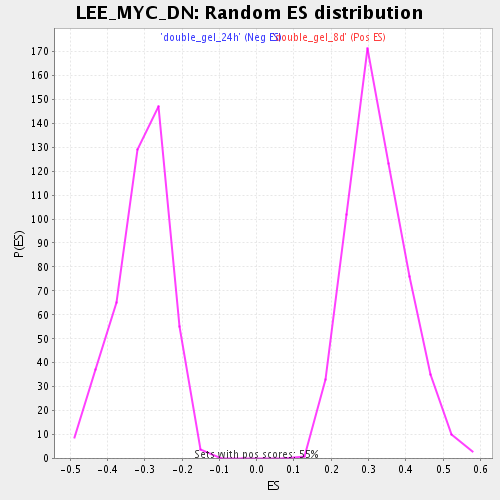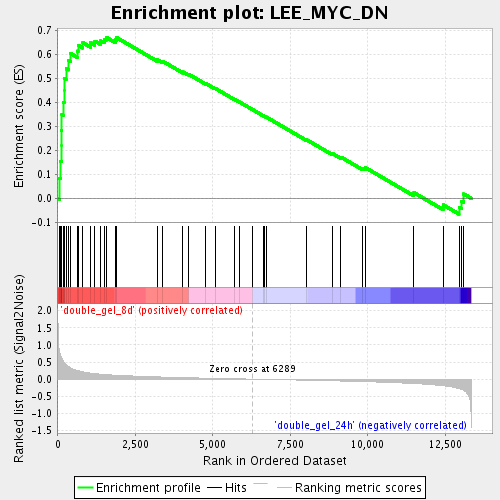
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | LEE_MYC_DN |
| Enrichment Score (ES) | 0.67052627 |
| Normalized Enrichment Score (NES) | 2.0723057 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0020569765 |
| FWER p-Value | 0.033 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 47 | 0.829 | 0.0837 | Yes |
| 2 | GOT1 | GOT1 Entrez, Source | glutamic-oxaloacetic transaminase 1, soluble (aspartate aminotransferase 1) | 76 | 0.697 | 0.1549 | Yes |
| 3 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 112 | 0.632 | 0.2187 | Yes |
| 4 | OTC | OTC Entrez, Source | ornithine carbamoyltransferase | 115 | 0.625 | 0.2843 | Yes |
| 5 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 127 | 0.610 | 0.3476 | Yes |
| 6 | TDO2 | TDO2 Entrez, Source | tryptophan 2,3-dioxygenase | 170 | 0.532 | 0.4005 | Yes |
| 7 | KLF15 | KLF15 Entrez, Source | Kruppel-like factor 15 | 202 | 0.486 | 0.4493 | Yes |
| 8 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 205 | 0.482 | 0.4999 | Yes |
| 9 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 267 | 0.426 | 0.5401 | Yes |
| 10 | G6PC | G6PC Entrez, Source | glucose-6-phosphatase, catalytic subunit | 347 | 0.366 | 0.5727 | Yes |
| 11 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 412 | 0.330 | 0.6027 | Yes |
| 12 | SLC37A4 | SLC37A4 Entrez, Source | solute carrier family 37 (glycerol-6-phosphate transporter), member 4 | 623 | 0.251 | 0.6133 | Yes |
| 13 | HRSP12 | HRSP12 Entrez, Source | heat-responsive protein 12 | 654 | 0.244 | 0.6367 | Yes |
| 14 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 804 | 0.213 | 0.6479 | Yes |
| 15 | BHMT | BHMT Entrez, Source | betaine-homocysteine methyltransferase | 1054 | 0.177 | 0.6478 | Yes |
| 16 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 1190 | 0.162 | 0.6546 | Yes |
| 17 | SFXN1 | SFXN1 Entrez, Source | sideroflexin 1 | 1384 | 0.144 | 0.6553 | Yes |
| 18 | DMBT1 | DMBT1 Entrez, Source | deleted in malignant brain tumors 1 | 1494 | 0.135 | 0.6613 | Yes |
| 19 | IGFBP2 | IGFBP2 Entrez, Source | insulin-like growth factor binding protein 2, 36kDa | 1556 | 0.131 | 0.6705 | Yes |
| 20 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 1861 | 0.114 | 0.6596 | No |
| 21 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 1883 | 0.113 | 0.6699 | No |
| 22 | C4BPA | C4BPA Entrez, Source | complement component 4 binding protein, alpha | 3211 | 0.065 | 0.5770 | No |
| 23 | EFEMP1 | EFEMP1 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 1 | 3378 | 0.061 | 0.5709 | No |
| 24 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4036 | 0.044 | 0.5261 | No |
| 25 | CFH | CFH Entrez, Source | complement factor H | 4209 | 0.039 | 0.5173 | No |
| 26 | ZG16 | ZG16 Entrez, Source | - | 4769 | 0.029 | 0.4783 | No |
| 27 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 5070 | 0.023 | 0.4581 | No |
| 28 | ELA2 | ELA2 Entrez, Source | elastase 2, neutrophil | 5704 | 0.011 | 0.4117 | No |
| 29 | HAL | HAL Entrez, Source | histidine ammonia-lyase | 5856 | 0.008 | 0.4012 | No |
| 30 | PNLIP | PNLIP Entrez, Source | pancreatic lipase | 6265 | 0.000 | 0.3705 | No |
| 31 | CTRC | CTRC Entrez, Source | chymotrypsin C (caldecrin) | 6631 | -0.006 | 0.3437 | No |
| 32 | PNLIPRP1 | PNLIPRP1 Entrez, Source | pancreatic lipase-related protein 1 | 6658 | -0.007 | 0.3425 | No |
| 33 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 6718 | -0.007 | 0.3388 | No |
| 34 | MEOX2 | MEOX2 Entrez, Source | mesenchyme homeobox 2 | 8026 | -0.032 | 0.2439 | No |
| 35 | AQP8 | AQP8 Entrez, Source | aquaporin 8 | 8862 | -0.049 | 0.1863 | No |
| 36 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 9134 | -0.055 | 0.1716 | No |
| 37 | FGG | FGG Entrez, Source | fibrinogen gamma chain | 9841 | -0.072 | 0.1262 | No |
| 38 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 9927 | -0.075 | 0.1276 | No |
| 39 | HBP1 | HBP1 Entrez, Source | HMG-box transcription factor 1 | 11489 | -0.129 | 0.0238 | No |
| 40 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 12434 | -0.191 | -0.0271 | No |
| 41 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 12946 | -0.276 | -0.0365 | No |
| 42 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 13031 | -0.302 | -0.0110 | No |
| 43 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 13097 | -0.326 | 0.0184 | No |