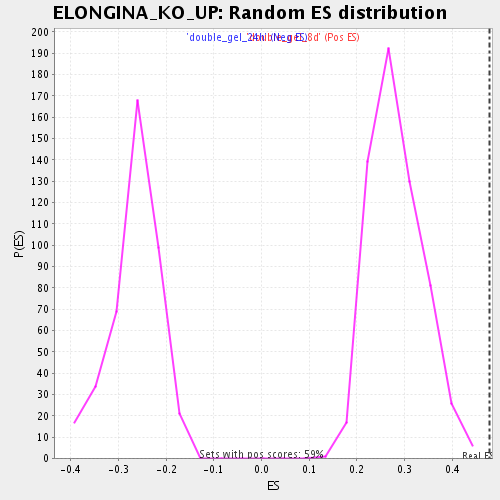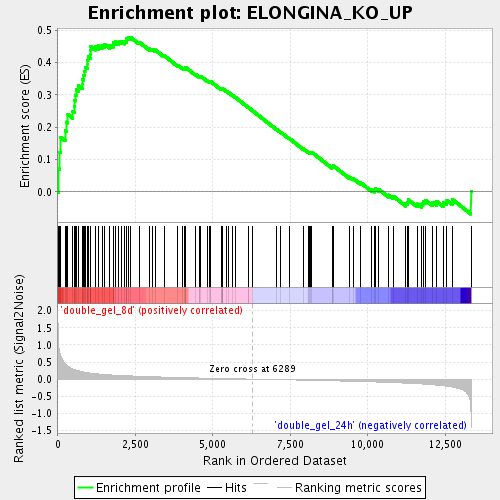
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ELONGINA_KO_UP |
| Enrichment Score (ES) | 0.4785084 |
| Normalized Enrichment Score (NES) | 1.7007505 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.054826222 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 21 | 1.039 | 0.0719 | Yes |
| 2 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 62 | 0.756 | 0.1224 | Yes |
| 3 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 89 | 0.680 | 0.1685 | Yes |
| 4 | SERPINH1 | SERPINH1 Entrez, Source | serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | 233 | 0.457 | 0.1900 | Yes |
| 5 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 276 | 0.416 | 0.2163 | Yes |
| 6 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 321 | 0.383 | 0.2400 | Yes |
| 7 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 480 | 0.292 | 0.2488 | Yes |
| 8 | LAMC1 | LAMC1 Entrez, Source | laminin, gamma 1 (formerly LAMB2) | 530 | 0.275 | 0.2646 | Yes |
| 9 | MFGE8 | MFGE8 Entrez, Source | milk fat globule-EGF factor 8 protein | 542 | 0.272 | 0.2830 | Yes |
| 10 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 567 | 0.265 | 0.2999 | Yes |
| 11 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 597 | 0.258 | 0.3159 | Yes |
| 12 | ENPEP | ENPEP Entrez, Source | glutamyl aminopeptidase (aminopeptidase A) | 658 | 0.244 | 0.3286 | Yes |
| 13 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 793 | 0.215 | 0.3338 | Yes |
| 14 | TTR | TTR Entrez, Source | transthyretin (prealbumin, amyloidosis type I) | 798 | 0.213 | 0.3486 | Yes |
| 15 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 834 | 0.207 | 0.3606 | Yes |
| 16 | EFEMP2 | EFEMP2 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 2 | 852 | 0.204 | 0.3737 | Yes |
| 17 | THBD | THBD Entrez, Source | thrombomodulin | 877 | 0.198 | 0.3860 | Yes |
| 18 | STARD10 | STARD10 Entrez, Source | START domain containing 10 | 953 | 0.188 | 0.3936 | Yes |
| 19 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 956 | 0.188 | 0.4067 | Yes |
| 20 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 973 | 0.186 | 0.4187 | Yes |
| 21 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 1052 | 0.177 | 0.4253 | Yes |
| 22 | GHR | GHR Entrez, Source | growth hormone receptor | 1057 | 0.176 | 0.4374 | Yes |
| 23 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.4495 | Yes |
| 24 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 1203 | 0.161 | 0.4503 | Yes |
| 25 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 1320 | 0.149 | 0.4520 | Yes |
| 26 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1445 | 0.139 | 0.4525 | Yes |
| 27 | CST3 | CST3 Entrez, Source | cystatin C (amyloid angiopathy and cerebral hemorrhage) | 1513 | 0.134 | 0.4570 | Yes |
| 28 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1676 | 0.124 | 0.4535 | Yes |
| 29 | VAMP8 | VAMP8 Entrez, Source | vesicle-associated membrane protein 8 (endobrevin) | 1782 | 0.118 | 0.4539 | Yes |
| 30 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1785 | 0.118 | 0.4621 | Yes |
| 31 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 1861 | 0.114 | 0.4644 | Yes |
| 32 | TST | TST Entrez, Source | thiosulfate sulfurtransferase (rhodanese) | 1969 | 0.109 | 0.4641 | Yes |
| 33 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 2048 | 0.106 | 0.4657 | Yes |
| 34 | KDELR2 | KDELR2 Entrez, Source | KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 2 | 2157 | 0.101 | 0.4647 | Yes |
| 35 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 2206 | 0.099 | 0.4680 | Yes |
| 36 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 2220 | 0.099 | 0.4740 | Yes |
| 37 | SGCE | SGCE Entrez, Source | sarcoglycan, epsilon | 2274 | 0.096 | 0.4769 | Yes |
| 38 | SNAI1 | SNAI1 Entrez, Source | snail homolog 1 (Drosophila) | 2341 | 0.094 | 0.4785 | Yes |
| 39 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 2618 | 0.083 | 0.4635 | No |
| 40 | ARF4 | ARF4 Entrez, Source | ADP-ribosylation factor 4 | 2963 | 0.072 | 0.4427 | No |
| 41 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 3039 | 0.070 | 0.4420 | No |
| 42 | SPINK1 | SPINK1 Entrez, Source | serine peptidase inhibitor, Kazal type 1 | 3138 | 0.068 | 0.4393 | No |
| 43 | CYBASC3 | CYBASC3 Entrez, Source | cytochrome b, ascorbate dependent 3 | 3424 | 0.060 | 0.4220 | No |
| 44 | PGA5 | PGA5 Entrez, Source | pepsinogen 5, group I (pepsinogen A) | 3874 | 0.048 | 0.3915 | No |
| 45 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 4025 | 0.044 | 0.3833 | No |
| 46 | MYL3 | MYL3 Entrez, Source | myosin, light chain 3, alkali; ventricular, skeletal, slow | 4081 | 0.043 | 0.3822 | No |
| 47 | FBLN2 | FBLN2 Entrez, Source | fibulin 2 | 4117 | 0.042 | 0.3825 | No |
| 48 | MESDC1 | MESDC1 Entrez, Source | mesoderm development candidate 1 | 4119 | 0.042 | 0.3854 | No |
| 49 | CD38 | CD38 Entrez, Source | CD38 molecule | 4439 | 0.035 | 0.3638 | No |
| 50 | KIT | KIT Entrez, Source | v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog | 4575 | 0.033 | 0.3559 | No |
| 51 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 4578 | 0.032 | 0.3580 | No |
| 52 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 4614 | 0.032 | 0.3576 | No |
| 53 | RAMP2 | RAMP2 Entrez, Source | receptor (calcitonin) activity modifying protein 2 | 4826 | 0.027 | 0.3436 | No |
| 54 | NID1 | NID1 Entrez, Source | nidogen 1 | 4891 | 0.026 | 0.3407 | No |
| 55 | CCNC | CCNC Entrez, Source | cyclin C | 4913 | 0.026 | 0.3409 | No |
| 56 | LGMN | LGMN Entrez, Source | legumain | 4915 | 0.026 | 0.3426 | No |
| 57 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 5264 | 0.019 | 0.3177 | No |
| 58 | JUND | JUND Entrez, Source | jun D proto-oncogene | 5289 | 0.019 | 0.3173 | No |
| 59 | PDE1B | PDE1B Entrez, Source | phosphodiesterase 1B, calmodulin-dependent | 5303 | 0.019 | 0.3176 | No |
| 60 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 5306 | 0.019 | 0.3188 | No |
| 61 | STRA6 | STRA6 Entrez, Source | stimulated by retinoic acid gene 6 homolog (mouse) | 5317 | 0.019 | 0.3193 | No |
| 62 | CLU | CLU Entrez, Source | clusterin | 5443 | 0.016 | 0.3110 | No |
| 63 | SURF4 | SURF4 Entrez, Source | surfeit 4 | 5514 | 0.014 | 0.3067 | No |
| 64 | SPOCK1 | SPOCK1 Entrez, Source | sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 1 | 5637 | 0.012 | 0.2984 | No |
| 65 | LRPAP1 | LRPAP1 Entrez, Source | low density lipoprotein receptor-related protein associated protein 1 | 5727 | 0.010 | 0.2924 | No |
| 66 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 6136 | 0.003 | 0.2618 | No |
| 67 | CITED1 | CITED1 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 | 6164 | 0.002 | 0.2599 | No |
| 68 | PTHR1 | PTHR1 Entrez, Source | parathyroid hormone receptor 1 | 6271 | 0.000 | 0.2519 | No |
| 69 | B2M | B2M Entrez, Source | beta-2-microglobulin | 7066 | -0.014 | 0.1929 | No |
| 70 | CREB3L1 | CREB3L1 Entrez, Source | cAMP responsive element binding protein 3-like 1 | 7184 | -0.016 | 0.1851 | No |
| 71 | GGTA1 | GGTA1 Entrez, Source | glycoprotein, alpha-galactosyltransferase 1 | 7462 | -0.021 | 0.1657 | No |
| 72 | FKBP9 | FKBP9 Entrez, Source | FK506 binding protein 9, 63 kDa | 7913 | -0.029 | 0.1338 | No |
| 73 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 8088 | -0.033 | 0.1230 | No |
| 74 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 8134 | -0.034 | 0.1220 | No |
| 75 | HSDL1 | HSDL1 Entrez, Source | hydroxysteroid dehydrogenase like 1 | 8151 | -0.034 | 0.1232 | No |
| 76 | ATP6AP1 | ATP6AP1 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 1 | 8196 | -0.035 | 0.1224 | No |
| 77 | TXNDC4 | TXNDC4 Entrez, Source | thioredoxin domain containing 4 (endoplasmic reticulum) | 8846 | -0.049 | 0.0768 | No |
| 78 | AQP8 | AQP8 Entrez, Source | aquaporin 8 | 8862 | -0.049 | 0.0792 | No |
| 79 | GNA12 | GNA12 Entrez, Source | guanine nucleotide binding protein (G protein) alpha 12 | 8878 | -0.049 | 0.0816 | No |
| 80 | TMED2 | TMED2 Entrez, Source | transmembrane emp24 domain trafficking protein 2 | 9403 | -0.061 | 0.0463 | No |
| 81 | GRINA | GRINA Entrez, Source | glutamate receptor, ionotropic, N-methyl D-asparate-associated protein 1 (glutamate binding) | 9530 | -0.063 | 0.0413 | No |
| 82 | PSPH | PSPH Entrez, Source | phosphoserine phosphatase | 9762 | -0.069 | 0.0287 | No |
| 83 | P4HB | P4HB Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), beta polypeptide | 10130 | -0.080 | 0.0067 | No |
| 84 | LMBRD1 | LMBRD1 Entrez, Source | LMBR1 domain containing 1 | 10226 | -0.082 | 0.0053 | No |
| 85 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 10239 | -0.083 | 0.0103 | No |
| 86 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 10356 | -0.086 | 0.0076 | No |
| 87 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 10674 | -0.096 | -0.0096 | No |
| 88 | ZYX | ZYX Entrez, Source | zyxin | 10834 | -0.102 | -0.0143 | No |
| 89 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 11229 | -0.117 | -0.0358 | No |
| 90 | CTSZ | CTSZ Entrez, Source | cathepsin Z | 11291 | -0.120 | -0.0319 | No |
| 91 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 11305 | -0.121 | -0.0244 | No |
| 92 | SRPRB | SRPRB Entrez, Source | signal recognition particle receptor, B subunit | 11601 | -0.133 | -0.0372 | No |
| 93 | P4HA1 | P4HA1 Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide I | 11736 | -0.140 | -0.0374 | No |
| 94 | FDX1 | FDX1 Entrez, Source | ferredoxin 1 | 11784 | -0.144 | -0.0308 | No |
| 95 | PDIA3 | PDIA3 Entrez, Source | protein disulfide isomerase family A, member 3 | 11852 | -0.147 | -0.0254 | No |
| 96 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 12088 | -0.163 | -0.0316 | No |
| 97 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 12221 | -0.173 | -0.0293 | No |
| 98 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 12434 | -0.191 | -0.0318 | No |
| 99 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 12549 | -0.204 | -0.0260 | No |
| 100 | SLC16A1 | SLC16A1 Entrez, Source | solute carrier family 16, member 1 (monocarboxylic acid transporter 1) | 12730 | -0.229 | -0.0233 | No |
| 101 | CYP26A1 | CYP26A1 Entrez, Source | cytochrome P450, family 26, subfamily A, polypeptide 1 | 13332 | -0.982 | 0.0008 | No |