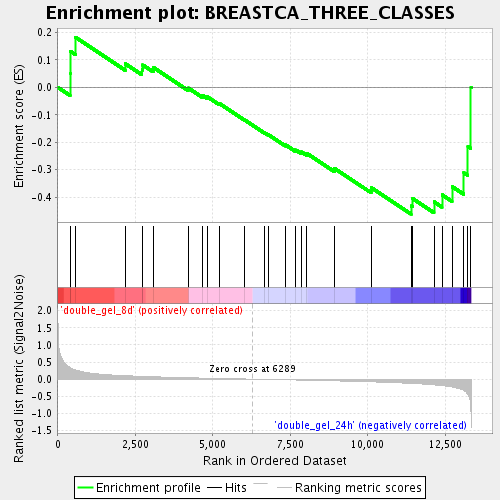
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | BREASTCA_THREE_CLASSES |
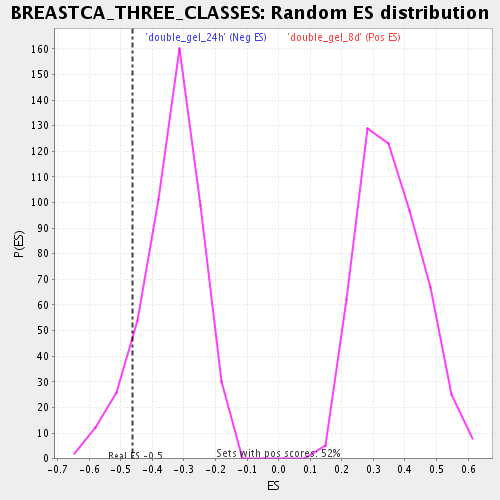
| Enrichment Score (ES) | -0.46216497 |
| Normalized Enrichment Score (NES) | -1.3584619 |
| Nominal p-value | 0.10123967 |
| FDR q-value | 0.3482962 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RBL2 | RBL2 Entrez, Source | retinoblastoma-like 2 (p130) | 394 | 0.340 | 0.0517 | No |
| 2 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 401 | 0.337 | 0.1318 | No |
| 3 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 566 | 0.265 | 0.1828 | No |
| 4 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2173 | 0.101 | 0.0863 | No |
| 5 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 2713 | 0.080 | 0.0648 | No |
| 6 | COX6C | COX6C Entrez, Source | cytochrome c oxidase subunit VIc | 2739 | 0.079 | 0.0818 | No |
| 7 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 3080 | 0.069 | 0.0727 | No |
| 8 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 4204 | 0.039 | -0.0022 | No |
| 9 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 4656 | 0.031 | -0.0286 | No |
| 10 | NT5DC2 | NT5DC2 Entrez, Source | 5'-nucleotidase domain containing 2 | 4819 | 0.028 | -0.0342 | No |
| 11 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5218 | 0.020 | -0.0593 | No |
| 12 | SLC35B1 | SLC35B1 Entrez, Source | solute carrier family 35, member B1 | 6036 | 0.004 | -0.1196 | No |
| 13 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 6679 | -0.007 | -0.1662 | No |
| 14 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6785 | -0.009 | -0.1720 | No |
| 15 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 7346 | -0.019 | -0.2096 | No |
| 16 | GDI2 | GDI2 Entrez, Source | GDP dissociation inhibitor 2 | 7674 | -0.025 | -0.2282 | No |
| 17 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 7860 | -0.028 | -0.2353 | No |
| 18 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 8028 | -0.032 | -0.2402 | No |
| 19 | C10ORF7 | C10ORF7 Entrez, Source | chromosome 10 open reading frame 7 | 8926 | -0.050 | -0.2956 | No |
| 20 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10112 | -0.079 | -0.3656 | No |
| 21 | SERBP1 | SERBP1 Entrez, Source | SERPINE1 mRNA binding protein 1 | 11399 | -0.125 | -0.4323 | Yes |
| 22 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | -0.4056 | Yes |
| 23 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 12139 | -0.167 | -0.4174 | Yes |
| 24 | ST13 | ST13 Entrez, Source | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | 12397 | -0.188 | -0.3918 | Yes |
| 25 | EIF2C2 | EIF2C2 Entrez, Source | eukaryotic translation initiation factor 2C, 2 | 12721 | -0.227 | -0.3617 | Yes |
| 26 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13101 | -0.328 | -0.3118 | Yes |
| 27 | VLDLR | VLDLR Entrez, Source | very low density lipoprotein receptor | 13233 | -0.442 | -0.2159 | Yes |
| 28 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 13331 | -0.937 | 0.0008 | Yes |