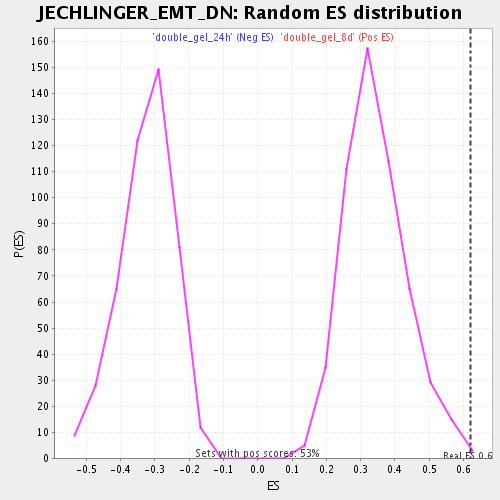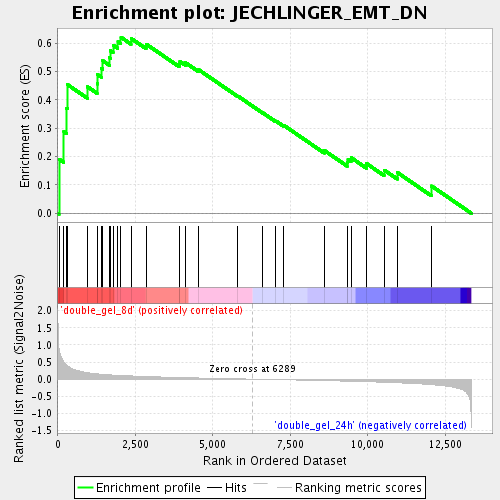
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | JECHLINGER_EMT_DN |
| Enrichment Score (ES) | 0.6212379 |
| Normalized Enrichment Score (NES) | 1.8102335 |
| Nominal p-value | 0.0018726592 |
| FDR q-value | 0.026325231 |
| FWER p-Value | 0.919 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 37 | 0.888 | 0.1907 | Yes |
| 2 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 191 | 0.498 | 0.2878 | Yes |
| 3 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 282 | 0.413 | 0.3711 | Yes |
| 4 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.4539 | Yes |
| 5 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 956 | 0.188 | 0.4465 | Yes |
| 6 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1266 | 0.154 | 0.4569 | Yes |
| 7 | TACSTD1 | TACSTD1 Entrez, Source | tumor-associated calcium signal transducer 1 | 1278 | 0.153 | 0.4893 | Yes |
| 8 | CTSH | CTSH Entrez, Source | cathepsin H | 1401 | 0.143 | 0.5113 | Yes |
| 9 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1445 | 0.139 | 0.5384 | Yes |
| 10 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 1666 | 0.124 | 0.5490 | Yes |
| 11 | SERPINB5 | SERPINB5 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 5 | 1692 | 0.123 | 0.5740 | Yes |
| 12 | VAMP8 | VAMP8 Entrez, Source | vesicle-associated membrane protein 8 (endobrevin) | 1782 | 0.118 | 0.5930 | Yes |
| 13 | NUMB | NUMB Entrez, Source | numb homolog (Drosophila) | 1938 | 0.110 | 0.6053 | Yes |
| 14 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 2035 | 0.106 | 0.6212 | Yes |
| 15 | ATP1A1 | ATP1A1 Entrez, Source | ATPase, Na+/K+ transporting, alpha 1 polypeptide | 2360 | 0.093 | 0.6172 | No |
| 16 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 2848 | 0.076 | 0.5971 | No |
| 17 | ARHGEF1 | ARHGEF1 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 1 | 3915 | 0.047 | 0.5272 | No |
| 18 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 3937 | 0.046 | 0.5357 | No |
| 19 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 4113 | 0.042 | 0.5317 | No |
| 20 | GRB7 | GRB7 Entrez, Source | growth factor receptor-bound protein 7 | 4549 | 0.033 | 0.5062 | No |
| 21 | PADI2 | PADI2 Entrez, Source | peptidyl arginine deiminase, type II | 5804 | 0.009 | 0.4140 | No |
| 22 | STAT5A | STAT5A Entrez, Source | signal transducer and activator of transcription 5A | 6613 | -0.006 | 0.3545 | No |
| 23 | FBP2 | FBP2 Entrez, Source | fructose-1,6-bisphosphatase 2 | 7034 | -0.013 | 0.3258 | No |
| 24 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 7282 | -0.018 | 0.3111 | No |
| 25 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 8595 | -0.043 | 0.2220 | No |
| 26 | ACTN4 | ACTN4 Entrez, Source | actinin, alpha 4 | 9335 | -0.059 | 0.1793 | No |
| 27 | KITLG | KITLG Entrez, Source | KIT ligand | 9361 | -0.060 | 0.1904 | No |
| 28 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 9463 | -0.062 | 0.1964 | No |
| 29 | JUP | JUP Entrez, Source | junction plakoglobin | 9963 | -0.075 | 0.1753 | No |
| 30 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 10528 | -0.091 | 0.1528 | No |
| 31 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 10955 | -0.107 | 0.1440 | No |
| 32 | MYH9 | MYH9 Entrez, Source | myosin, heavy chain 9, non-muscle | 12058 | -0.161 | 0.0965 | No |