Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | KTGGYRSGAA_UNKNOWN |
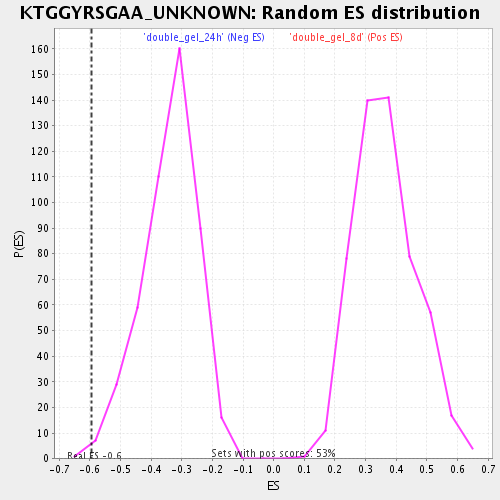
| Enrichment Score (ES) | -0.5953538 |
| Normalized Enrichment Score (NES) | -1.7401326 |
| Nominal p-value | 0.004237288 |
| FDR q-value | 0.07644372 |
| FWER p-Value | 0.974 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 932 | 0.190 | -0.0188 | No |
| 2 | SH3KBP1 | SH3KBP1 Entrez, Source | SH3-domain kinase binding protein 1 | 1293 | 0.151 | -0.0052 | No |
| 3 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 1722 | 0.122 | -0.0046 | No |
| 4 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1796 | 0.117 | 0.0214 | No |
| 5 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 2867 | 0.075 | -0.0388 | No |
| 6 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 3971 | 0.045 | -0.1094 | No |
| 7 | SEC61A1 | SEC61A1 Entrez, Source | Sec61 alpha 1 subunit (S. cerevisiae) | 6253 | 0.001 | -0.2806 | No |
| 8 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 7401 | -0.020 | -0.3613 | No |
| 9 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 7584 | -0.023 | -0.3688 | No |
| 10 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 7775 | -0.027 | -0.3758 | No |
| 11 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9009 | -0.052 | -0.4544 | No |
| 12 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 9277 | -0.058 | -0.4589 | No |
| 13 | GDI1 | GDI1 Entrez, Source | GDP dissociation inhibitor 1 | 9910 | -0.074 | -0.4865 | No |
| 14 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 9947 | -0.075 | -0.4690 | No |
| 15 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10438 | -0.088 | -0.4820 | No |
| 16 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 11948 | -0.154 | -0.5541 | Yes |
| 17 | HTF9C | HTF9C Entrez, Source | - | 11970 | -0.155 | -0.5139 | Yes |
| 18 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 12249 | -0.175 | -0.4877 | Yes |
| 19 | TUBA4 | TUBA4 Entrez, Source | tubulin, alpha 4 | 12345 | -0.183 | -0.4456 | Yes |
| 20 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 12783 | -0.238 | -0.4144 | Yes |
| 21 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 12830 | -0.249 | -0.3508 | Yes |
| 22 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12917 | -0.270 | -0.2847 | Yes |
| 23 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 13014 | -0.296 | -0.2123 | Yes |
| 24 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13101 | -0.328 | -0.1307 | Yes |
| 25 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13288 | -0.553 | 0.0041 | Yes |