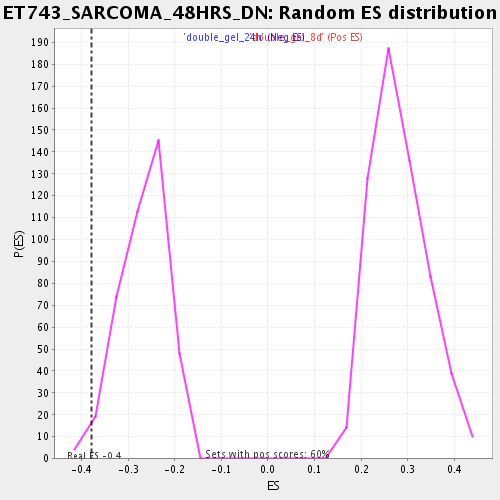
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | ET743_SARCOMA_48HRS_DN |
| Enrichment Score (ES) | -0.37834287 |
| Normalized Enrichment Score (NES) | -1.4181098 |
| Nominal p-value | 0.014888338 |
| FDR q-value | 0.27673417 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 37 | 0.888 | 0.0824 | No |
| 2 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 433 | 0.319 | 0.0832 | No |
| 3 | CYHR1 | CYHR1 Entrez, Source | cysteine/histidine-rich 1 | 498 | 0.287 | 0.1059 | No |
| 4 | FZD1 | FZD1 Entrez, Source | frizzled homolog 1 (Drosophila) | 708 | 0.233 | 0.1125 | No |
| 5 | RNF14 | RNF14 Entrez, Source | ring finger protein 14 | 836 | 0.207 | 0.1228 | No |
| 6 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1115 | 0.169 | 0.1180 | No |
| 7 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 1410 | 0.142 | 0.1094 | No |
| 8 | RAP1B | RAP1B Entrez, Source | RAP1B, member of RAS oncogene family | 2132 | 0.102 | 0.0649 | No |
| 9 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 2686 | 0.081 | 0.0309 | No |
| 10 | MCTS1 | MCTS1 Entrez, Source | malignant T cell amplified sequence 1 | 3084 | 0.069 | 0.0075 | No |
| 11 | MAPK14 | MAPK14 Entrez, Source | mitogen-activated protein kinase 14 | 3602 | 0.055 | -0.0263 | No |
| 12 | CDH11 | CDH11 Entrez, Source | cadherin 11, type 2, OB-cadherin (osteoblast) | 3983 | 0.045 | -0.0506 | No |
| 13 | TMPO | TMPO Entrez, Source | thymopoietin | 4229 | 0.039 | -0.0654 | No |
| 14 | HERC4 | HERC4 Entrez, Source | hect domain and RLD 4 | 4484 | 0.034 | -0.0813 | No |
| 15 | WNT5B | WNT5B Entrez, Source | wingless-type MMTV integration site family, member 5B | 4602 | 0.032 | -0.0870 | No |
| 16 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 4996 | 0.024 | -0.1144 | No |
| 17 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 5186 | 0.020 | -0.1267 | No |
| 18 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 5378 | 0.017 | -0.1395 | No |
| 19 | MID1IP1 | MID1IP1 Entrez, Source | MID1 interacting protein 1 (gastrulation specific G12 homolog (zebrafish)) | 5440 | 0.016 | -0.1425 | No |
| 20 | SIRPA | SIRPA Entrez, Source | signal-regulatory protein alpha | 5901 | 0.007 | -0.1766 | No |
| 21 | CXXC5 | CXXC5 Entrez, Source | CXXC finger 5 | 5992 | 0.005 | -0.1828 | No |
| 22 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 6034 | 0.005 | -0.1855 | No |
| 23 | CDC5L | CDC5L Entrez, Source | CDC5 cell division cycle 5-like (S. pombe) | 6411 | -0.002 | -0.2137 | No |
| 24 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 6729 | -0.008 | -0.2369 | No |
| 25 | PSMB1 | PSMB1 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 1 | 6762 | -0.008 | -0.2385 | No |
| 26 | GTF2B | GTF2B Entrez, Source | general transcription factor IIB | 6968 | -0.012 | -0.2528 | No |
| 27 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 7017 | -0.013 | -0.2553 | No |
| 28 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 7106 | -0.014 | -0.2606 | No |
| 29 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 7273 | -0.017 | -0.2714 | No |
| 30 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 7458 | -0.021 | -0.2833 | No |
| 31 | BCL2L1 | BCL2L1 Entrez, Source | BCL2-like 1 | 7544 | -0.022 | -0.2876 | No |
| 32 | EIF3S10 | EIF3S10 Entrez, Source | eukaryotic translation initiation factor 3, subunit 10 theta, 150/170kDa | 7771 | -0.027 | -0.3021 | No |
| 33 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 7813 | -0.028 | -0.3025 | No |
| 34 | BIRC2 | BIRC2 Entrez, Source | baculoviral IAP repeat-containing 2 | 7883 | -0.029 | -0.3049 | No |
| 35 | CDC42BPA | CDC42BPA Entrez, Source | CDC42 binding protein kinase alpha (DMPK-like) | 8008 | -0.031 | -0.3113 | No |
| 36 | DNAJA2 | DNAJA2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 2 | 8188 | -0.035 | -0.3214 | No |
| 37 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 8227 | -0.036 | -0.3209 | No |
| 38 | FNTA | FNTA Entrez, Source | farnesyltransferase, CAAX box, alpha | 8270 | -0.037 | -0.3205 | No |
| 39 | NBN | NBN Entrez, Source | nibrin | 8672 | -0.045 | -0.3464 | No |
| 40 | PJA2 | PJA2 Entrez, Source | praja 2, RING-H2 motif containing | 8736 | -0.046 | -0.3468 | No |
| 41 | CRLF3 | CRLF3 Entrez, Source | cytokine receptor-like factor 3 | 8904 | -0.050 | -0.3546 | No |
| 42 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 9220 | -0.056 | -0.3729 | Yes |
| 43 | SON | SON Entrez, Source | SON DNA binding protein | 9229 | -0.057 | -0.3681 | Yes |
| 44 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 9290 | -0.058 | -0.3671 | Yes |
| 45 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 9296 | -0.058 | -0.3619 | Yes |
| 46 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 9349 | -0.059 | -0.3601 | Yes |
| 47 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 9365 | -0.060 | -0.3555 | Yes |
| 48 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 9449 | -0.062 | -0.3559 | Yes |
| 49 | SFRS9 | SFRS9 Entrez, Source | splicing factor, arginine/serine-rich 9 | 9480 | -0.062 | -0.3522 | Yes |
| 50 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 9489 | -0.063 | -0.3468 | Yes |
| 51 | PSMC2 | PSMC2 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 2 | 9566 | -0.064 | -0.3463 | Yes |
| 52 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 9615 | -0.066 | -0.3436 | Yes |
| 53 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 9619 | -0.066 | -0.3375 | Yes |
| 54 | USP3 | USP3 Entrez, Source | ubiquitin specific peptidase 3 | 9792 | -0.070 | -0.3438 | Yes |
| 55 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 9797 | -0.071 | -0.3373 | Yes |
| 56 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 9987 | -0.076 | -0.3442 | Yes |
| 57 | CLCN7 | CLCN7 Entrez, Source | chloride channel 7 | 10076 | -0.079 | -0.3433 | Yes |
| 58 | RAB2 | RAB2 Entrez, Source | RAB2, member RAS oncogene family | 10106 | -0.079 | -0.3379 | Yes |
| 59 | LBR | LBR Entrez, Source | lamin B receptor | 10137 | -0.080 | -0.3325 | Yes |
| 60 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 10351 | -0.086 | -0.3403 | Yes |
| 61 | ATF1 | ATF1 Entrez, Source | activating transcription factor 1 | 10381 | -0.087 | -0.3342 | Yes |
| 62 | NCOA6 | NCOA6 Entrez, Source | nuclear receptor coactivator 6 | 10541 | -0.091 | -0.3374 | Yes |
| 63 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 10574 | -0.093 | -0.3309 | Yes |
| 64 | CRSP6 | CRSP6 Entrez, Source | cofactor required for Sp1 transcriptional activation, subunit 6, 77kDa | 10881 | -0.104 | -0.3441 | Yes |
| 65 | CCNH | CCNH Entrez, Source | cyclin H | 10890 | -0.105 | -0.3346 | Yes |
| 66 | PSMA5 | PSMA5 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 5 | 11084 | -0.111 | -0.3385 | Yes |
| 67 | RABGGTB | RABGGTB Entrez, Source | Rab geranylgeranyltransferase, beta subunit | 11200 | -0.116 | -0.3360 | Yes |
| 68 | RBM17 | RBM17 Entrez, Source | RNA binding motif protein 17 | 11216 | -0.117 | -0.3260 | Yes |
| 69 | SNIP1 | SNIP1 Entrez, Source | Smad nuclear interacting protein 1 | 11379 | -0.124 | -0.3263 | Yes |
| 70 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 11393 | -0.124 | -0.3154 | Yes |
| 71 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 11410 | -0.125 | -0.3046 | Yes |
| 72 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | -0.2953 | Yes |
| 73 | CCT4 | CCT4 Entrez, Source | chaperonin containing TCP1, subunit 4 (delta) | 11585 | -0.133 | -0.2927 | Yes |
| 74 | FBXL3 | FBXL3 Entrez, Source | F-box and leucine-rich repeat protein 3 | 11676 | -0.137 | -0.2864 | Yes |
| 75 | ARFGAP1 | ARFGAP1 Entrez, Source | ADP-ribosylation factor GTPase activating protein 1 | 11713 | -0.139 | -0.2757 | Yes |
| 76 | PSMC6 | PSMC6 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 6 | 11752 | -0.141 | -0.2651 | Yes |
| 77 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 11795 | -0.144 | -0.2544 | Yes |
| 78 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 11813 | -0.145 | -0.2417 | Yes |
| 79 | NUFIP1 | NUFIP1 Entrez, Source | nuclear fragile X mental retardation protein interacting protein 1 | 11965 | -0.155 | -0.2383 | Yes |
| 80 | DDX20 | DDX20 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 | 12161 | -0.169 | -0.2368 | Yes |
| 81 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 12178 | -0.170 | -0.2218 | Yes |
| 82 | PANX1 | PANX1 Entrez, Source | pannexin 1 | 12336 | -0.182 | -0.2161 | Yes |
| 83 | MTIF2 | MTIF2 Entrez, Source | mitochondrial translational initiation factor 2 | 12340 | -0.183 | -0.1988 | Yes |
| 84 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 12536 | -0.203 | -0.1941 | Yes |
| 85 | PSME3 | PSME3 Entrez, Source | proteasome (prosome, macropain) activator subunit 3 (PA28 gamma; Ki) | 12583 | -0.209 | -0.1775 | Yes |
| 86 | PRKCBP1 | PRKCBP1 Entrez, Source | protein kinase C binding protein 1 | 12589 | -0.209 | -0.1579 | Yes |
| 87 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 12599 | -0.210 | -0.1384 | Yes |
| 88 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12761 | -0.235 | -0.1280 | Yes |
| 89 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 12920 | -0.270 | -0.1140 | Yes |
| 90 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 12967 | -0.282 | -0.0904 | Yes |
| 91 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 13031 | -0.302 | -0.0662 | Yes |
| 92 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 13108 | -0.331 | -0.0401 | Yes |
| 93 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13299 | -0.601 | 0.0032 | Yes |