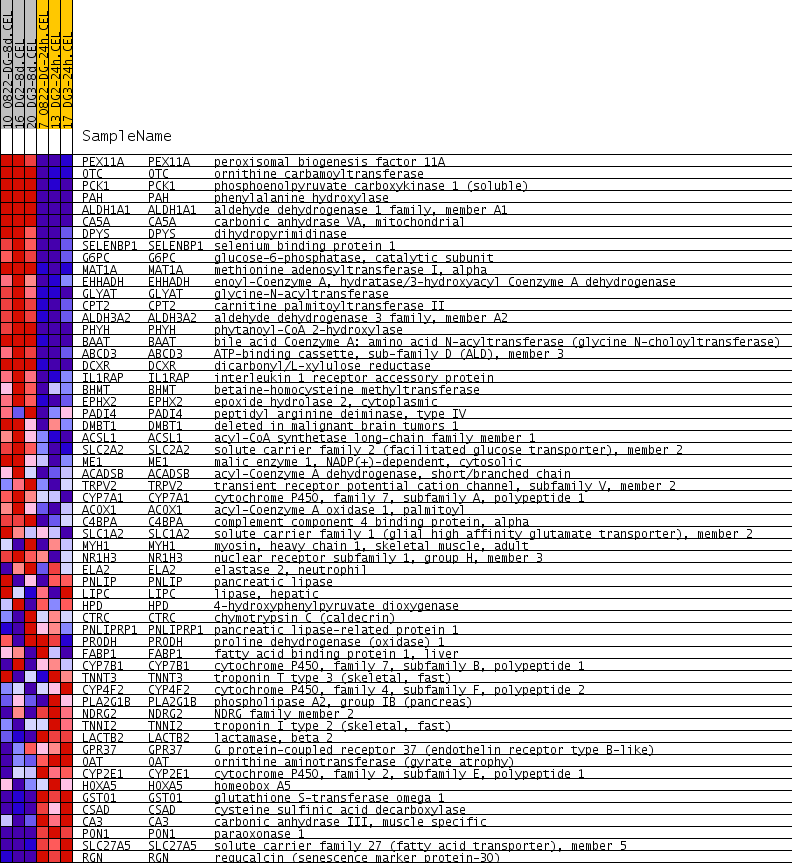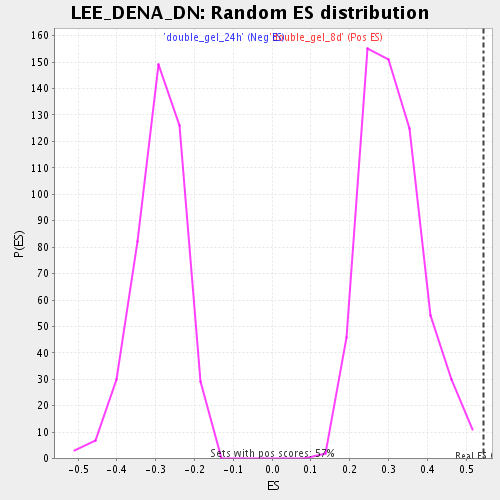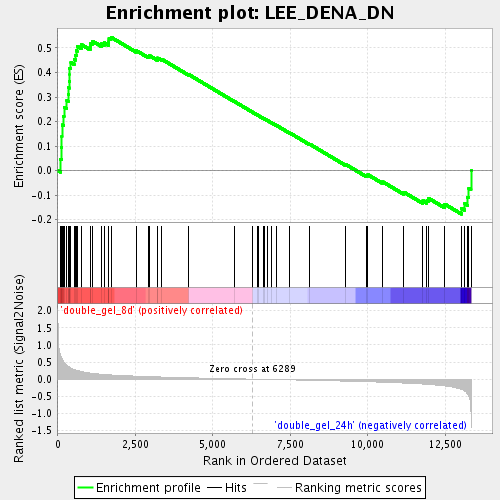
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | LEE_DENA_DN |
| Enrichment Score (ES) | 0.54384035 |
| Normalized Enrichment Score (NES) | 1.7513038 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.039334044 |
| FWER p-Value | 0.995 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PEX11A | PEX11A Entrez, Source | peroxisomal biogenesis factor 11A | 86 | 0.682 | 0.0467 | Yes |
| 2 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 115 | 0.625 | 0.0932 | Yes |
| 3 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 127 | 0.610 | 0.1399 | Yes |
| 4 | PAH | PAH Entrez, Source | phenylalanine hydroxylase | 133 | 0.597 | 0.1861 | Yes |
| 5 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.2220 | Yes |
| 6 | CA5A | CA5A Entrez, Source | carbonic anhydrase VA, mitochondrial | 199 | 0.487 | 0.2590 | Yes |
| 7 | DPYS | DPYS Entrez, Source | dihydropyrimidinase | 287 | 0.410 | 0.2844 | Yes |
| 8 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 336 | 0.371 | 0.3097 | Yes |
| 9 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 347 | 0.366 | 0.3375 | Yes |
| 10 | MAT1A | MAT1A Entrez, Source | methionine adenosyltransferase I, alpha | 362 | 0.357 | 0.3643 | Yes |
| 11 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 375 | 0.349 | 0.3906 | Yes |
| 12 | GLYAT | GLYAT Entrez, Source | glycine-N-acyltransferase | 379 | 0.348 | 0.4175 | Yes |
| 13 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 419 | 0.326 | 0.4399 | Yes |
| 14 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 526 | 0.277 | 0.4535 | Yes |
| 15 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 566 | 0.265 | 0.4712 | Yes |
| 16 | BAAT | BAAT Entrez, Source | bile acid Coenzyme A: amino acid N-acyltransferase (glycine N-choloyltransferase) | 602 | 0.257 | 0.4885 | Yes |
| 17 | ABCD3 | ABCD3 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 3 | 622 | 0.251 | 0.5067 | Yes |
| 18 | DCXR | DCXR Entrez, Source | dicarbonyl/L-xylulose reductase | 765 | 0.222 | 0.5133 | Yes |
| 19 | IL1RAP | IL1RAP Entrez, Source | interleukin 1 receptor accessory protein | 1043 | 0.177 | 0.5063 | Yes |
| 20 | BHMT | BHMT Entrez, Source | betaine-homocysteine methyltransferase | 1054 | 0.177 | 0.5193 | Yes |
| 21 | EPHX2 | EPHX2 Entrez, Source | epoxide hydrolase 2, cytoplasmic | 1108 | 0.170 | 0.5285 | Yes |
| 22 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 1411 | 0.142 | 0.5169 | Yes |
| 23 | DMBT1 | DMBT1 Entrez, Source | deleted in malignant brain tumors 1 | 1494 | 0.135 | 0.5212 | Yes |
| 24 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 1627 | 0.127 | 0.5212 | Yes |
| 25 | SLC2A2 | SLC2A2 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 2 | 1645 | 0.126 | 0.5297 | Yes |
| 26 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1647 | 0.125 | 0.5394 | Yes |
| 27 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 1715 | 0.122 | 0.5438 | Yes |
| 28 | TRPV2 | TRPV2 Entrez, Source | transient receptor potential cation channel, subfamily V, member 2 | 2522 | 0.087 | 0.4899 | No |
| 29 | CYP7A1 | CYP7A1 Entrez, Source | cytochrome P450, family 7, subfamily A, polypeptide 1 | 2927 | 0.073 | 0.4653 | No |
| 30 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 2947 | 0.073 | 0.4695 | No |
| 31 | C4BPA | C4BPA Entrez, Source | complement component 4 binding protein, alpha | 3211 | 0.065 | 0.4548 | No |
| 32 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 3219 | 0.065 | 0.4593 | No |
| 33 | MYH1 | MYH1 Entrez, Source | myosin, heavy chain 1, skeletal muscle, adult | 3349 | 0.061 | 0.4544 | No |
| 34 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 4228 | 0.039 | 0.3913 | No |
| 35 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 5704 | 0.011 | 0.2811 | No |
| 36 | PNLIP | PNLIP Entrez, Source | pancreatic lipase | 6265 | 0.000 | 0.2390 | No |
| 37 | LIPC | LIPC Entrez, Source | lipase, hepatic | 6430 | -0.002 | 0.2268 | No |
| 38 | HPD | HPD Entrez, Source | 4-hydroxyphenylpyruvate dioxygenase | 6481 | -0.003 | 0.2233 | No |
| 39 | CTRC | CTRC Entrez, Source | chymotrypsin C (caldecrin) | 6631 | -0.006 | 0.2126 | No |
| 40 | PNLIPRP1 | PNLIPRP1 Entrez, Source | pancreatic lipase-related protein 1 | 6658 | -0.007 | 0.2111 | No |
| 41 | PRODH | PRODH Entrez, Source | proline dehydrogenase (oxidase) 1 | 6759 | -0.008 | 0.2042 | No |
| 42 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 6898 | -0.010 | 0.1947 | No |
| 43 | CYP7B1 | CYP7B1 Entrez, Source | cytochrome P450, family 7, subfamily B, polypeptide 1 | 7069 | -0.014 | 0.1829 | No |
| 44 | TNNT3 | TNNT3 Entrez, Source | troponin T type 3 (skeletal, fast) | 7488 | -0.021 | 0.1531 | No |
| 45 | CYP4F2 | CYP4F2 Entrez, Source | cytochrome P450, family 4, subfamily F, polypeptide 2 | 8129 | -0.034 | 0.1076 | No |
| 46 | PLA2G1B | PLA2G1B Entrez, Source | phospholipase A2, group IB (pancreas) | 9289 | -0.058 | 0.0249 | No |
| 47 | NDRG2 | NDRG2 Entrez, Source | NDRG family member 2 | 9944 | -0.075 | -0.0185 | No |
| 48 | TNNI2 | TNNI2 Entrez, Source | troponin I type 2 (skeletal, fast) | 10004 | -0.077 | -0.0170 | No |
| 49 | LACTB2 | LACTB2 Entrez, Source | lactamase, beta 2 | 10489 | -0.090 | -0.0464 | No |
| 50 | GPR37 | GPR37 Entrez, Source | G protein-coupled receptor 37 (endothelin receptor type B-like) | 11157 | -0.115 | -0.0877 | No |
| 51 | OAT | OAT Entrez, Source | ornithine aminotransferase (gyrate atrophy) | 11781 | -0.143 | -0.1234 | No |
| 52 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 11911 | -0.151 | -0.1214 | No |
| 53 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 11961 | -0.154 | -0.1131 | No |
| 54 | GSTO1 | GSTO1 Entrez, Source | glutathione S-transferase omega 1 | 12484 | -0.197 | -0.1370 | No |
| 55 | CSAD | CSAD Entrez, Source | cysteine sulfinic acid decarboxylase | 13026 | -0.301 | -0.1543 | No |
| 56 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13129 | -0.343 | -0.1353 | No |
| 57 | PON1 | PON1 Entrez, Source | paraoxonase 1 | 13224 | -0.431 | -0.1087 | No |
| 58 | SLC27A5 | SLC27A5 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 5 | 13261 | -0.509 | -0.0718 | No |
| 59 | RGN | RGN Entrez, Source | regucalcin (senescence marker protein-30) | 13333 | -0.998 | 0.0007 | No |