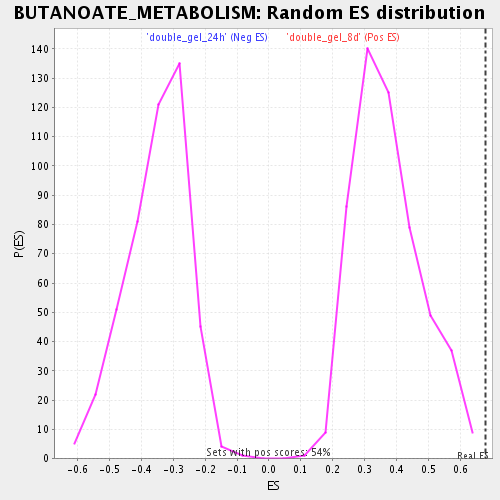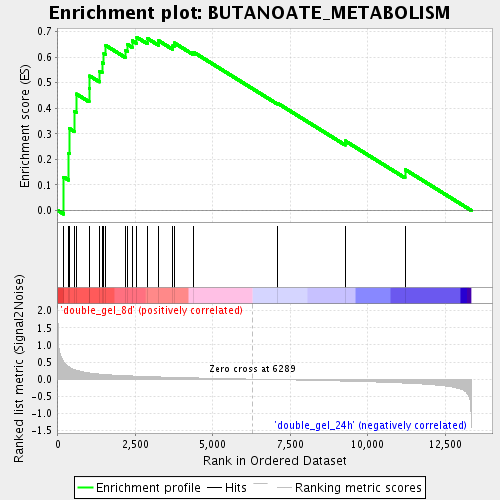
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | BUTANOATE_METABOLISM |
| Enrichment Score (ES) | 0.67789793 |
| Normalized Enrichment Score (NES) | 1.8164523 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.024780259 |
| FWER p-Value | 0.898 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.1311 | Yes |
| 2 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 346 | 0.367 | 0.2235 | Yes |
| 3 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 375 | 0.349 | 0.3205 | Yes |
| 4 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 526 | 0.277 | 0.3879 | Yes |
| 5 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 595 | 0.258 | 0.4561 | Yes |
| 6 | SDS | SDS Entrez, Source | serine dehydratase | 1007 | 0.182 | 0.4770 | Yes |
| 7 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 1026 | 0.179 | 0.5265 | Yes |
| 8 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 1343 | 0.147 | 0.5447 | Yes |
| 9 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 1428 | 0.140 | 0.5782 | Yes |
| 10 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 1482 | 0.136 | 0.6131 | Yes |
| 11 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1544 | 0.132 | 0.6459 | Yes |
| 12 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 2173 | 0.101 | 0.6273 | Yes |
| 13 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 2249 | 0.097 | 0.6494 | Yes |
| 14 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 2398 | 0.092 | 0.6644 | Yes |
| 15 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 2544 | 0.086 | 0.6779 | Yes |
| 16 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 2893 | 0.074 | 0.6729 | No |
| 17 | AACS | AACS Entrez, Source | acetoacetyl-CoA synthetase | 3252 | 0.064 | 0.6642 | No |
| 18 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 3710 | 0.052 | 0.6447 | No |
| 19 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 3755 | 0.051 | 0.6559 | No |
| 20 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 4375 | 0.036 | 0.6197 | No |
| 21 | HMGCL | HMGCL Entrez, Source | 3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria) | 7083 | -0.014 | 0.4204 | No |
| 22 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 9270 | -0.057 | 0.2726 | No |
| 23 | GAD2 | GAD2 Entrez, Source | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | 11204 | -0.116 | 0.1605 | No |