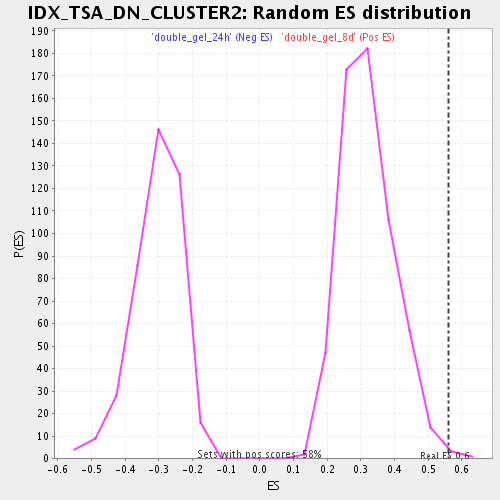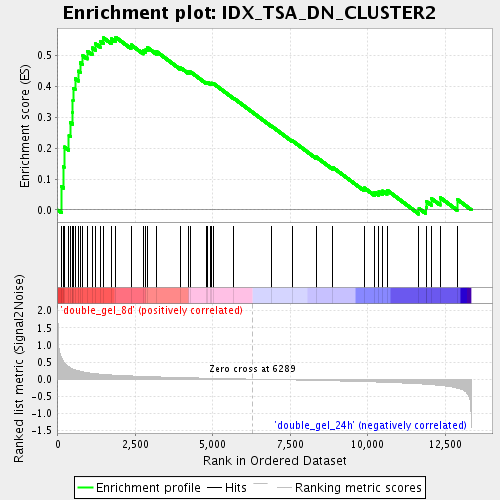
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | IDX_TSA_DN_CLUSTER2 |
| Enrichment Score (ES) | 0.5594627 |
| Normalized Enrichment Score (NES) | 1.7495577 |
| Nominal p-value | 0.0034188034 |
| FDR q-value | 0.039497852 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 114 | 0.625 | 0.0757 | Yes |
| 2 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 183 | 0.515 | 0.1398 | Yes |
| 3 | PRSS23 | PRSS23 Entrez, Source | protease, serine, 23 | 201 | 0.487 | 0.2041 | Yes |
| 4 | FXYD5 | FXYD5 Entrez, Source | FXYD domain containing ion transport regulator 5 | 356 | 0.362 | 0.2413 | Yes |
| 5 | PGCP | PGCP Entrez, Source | - | 406 | 0.335 | 0.2827 | Yes |
| 6 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 476 | 0.294 | 0.3171 | Yes |
| 7 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 484 | 0.291 | 0.3558 | Yes |
| 8 | CD276 | CD276 Entrez, Source | CD276 molecule | 492 | 0.288 | 0.3941 | Yes |
| 9 | LOX | LOX Entrez, Source | lysyl oxidase | 558 | 0.267 | 0.4252 | Yes |
| 10 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 662 | 0.243 | 0.4501 | Yes |
| 11 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 722 | 0.231 | 0.4768 | Yes |
| 12 | PDPN | PDPN Entrez, Source | podoplanin | 803 | 0.213 | 0.4995 | Yes |
| 13 | SLC38A4 | SLC38A4 Entrez, Source | solute carrier family 38, member 4 | 944 | 0.189 | 0.5144 | Yes |
| 14 | PTGIS | PTGIS Entrez, Source | prostaglandin I2 (prostacyclin) synthase | 1107 | 0.170 | 0.5250 | Yes |
| 15 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 1218 | 0.159 | 0.5382 | Yes |
| 16 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1367 | 0.146 | 0.5467 | Yes |
| 17 | PLD3 | PLD3 Entrez, Source | phospholipase D family, member 3 | 1469 | 0.137 | 0.5576 | Yes |
| 18 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 1730 | 0.121 | 0.5543 | Yes |
| 19 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 1866 | 0.113 | 0.5595 | Yes |
| 20 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 2361 | 0.093 | 0.5348 | No |
| 21 | TWIST2 | TWIST2 Entrez, Source | twist homolog 2 (Drosophila) | 2757 | 0.078 | 0.5156 | No |
| 22 | MAP1LC3B | MAP1LC3B Entrez, Source | microtubule-associated protein 1 light chain 3 beta | 2840 | 0.076 | 0.5197 | No |
| 23 | CAPG | CAPG Entrez, Source | capping protein (actin filament), gelsolin-like | 2891 | 0.074 | 0.5260 | No |
| 24 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 3188 | 0.066 | 0.5126 | No |
| 25 | TES | TES Entrez, Source | testis derived transcript (3 LIM domains) | 3963 | 0.045 | 0.4605 | No |
| 26 | DSTN | DSTN Entrez, Source | destrin (actin depolymerizing factor) | 4198 | 0.040 | 0.4482 | No |
| 27 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 4280 | 0.038 | 0.4473 | No |
| 28 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 4800 | 0.028 | 0.4120 | No |
| 29 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 4827 | 0.027 | 0.4137 | No |
| 30 | SCPEP1 | SCPEP1 Entrez, Source | serine carboxypeptidase 1 | 4916 | 0.026 | 0.4106 | No |
| 31 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 4953 | 0.025 | 0.4112 | No |
| 32 | SERTAD1 | SERTAD1 Entrez, Source | SERTA domain containing 1 | 5017 | 0.024 | 0.4097 | No |
| 33 | LEPROT | LEPROT Entrez, Source | leptin receptor overlapping transcript | 5678 | 0.011 | 0.3616 | No |
| 34 | NCAM1 | NCAM1 Entrez, Source | neural cell adhesion molecule 1 | 6887 | -0.010 | 0.2721 | No |
| 35 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 7565 | -0.023 | 0.2243 | No |
| 36 | NQO2 | NQO2 Entrez, Source | NAD(P)H dehydrogenase, quinone 2 | 8329 | -0.038 | 0.1720 | No |
| 37 | PPGB | PPGB Entrez, Source | protective protein for beta-galactosidase (galactosialidosis) | 8872 | -0.049 | 0.1379 | No |
| 38 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 9887 | -0.073 | 0.0715 | No |
| 39 | TINP1 | TINP1 Entrez, Source | - | 10223 | -0.082 | 0.0574 | No |
| 40 | PLXNB2 | PLXNB2 Entrez, Source | plexin B2 | 10361 | -0.086 | 0.0586 | No |
| 41 | BAX | BAX Entrez, Source | BCL2-associated X protein | 10485 | -0.090 | 0.0615 | No |
| 42 | CD99 | CD99 Entrez, Source | CD99 molecule | 10623 | -0.094 | 0.0639 | No |
| 43 | TSPAN6 | TSPAN6 Entrez, Source | tetraspanin 6 | 11653 | -0.136 | 0.0048 | No |
| 44 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 11879 | -0.149 | 0.0080 | No |
| 45 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 11900 | -0.150 | 0.0267 | No |
| 46 | MYH9 | MYH9 Entrez, Source | myosin, heavy chain 9, non-muscle | 12058 | -0.161 | 0.0366 | No |
| 47 | MSN | MSN Entrez, Source | moesin | 12335 | -0.182 | 0.0404 | No |
| 48 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 12884 | -0.262 | 0.0344 | No |