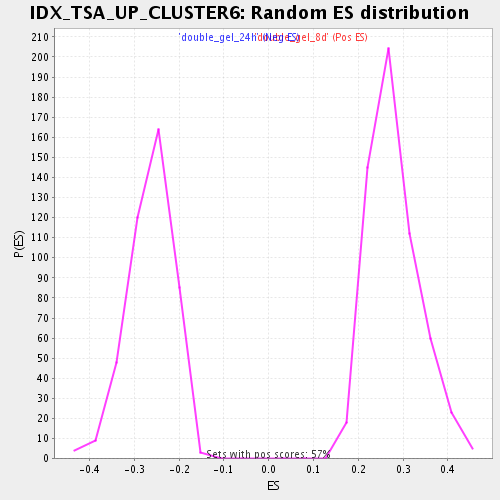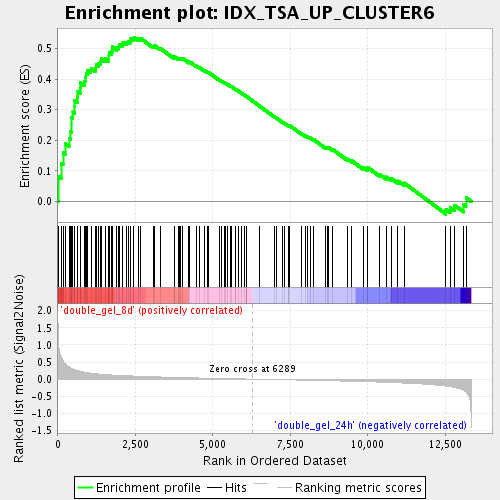
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | IDX_TSA_UP_CLUSTER6 |
| Enrichment Score (ES) | 0.53667766 |
| Normalized Enrichment Score (NES) | 1.9207108 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010332117 |
| FWER p-Value | 0.431 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 15 | 1.102 | 0.0819 | Yes |
| 2 | LPL | LPL Entrez, Source | lipoprotein lipase | 107 | 0.646 | 0.1237 | Yes |
| 3 | AQP7 | AQP7 Entrez, Source | aquaporin 7 | 167 | 0.536 | 0.1596 | Yes |
| 4 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 232 | 0.457 | 0.1892 | Yes |
| 5 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 359 | 0.359 | 0.2068 | Yes |
| 6 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 411 | 0.331 | 0.2278 | Yes |
| 7 | SCARB1 | SCARB1 Entrez, Source | scavenger receptor class B, member 1 | 431 | 0.321 | 0.2505 | Yes |
| 8 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 432 | 0.320 | 0.2746 | Yes |
| 9 | DECR1 | DECR1 Entrez, Source | 2,4-dienoyl CoA reductase 1, mitochondrial | 486 | 0.291 | 0.2926 | Yes |
| 10 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 541 | 0.272 | 0.3090 | Yes |
| 11 | RARRES2 | RARRES2 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 2 | 544 | 0.271 | 0.3292 | Yes |
| 12 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 624 | 0.251 | 0.3422 | Yes |
| 13 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 644 | 0.246 | 0.3593 | Yes |
| 14 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 718 | 0.232 | 0.3712 | Yes |
| 15 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 725 | 0.231 | 0.3881 | Yes |
| 16 | HSD17B12 | HSD17B12 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 12 | 857 | 0.203 | 0.3935 | Yes |
| 17 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 890 | 0.197 | 0.4059 | Yes |
| 18 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 906 | 0.195 | 0.4195 | Yes |
| 19 | ETFDH | ETFDH Entrez, Source | electron-transferring-flavoprotein dehydrogenase | 962 | 0.187 | 0.4294 | Yes |
| 20 | DGAT1 | DGAT1 Entrez, Source | diacylglycerol O-acyltransferase homolog 1 (mouse) | 1067 | 0.174 | 0.4347 | Yes |
| 21 | RETSAT | RETSAT Entrez, Source | retinol saturase (all-trans-retinol 13,14-reductase) | 1210 | 0.160 | 0.4360 | Yes |
| 22 | HIBADH | HIBADH Entrez, Source | 3-hydroxyisobutyrate dehydrogenase | 1229 | 0.158 | 0.4466 | Yes |
| 23 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 1319 | 0.149 | 0.4511 | Yes |
| 24 | SFXN1 | SFXN1 Entrez, Source | sideroflexin 1 | 1384 | 0.144 | 0.4571 | Yes |
| 25 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 1402 | 0.143 | 0.4666 | Yes |
| 26 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1544 | 0.132 | 0.4658 | Yes |
| 27 | CYB5B | CYB5B Entrez, Source | cytochrome b5 type B (outer mitochondrial membrane) | 1646 | 0.126 | 0.4677 | Yes |
| 28 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1647 | 0.125 | 0.4771 | Yes |
| 29 | ADCY6 | ADCY6 Entrez, Source | adenylate cyclase 6 | 1649 | 0.125 | 0.4865 | Yes |
| 30 | ACADL | ACADL Entrez, Source | acyl-Coenzyme A dehydrogenase, long chain | 1720 | 0.122 | 0.4904 | Yes |
| 31 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 1747 | 0.119 | 0.4974 | Yes |
| 32 | SDPR | SDPR Entrez, Source | serum deprivation response (phosphatidylserine binding protein) | 1754 | 0.119 | 0.5059 | Yes |
| 33 | UQCRB | UQCRB Entrez, Source | ubiquinol-cytochrome c reductase binding protein | 1905 | 0.111 | 0.5030 | Yes |
| 34 | TST | TST Entrez, Source | thiosulfate sulfurtransferase (rhodanese) | 1969 | 0.109 | 0.5064 | Yes |
| 35 | MOSC2 | MOSC2 Entrez, Source | MOCO sulphurase C-terminal domain containing 2 | 1987 | 0.108 | 0.5133 | Yes |
| 36 | TSPAN12 | TSPAN12 Entrez, Source | tetraspanin 12 | 2091 | 0.104 | 0.5133 | Yes |
| 37 | FASN | FASN Entrez, Source | fatty acid synthase | 2099 | 0.104 | 0.5206 | Yes |
| 38 | RAB3D | RAB3D Entrez, Source | RAB3D, member RAS oncogene family | 2207 | 0.099 | 0.5200 | Yes |
| 39 | ISOC1 | ISOC1 Entrez, Source | isochorismatase domain containing 1 | 2278 | 0.096 | 0.5220 | Yes |
| 40 | REEP6 | REEP6 Entrez, Source | receptor accessory protein 6 | 2333 | 0.094 | 0.5250 | Yes |
| 41 | HP | HP Entrez, Source | haptoglobin | 2342 | 0.094 | 0.5314 | Yes |
| 42 | EHD1 | EHD1 Entrez, Source | EH-domain containing 1 | 2429 | 0.090 | 0.5317 | Yes |
| 43 | MCCC1 | MCCC1 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 1 (alpha) | 2454 | 0.090 | 0.5367 | Yes |
| 44 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 2593 | 0.084 | 0.5326 | No |
| 45 | MGST1 | MGST1 Entrez, Source | microsomal glutathione S-transferase 1 | 2666 | 0.081 | 0.5332 | No |
| 46 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 3083 | 0.069 | 0.5070 | No |
| 47 | COX7A2 | COX7A2 Entrez, Source | cytochrome c oxidase subunit VIIa polypeptide 2 (liver) | 3123 | 0.068 | 0.5092 | No |
| 48 | SDHC | SDHC Entrez, Source | succinate dehydrogenase complex, subunit C, integral membrane protein, 15kDa | 3311 | 0.062 | 0.4997 | No |
| 49 | SUCLG1 | SUCLG1 Entrez, Source | succinate-CoA ligase, GDP-forming, alpha subunit | 3754 | 0.051 | 0.4702 | No |
| 50 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 3755 | 0.051 | 0.4740 | No |
| 51 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 3905 | 0.047 | 0.4663 | No |
| 52 | DHRS7B | DHRS7B Entrez, Source | dehydrogenase/reductase (SDR family) member 7B | 3919 | 0.047 | 0.4688 | No |
| 53 | SLC25A10 | SLC25A10 Entrez, Source | solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 | 3966 | 0.045 | 0.4688 | No |
| 54 | PCYOX1 | PCYOX1 Entrez, Source | prenylcysteine oxidase 1 | 4016 | 0.044 | 0.4684 | No |
| 55 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 4228 | 0.039 | 0.4554 | No |
| 56 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 4257 | 0.038 | 0.4562 | No |
| 57 | SLC25A1 | SLC25A1 Entrez, Source | solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 | 4469 | 0.035 | 0.4428 | No |
| 58 | SLC19A1 | SLC19A1 Entrez, Source | solute carrier family 19 (folate transporter), member 1 | 4566 | 0.033 | 0.4380 | No |
| 59 | PSEN2 | PSEN2 Entrez, Source | presenilin 2 (Alzheimer disease 4) | 4726 | 0.030 | 0.4283 | No |
| 60 | ADIPOQ | ADIPOQ Entrez, Source | adiponectin, C1Q and collagen domain containing | 4838 | 0.027 | 0.4219 | No |
| 61 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 4853 | 0.027 | 0.4229 | No |
| 62 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 5216 | 0.020 | 0.3971 | No |
| 63 | SLC2A4 | SLC2A4 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 4 | 5283 | 0.019 | 0.3935 | No |
| 64 | PCAF | PCAF Entrez, Source | p300/CBP-associated factor | 5363 | 0.017 | 0.3889 | No |
| 65 | COX5B | COX5B Entrez, Source | cytochrome c oxidase subunit Vb | 5409 | 0.017 | 0.3867 | No |
| 66 | UQCRC2 | UQCRC2 Entrez, Source | ubiquinol-cytochrome c reductase core protein II | 5487 | 0.015 | 0.3820 | No |
| 67 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 5578 | 0.013 | 0.3762 | No |
| 68 | ORM1 | ORM1 Entrez, Source | orosomucoid 1 | 5606 | 0.013 | 0.3751 | No |
| 69 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 5728 | 0.010 | 0.3668 | No |
| 70 | ACAD9 | ACAD9 Entrez, Source | acyl-Coenzyme A dehydrogenase family, member 9 | 5744 | 0.010 | 0.3664 | No |
| 71 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 5816 | 0.009 | 0.3617 | No |
| 72 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 5927 | 0.007 | 0.3539 | No |
| 73 | NDUFA5 | NDUFA5 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5, 13kDa | 6009 | 0.005 | 0.3481 | No |
| 74 | SLC25A5 | SLC25A5 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 | 6080 | 0.004 | 0.3431 | No |
| 75 | BCAT2 | BCAT2 Entrez, Source | branched chain aminotransferase 2, mitochondrial | 6086 | 0.004 | 0.3430 | No |
| 76 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 6088 | 0.003 | 0.3432 | No |
| 77 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 6502 | -0.004 | 0.3123 | No |
| 78 | COL15A1 | COL15A1 Entrez, Source | collagen, type XV, alpha 1 | 6979 | -0.012 | 0.2772 | No |
| 79 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 7051 | -0.013 | 0.2728 | No |
| 80 | ALAD | ALAD Entrez, Source | aminolevulinate, delta-, dehydratase | 7244 | -0.017 | 0.2596 | No |
| 81 | PEX19 | PEX19 Entrez, Source | peroxisomal biogenesis factor 19 | 7306 | -0.018 | 0.2563 | No |
| 82 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 7441 | -0.021 | 0.2478 | No |
| 83 | UNC119 | UNC119 Entrez, Source | unc-119 homolog (C. elegans) | 7443 | -0.021 | 0.2492 | No |
| 84 | LIAS | LIAS Entrez, Source | lipoic acid synthetase | 7484 | -0.021 | 0.2478 | No |
| 85 | SAMM50 | SAMM50 Entrez, Source | sorting and assembly machinery component 50 homolog (S. cerevisiae) | 7865 | -0.028 | 0.2213 | No |
| 86 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 7978 | -0.031 | 0.2151 | No |
| 87 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 8051 | -0.032 | 0.2121 | No |
| 88 | HIBCH | HIBCH Entrez, Source | 3-hydroxyisobutyryl-Coenzyme A hydrolase | 8146 | -0.034 | 0.2076 | No |
| 89 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 8263 | -0.037 | 0.2016 | No |
| 90 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 8635 | -0.044 | 0.1769 | No |
| 91 | CAMP | CAMP Entrez, Source | cathelicidin antimicrobial peptide | 8702 | -0.045 | 0.1753 | No |
| 92 | HBLD2 | HBLD2 Entrez, Source | HESB like domain containing 2 | 8732 | -0.046 | 0.1766 | No |
| 93 | CAT | CAT Entrez, Source | catalase | 8873 | -0.049 | 0.1697 | No |
| 94 | LTC4S | LTC4S Entrez, Source | leukotriene C4 synthase | 9342 | -0.059 | 0.1388 | No |
| 95 | PLA2G6 | PLA2G6 Entrez, Source | phospholipase A2, group VI (cytosolic, calcium-independent) | 9476 | -0.062 | 0.1335 | No |
| 96 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 9855 | -0.073 | 0.1104 | No |
| 97 | S100A1 | S100A1 Entrez, Source | S100 calcium binding protein A1 | 9993 | -0.076 | 0.1058 | No |
| 98 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 9995 | -0.076 | 0.1114 | No |
| 99 | CIB2 | CIB2 Entrez, Source | calcium and integrin binding family member 2 | 10393 | -0.087 | 0.0880 | No |
| 100 | PDAP1 | PDAP1 Entrez, Source | PDGFA associated protein 1 | 10592 | -0.093 | 0.0801 | No |
| 101 | CA2 | CA2 Entrez, Source | carbonic anhydrase II | 10759 | -0.099 | 0.0750 | No |
| 102 | RTN3 | RTN3 Entrez, Source | reticulon 3 | 10973 | -0.107 | 0.0670 | No |
| 103 | CBR1 | CBR1 Entrez, Source | carbonyl reductase 1 | 11177 | -0.115 | 0.0604 | No |
| 104 | TKT | TKT Entrez, Source | transketolase (Wernicke-Korsakoff syndrome) | 12523 | -0.202 | -0.0261 | No |
| 105 | CS | CS Entrez, Source | citrate synthase | 12671 | -0.220 | -0.0206 | No |
| 106 | GBE1 | GBE1 Entrez, Source | glucan (1,4-alpha-), branching enzyme 1 (glycogen branching enzyme, Andersen disease, glycogen storage disease type IV) | 12796 | -0.240 | -0.0118 | No |
| 107 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 13097 | -0.326 | -0.0099 | No |
| 108 | GSR | GSR Entrez, Source | glutathione reductase | 13171 | -0.376 | 0.0129 | No |