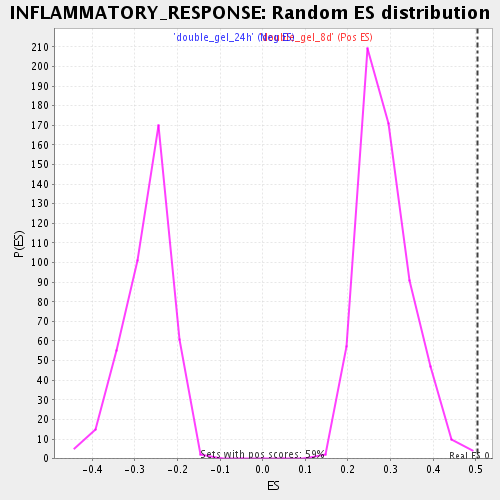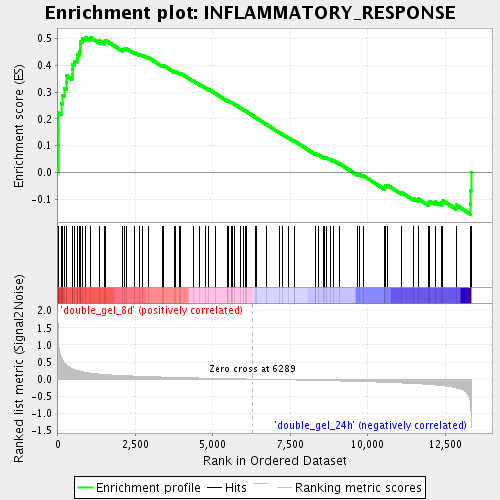
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | INFLAMMATORY_RESPONSE |
| Enrichment Score (ES) | 0.5037141 |
| Normalized Enrichment Score (NES) | 1.7539337 |
| Nominal p-value | 0.0016920473 |
| FDR q-value | 0.039468013 |
| FWER p-Value | 0.995 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.1013 | Yes |
| 2 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 25 | 0.995 | 0.1620 | Yes |
| 3 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 27 | 0.963 | 0.2221 | Yes |
| 4 | LYZ | LYZ Entrez, Source | lysozyme (renal amyloidosis) | 103 | 0.659 | 0.2577 | Yes |
| 5 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 159 | 0.550 | 0.2880 | Yes |
| 6 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 218 | 0.471 | 0.3131 | Yes |
| 7 | CRP | CRP Entrez, Source | C-reactive protein, pentraxin-related | 272 | 0.422 | 0.3355 | Yes |
| 8 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 279 | 0.415 | 0.3610 | Yes |
| 9 | KLRG1 | KLRG1 Entrez, Source | killer cell lectin-like receptor subfamily G, member 1 | 457 | 0.304 | 0.3666 | Yes |
| 10 | IRAK2 | IRAK2 Entrez, Source | interleukin-1 receptor-associated kinase 2 | 464 | 0.300 | 0.3850 | Yes |
| 11 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 474 | 0.294 | 0.4027 | Yes |
| 12 | CD40 | CD40 Entrez, Source | CD40 molecule, TNF receptor superfamily member 5 | 539 | 0.273 | 0.4149 | Yes |
| 13 | TPST1 | TPST1 Entrez, Source | tyrosylprotein sulfotransferase 1 | 617 | 0.253 | 0.4250 | Yes |
| 14 | LBP | LBP Entrez, Source | lipopolysaccharide binding protein | 636 | 0.247 | 0.4391 | Yes |
| 15 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 689 | 0.237 | 0.4500 | Yes |
| 16 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 718 | 0.232 | 0.4624 | Yes |
| 17 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 725 | 0.231 | 0.4764 | Yes |
| 18 | ALOX5AP | ALOX5AP Entrez, Source | arachidonate 5-lipoxygenase-activating protein | 740 | 0.229 | 0.4896 | Yes |
| 19 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 796 | 0.215 | 0.4989 | Yes |
| 20 | ADORA1 | ADORA1 Entrez, Source | adenosine A1 receptor | 901 | 0.195 | 0.5033 | Yes |
| 21 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 1043 | 0.177 | 0.5037 | Yes |
| 22 | IL8RB | IL8RB Entrez, Source | interleukin 8 receptor, beta | 1345 | 0.147 | 0.4902 | No |
| 23 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 1508 | 0.134 | 0.4864 | No |
| 24 | IL8RA | IL8RA Entrez, Source | interleukin 8 receptor, alpha | 1527 | 0.133 | 0.4933 | No |
| 25 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 2076 | 0.105 | 0.4585 | No |
| 26 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 2133 | 0.102 | 0.4607 | No |
| 27 | ALOX15 | ALOX15 Entrez, Source | arachidonate 15-lipoxygenase | 2217 | 0.099 | 0.4606 | No |
| 28 | AHSG | AHSG Entrez, Source | alpha-2-HS-glycoprotein | 2467 | 0.089 | 0.4474 | No |
| 29 | ADORA3 | ADORA3 Entrez, Source | adenosine A3 receptor | 2617 | 0.083 | 0.4413 | No |
| 30 | BLNK | BLNK Entrez, Source | B-cell linker | 2743 | 0.079 | 0.4368 | No |
| 31 | GHSR | GHSR Entrez, Source | growth hormone secretagogue receptor | 2907 | 0.074 | 0.4292 | No |
| 32 | CCL11 | CCL11 Entrez, Source | chemokine (C-C motif) ligand 11 | 3370 | 0.061 | 0.3981 | No |
| 33 | PARP4 | PARP4 Entrez, Source | poly (ADP-ribose) polymerase family, member 4 | 3401 | 0.060 | 0.3996 | No |
| 34 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 3767 | 0.051 | 0.3753 | No |
| 35 | AGER | AGER Entrez, Source | advanced glycosylation end product-specific receptor | 3800 | 0.050 | 0.3760 | No |
| 36 | CHRNA7 | CHRNA7 Entrez, Source | cholinergic receptor, nicotinic, alpha 7 | 3936 | 0.046 | 0.3687 | No |
| 37 | CCR2 | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 3950 | 0.046 | 0.3706 | No |
| 38 | F11R | F11R Entrez, Source | F11 receptor | 4372 | 0.036 | 0.3411 | No |
| 39 | CCR5 | CCR5 Entrez, Source | chemokine (C-C motif) receptor 5 | 4583 | 0.032 | 0.3273 | No |
| 40 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 4778 | 0.028 | 0.3144 | No |
| 41 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 4875 | 0.027 | 0.3088 | No |
| 42 | TACR1 | TACR1 Entrez, Source | tachykinin receptor 1 | 5085 | 0.023 | 0.2945 | No |
| 43 | MEFV | MEFV Entrez, Source | Mediterranean fever | 5464 | 0.016 | 0.2669 | No |
| 44 | NOX4 | NOX4 Entrez, Source | NADPH oxidase 4 | 5475 | 0.015 | 0.2671 | No |
| 45 | CCR4 | CCR4 Entrez, Source | chemokine (C-C motif) receptor 4 | 5494 | 0.015 | 0.2667 | No |
| 46 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 5594 | 0.013 | 0.2600 | No |
| 47 | ORM1 | ORM1 Entrez, Source | orosomucoid 1 | 5606 | 0.013 | 0.2600 | No |
| 48 | SIGIRR | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 5629 | 0.012 | 0.2591 | No |
| 49 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 5694 | 0.011 | 0.2550 | No |
| 50 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 5892 | 0.007 | 0.2406 | No |
| 51 | CD97 | CD97 Entrez, Source | CD97 molecule | 5995 | 0.005 | 0.2332 | No |
| 52 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 6062 | 0.004 | 0.2285 | No |
| 53 | IL5 | IL5 Entrez, Source | interleukin 5 (colony-stimulating factor, eosinophil) | 6076 | 0.004 | 0.2277 | No |
| 54 | SCG2 | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 6379 | -0.001 | 0.2050 | No |
| 55 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 6393 | -0.002 | 0.2041 | No |
| 56 | IL13 | IL13 Entrez, Source | interleukin 13 | 6740 | -0.008 | 0.1785 | No |
| 57 | CCL22 | CCL22 Entrez, Source | chemokine (C-C motif) ligand 22 | 7142 | -0.015 | 0.1492 | No |
| 58 | CD40LG | CD40LG Entrez, Source | CD40 ligand (TNF superfamily, member 5, hyper-IgM syndrome) | 7233 | -0.017 | 0.1434 | No |
| 59 | ELF3 | ELF3 Entrez, Source | E74-like factor 3 (ets domain transcription factor, epithelial-specific ) | 7442 | -0.021 | 0.1290 | No |
| 60 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 7625 | -0.024 | 0.1168 | No |
| 61 | PTAFR | PTAFR Entrez, Source | platelet-activating factor receptor | 7629 | -0.024 | 0.1181 | No |
| 62 | LTB4R | LTB4R Entrez, Source | leukotriene B4 receptor | 8298 | -0.037 | 0.0700 | No |
| 63 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 8306 | -0.037 | 0.0718 | No |
| 64 | RIPK2 | RIPK2 Entrez, Source | receptor-interacting serine-threonine kinase 2 | 8400 | -0.039 | 0.0673 | No |
| 65 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 8582 | -0.043 | 0.0563 | No |
| 66 | HRH1 | HRH1 Entrez, Source | histamine receptor H1 | 8600 | -0.043 | 0.0577 | No |
| 67 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 8654 | -0.044 | 0.0565 | No |
| 68 | CCR3 | CCR3 Entrez, Source | chemokine (C-C motif) receptor 3 | 8807 | -0.048 | 0.0480 | No |
| 69 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 8894 | -0.050 | 0.0447 | No |
| 70 | RAC1 | RAC1 Entrez, Source | ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) | 9101 | -0.054 | 0.0325 | No |
| 71 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 9683 | -0.067 | -0.0071 | No |
| 72 | PLA2G2D | PLA2G2D Entrez, Source | phospholipase A2, group IID | 9730 | -0.069 | -0.0063 | No |
| 73 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 9852 | -0.072 | -0.0109 | No |
| 74 | C2 | C2 Entrez, Source | complement component 2 | 10542 | -0.091 | -0.0571 | No |
| 75 | PRDX5 | PRDX5 Entrez, Source | peroxiredoxin 5 | 10551 | -0.092 | -0.0520 | No |
| 76 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 10574 | -0.093 | -0.0479 | No |
| 77 | SELE | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 10650 | -0.095 | -0.0476 | No |
| 78 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 11087 | -0.111 | -0.0735 | No |
| 79 | CFHR1 | CFHR1 Entrez, Source | complement factor H-related 1 | 11475 | -0.128 | -0.0947 | No |
| 80 | APCS | APCS Entrez, Source | amyloid P component, serum | 11626 | -0.135 | -0.0976 | No |
| 81 | MBL2 | MBL2 Entrez, Source | mannose-binding lectin (protein C) 2, soluble (opsonic defect) | 11957 | -0.154 | -0.1128 | No |
| 82 | ABCF1 | ABCF1 Entrez, Source | ATP-binding cassette, sub-family F (GCN20), member 1 | 11995 | -0.157 | -0.1058 | No |
| 83 | NFX1 | NFX1 Entrez, Source | nuclear transcription factor, X-box binding 1 | 12176 | -0.170 | -0.1088 | No |
| 84 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 12379 | -0.186 | -0.1124 | No |
| 85 | SCYE1 | SCYE1 Entrez, Source | small inducible cytokine subfamily E, member 1 (endothelial monocyte-activating) | 12401 | -0.188 | -0.1022 | No |
| 86 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 12851 | -0.254 | -0.1202 | No |
| 87 | AOX1 | AOX1 Entrez, Source | aldehyde oxidase 1 | 13304 | -0.615 | -0.1158 | No |
| 88 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 13322 | -0.789 | -0.0677 | No |
| 89 | CXCL9 | CXCL9 Entrez, Source | chemokine (C-X-C motif) ligand 9 | 13337 | -1.105 | 0.0004 | No |