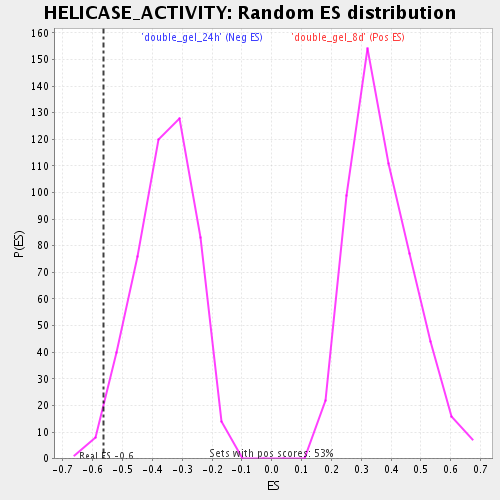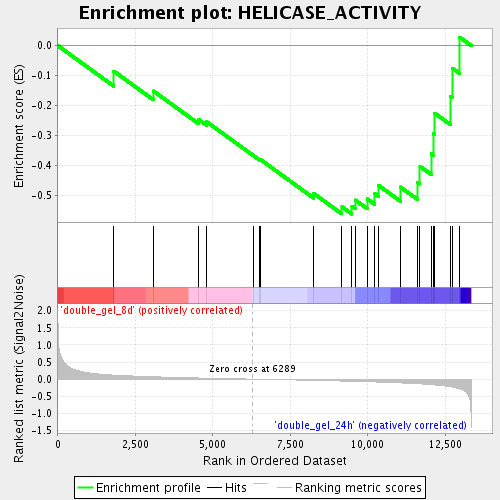
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | HELICASE_ACTIVITY |
| Enrichment Score (ES) | -0.5623577 |
| Normalized Enrichment Score (NES) | -1.5767094 |
| Nominal p-value | 0.012765957 |
| FDR q-value | 0.18481866 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 1801 | 0.117 | -0.0849 | No |
| 2 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 3080 | 0.069 | -0.1513 | No |
| 3 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 4548 | 0.033 | -0.2472 | No |
| 4 | DDX25 | DDX25 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 25 | 4802 | 0.028 | -0.2542 | No |
| 5 | RECQL | RECQL Entrez, Source | RecQ protein-like (DNA helicase Q1-like) | 6311 | -0.000 | -0.3673 | No |
| 6 | SKIV2L | SKIV2L Entrez, Source | superkiller viralicidic activity 2-like (S. cerevisiae) | 6509 | -0.004 | -0.3804 | No |
| 7 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 6550 | -0.005 | -0.3814 | No |
| 8 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8251 | -0.036 | -0.4933 | No |
| 9 | DHX8 | DHX8 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 8 | 9167 | -0.055 | -0.5383 | Yes |
| 10 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 9489 | -0.063 | -0.5355 | Yes |
| 11 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9594 | -0.065 | -0.5152 | Yes |
| 12 | IGHMBP2 | IGHMBP2 Entrez, Source | immunoglobulin mu binding protein 2 | 9982 | -0.076 | -0.5116 | Yes |
| 13 | DHX16 | DHX16 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 16 | 10229 | -0.082 | -0.4945 | Yes |
| 14 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 10351 | -0.086 | -0.4667 | Yes |
| 15 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11050 | -0.110 | -0.4718 | Yes |
| 16 | DDX1 | DDX1 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 1 | 11603 | -0.133 | -0.4559 | Yes |
| 17 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 11685 | -0.137 | -0.4029 | Yes |
| 18 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 12048 | -0.161 | -0.3610 | Yes |
| 19 | DDX18 | DDX18 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 18 | 12109 | -0.165 | -0.2944 | Yes |
| 20 | DDX20 | DDX20 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 | 12161 | -0.169 | -0.2256 | Yes |
| 21 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 12667 | -0.220 | -0.1690 | Yes |
| 22 | DDX56 | DDX56 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 56 | 12725 | -0.229 | -0.0750 | Yes |
| 23 | ASCC3L1 | ASCC3L1 Entrez, Source | activating signal cointegrator 1 complex subunit 3-like 1 | 12968 | -0.282 | 0.0281 | Yes |