49 | WDR1 | WDR1 Entrez, Source | WD repeat domain 1 | 8991 | -0.052 | 0.1427 | No |
| 50 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 9971 | -0.076 | 0.0775 | No |
| 51 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 9991 | -0.076 | 0.0847 | No |
| 52 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 10248 | -0.083 | 0.0748 | No |
| 53 | PLXNB2 | PLXNB2 Entrez, Source | plexin B2 | 10361 | -0.086 | 0.0761 | No |
| 54 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 10664 | -0.096 | 0.0642 | No |
| 55 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 11087 | -0.111 | 0.0450 | No |
| 56 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 11229 | -0.117 | 0.0476 | No |
| 57 | GPX1 | GPX1 Entrez, Source | glutathione peroxidase 1 | 12098 | -0.164 | 0.0008 | No |
| 58 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12298 | -0.179 | 0.0061 | No |
| 59 | MYO5A | MYO5A Entrez, Source | myosin VA (heavy chain 12, myoxin) | 12548 | -0.204 | 0.0104 | No |
| 60 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 12556 | -0.205 | 0.0331 | No |
| 61 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 12738 | -0.231 | 0.0455 | No |