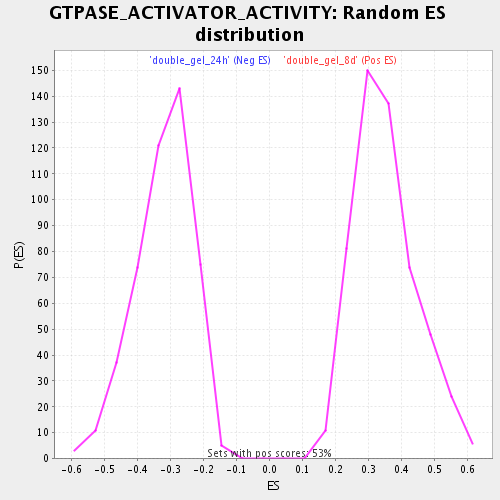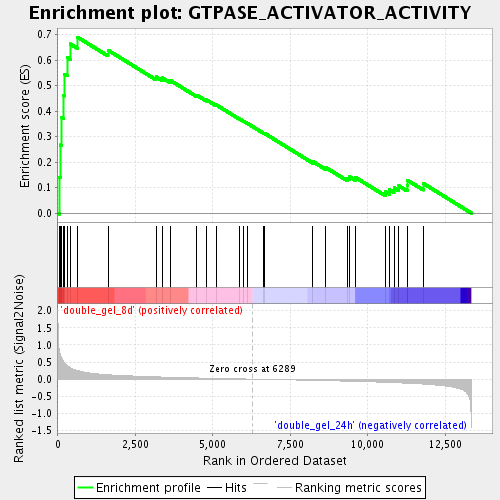
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | GTPASE_ACTIVATOR_ACTIVITY |
| Enrichment Score (ES) | 0.68948746 |
| Normalized Enrichment Score (NES) | 1.9654624 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.007301701 |
| FWER p-Value | 0.25 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 50 | 0.810 | 0.1411 | Yes |
| 2 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 73 | 0.715 | 0.2673 | Yes |
| 3 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 125 | 0.616 | 0.3738 | Yes |
| 4 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.4606 | Yes |
| 5 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 205 | 0.482 | 0.5454 | Yes |
| 6 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 301 | 0.402 | 0.6102 | Yes |
| 7 | CDC42EP2 | CDC42EP2 Entrez, Source | CDC42 effector protein (Rho GTPase binding) 2 | 399 | 0.337 | 0.6633 | Yes |
| 8 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 639 | 0.247 | 0.6895 | Yes |
| 9 | RALBP1 | RALBP1 Entrez, Source | ralA binding protein 1 | 1617 | 0.127 | 0.6388 | No |
| 10 | CENTA1 | CENTA1 Entrez, Source | centaurin, alpha 1 | 3167 | 0.067 | 0.5344 | No |
| 11 | RGS6 | RGS6 Entrez, Source | regulator of G-protein signalling 6 | 3368 | 0.061 | 0.5303 | No |
| 12 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 3633 | 0.054 | 0.5201 | No |
| 13 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 4478 | 0.034 | 0.4628 | No |
| 14 | ARHGAP4 | ARHGAP4 Entrez, Source | Rho GTPase activating protein 4 | 4793 | 0.028 | 0.4442 | No |
| 15 | RAP1GAP | RAP1GAP Entrez, Source | RAP1 GTPase activating protein | 5106 | 0.022 | 0.4247 | No |
| 16 | RGS9 | RGS9 Entrez, Source | regulator of G-protein signalling 9 | 5873 | 0.008 | 0.3686 | No |
| 17 | ARHGAP27 | ARHGAP27 Entrez, Source | Rho GTPase activating protein 27 | 6001 | 0.005 | 0.3599 | No |
| 18 | CENTD2 | CENTD2 Entrez, Source | centaurin, delta 2 | 6123 | 0.003 | 0.3514 | No |
| 19 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 6645 | -0.006 | 0.3134 | No |
| 20 | RGS12 | RGS12 Entrez, Source | regulator of G-protein signalling 12 | 6660 | -0.007 | 0.3135 | No |
| 21 | RGS14 | RGS14 Entrez, Source | regulator of G-protein signalling 14 | 8231 | -0.036 | 0.2020 | No |
| 22 | CHN2 | CHN2 Entrez, Source | chimerin (chimaerin) 2 | 8642 | -0.044 | 0.1791 | No |
| 23 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 9336 | -0.059 | 0.1376 | No |
| 24 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 9408 | -0.061 | 0.1431 | No |
| 25 | RAB3GAP2 | RAB3GAP2 Entrez, Source | RAB3 GTPase activating protein subunit 2 (non-catalytic) | 9593 | -0.065 | 0.1410 | No |
| 26 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 10565 | -0.092 | 0.0845 | No |
| 27 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 10704 | -0.097 | 0.0915 | No |
| 28 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 10848 | -0.103 | 0.0992 | No |
| 29 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11004 | -0.108 | 0.1069 | No |
| 30 | RASA3 | RASA3 Entrez, Source | RAS p21 protein activator 3 | 11272 | -0.119 | 0.1082 | No |
| 31 | MYO9B | MYO9B Entrez, Source | myosin IXB | 11288 | -0.120 | 0.1285 | No |
| 32 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 11793 | -0.144 | 0.1164 | No |