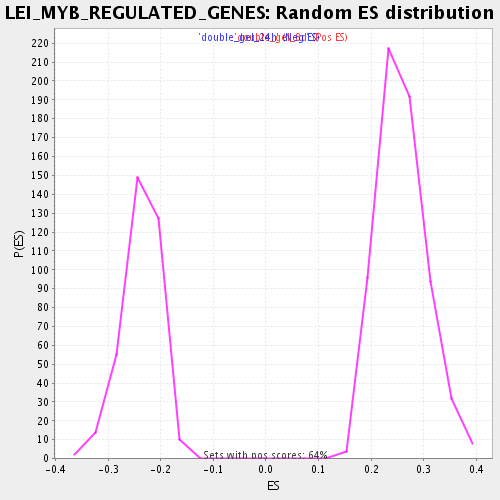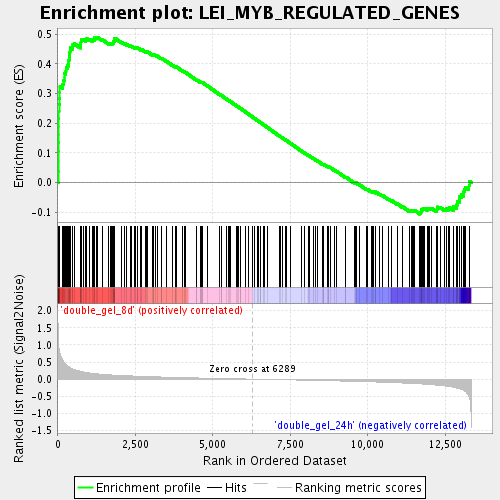
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | LEI_MYB_REGULATED_GENES |
| Enrichment Score (ES) | 0.4900807 |
| Normalized Enrichment Score (NES) | 1.901932 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.01243502 |
| FWER p-Value | 0.525 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 7 | 1.391 | 0.0369 | Yes |
| 2 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 11 | 1.276 | 0.0711 | Yes |
| 3 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 12 | 1.254 | 0.1048 | Yes |
| 4 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 15 | 1.102 | 0.1344 | Yes |
| 5 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 21 | 1.039 | 0.1619 | Yes |
| 6 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 23 | 1.021 | 0.1894 | Yes |
| 7 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 25 | 0.995 | 0.2161 | Yes |
| 8 | DAB2 | DAB2 Entrez, Source | disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila) | 32 | 0.925 | 0.2405 | Yes |
| 9 | CLDN7 | CLDN7 Entrez, Source | claudin 7 | 36 | 0.888 | 0.2642 | Yes |
| 10 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 50 | 0.810 | 0.2850 | Yes |
| 11 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 62 | 0.756 | 0.3046 | Yes |
| 12 | CD24 | CD24 Entrez, Source | CD24 molecule | 66 | 0.745 | 0.3244 | Yes |
| 13 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 159 | 0.550 | 0.3322 | Yes |
| 14 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 190 | 0.499 | 0.3434 | Yes |
| 15 | PRSS23 | PRSS23 Entrez, Source | protease, serine, 23 | 201 | 0.487 | 0.3557 | Yes |
| 16 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 203 | 0.484 | 0.3687 | Yes |
| 17 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 257 | 0.432 | 0.3763 | Yes |
| 18 | DDR1 | DDR1 Entrez, Source | discoidin domain receptor family, member 1 | 262 | 0.429 | 0.3875 | Yes |
| 19 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 316 | 0.387 | 0.3939 | Yes |
| 20 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 330 | 0.373 | 0.4030 | Yes |
| 21 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 336 | 0.371 | 0.4126 | Yes |
| 22 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 361 | 0.358 | 0.4204 | Yes |
| 23 | LAMA5 | LAMA5 Entrez, Source | laminin, alpha 5 | 368 | 0.352 | 0.4294 | Yes |
| 24 | IFITM2 | IFITM2 Entrez, Source | interferon induced transmembrane protein 2 (1-8D) | 381 | 0.346 | 0.4378 | Yes |
| 25 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 391 | 0.341 | 0.4463 | Yes |
| 26 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 405 | 0.336 | 0.4544 | Yes |
| 27 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 480 | 0.292 | 0.4566 | Yes |
| 28 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 484 | 0.291 | 0.4643 | Yes |
| 29 | POR | POR Entrez, Source | P450 (cytochrome) oxidoreductase | 523 | 0.278 | 0.4688 | Yes |
| 30 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 722 | 0.231 | 0.4600 | Yes |
| 31 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 725 | 0.231 | 0.4661 | Yes |
| 32 | EBP | EBP Entrez, Source | emopamil binding protein (sterol isomerase) | 743 | 0.228 | 0.4709 | Yes |
| 33 | CLDN3 | CLDN3 Entrez, Source | claudin 3 | 745 | 0.228 | 0.4770 | Yes |
| 34 | BAIAP2 | BAIAP2 Entrez, Source | BAI1-associated protein 2 | 768 | 0.222 | 0.4813 | Yes |
| 35 | CD59 | CD59 Entrez, Source | CD59 molecule, complement regulatory protein | 818 | 0.210 | 0.4832 | Yes |
| 36 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 905 | 0.195 | 0.4819 | Yes |
| 37 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 934 | 0.190 | 0.4849 | Yes |
| 38 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 1031 | 0.178 | 0.4824 | Yes |
| 39 | SLC20A1 | SLC20A1 Entrez, Source | solute carrier family 20 (phosphate transporter), member 1 | 1123 | 0.168 | 0.4800 | Yes |
| 40 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 1152 | 0.165 | 0.4823 | Yes |
| 41 | FGFBP1 | FGFBP1 Entrez, Source | fibroblast growth factor binding protein 1 | 1178 | 0.163 | 0.4848 | Yes |
| 42 | MAGED2 | MAGED2 Entrez, Source | melanoma antigen family D, 2 | 1191 | 0.162 | 0.4882 | Yes |
| 43 | FN1 | FN1 Entrez, Source | fibronectin 1 | 1246 | 0.156 | 0.4883 | Yes |
| 44 | TACSTD1 | TACSTD1 Entrez, Source | tumor-associated calcium signal transducer 1 | 1278 | 0.153 | 0.4901 | Yes |
| 45 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1445 | 0.139 | 0.4812 | No |
| 46 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1647 | 0.125 | 0.4693 | No |
| 47 | SERPINB5 | SERPINB5 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 5 | 1692 | 0.123 | 0.4692 | No |
| 48 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 1730 | 0.121 | 0.4697 | No |
| 49 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 1766 | 0.118 | 0.4702 | No |
| 50 | BICD2 | BICD2 Entrez, Source | bicaudal D homolog 2 (Drosophila) | 1794 | 0.117 | 0.4713 | No |
| 51 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1796 | 0.117 | 0.4744 | No |
| 52 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 1801 | 0.117 | 0.4772 | No |
| 53 | TM7SF2 | TM7SF2 Entrez, Source | transmembrane 7 superfamily member 2 | 1828 | 0.115 | 0.4784 | No |
| 54 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 1829 | 0.115 | 0.4815 | No |
| 55 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 1830 | 0.115 | 0.4846 | No |
| 56 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 1837 | 0.115 | 0.4872 | No |
| 57 | GOSR1 | GOSR1 Entrez, Source | golgi SNAP receptor complex member 1 | 2055 | 0.105 | 0.4735 | No |
| 58 | MAOA | MAOA Entrez, Source | monoamine oxidase A | 2159 | 0.101 | 0.4684 | No |
| 59 | FASTK | FASTK Entrez, Source | Fas-activated serine/threonine kinase | 2227 | 0.098 | 0.4660 | No |
| 60 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 2327 | 0.094 | 0.4610 | No |
| 61 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 2376 | 0.093 | 0.4598 | No |
| 62 | TFF1 | TFF1 Entrez, Source | trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) | 2482 | 0.089 | 0.4542 | No |
| 63 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 2496 | 0.088 | 0.4556 | No |
| 64 | TMEM106C | TMEM106C Entrez, Source | transmembrane protein 106C | 2557 | 0.085 | 0.4533 | No |
| 65 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 2561 | 0.085 | 0.4554 | No |
| 66 | DPM2 | DPM2 Entrez, Source | dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit | 2678 | 0.081 | 0.4487 | No |
| 67 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 2686 | 0.081 | 0.4504 | No |
| 68 | MAP1LC3B | MAP1LC3B Entrez, Source | microtubule-associated protein 1 light chain 3 beta | 2840 | 0.076 | 0.4408 | No |
| 69 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 2848 | 0.076 | 0.4423 | No |
| 70 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 2894 | 0.074 | 0.4409 | No |
| 71 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 3039 | 0.070 | 0.4318 | No |
| 72 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 3080 | 0.069 | 0.4306 | No |
| 73 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 3086 | 0.069 | 0.4321 | No |
| 74 | SPINK1 | SPINK1 Entrez, Source | serine peptidase inhibitor, Kazal type 1 | 3138 | 0.068 | 0.4300 | No |
| 75 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 3200 | 0.066 | 0.4271 | No |
| 76 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 3329 | 0.062 | 0.4191 | No |
| 77 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 3344 | 0.061 | 0.4197 | No |
| 78 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 3505 | 0.057 | 0.4090 | No |
| 79 | FLOT1 | FLOT1 Entrez, Source | flotillin 1 | 3693 | 0.052 | 0.3962 | No |
| 80 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 3790 | 0.050 | 0.3903 | No |
| 81 | SERPINB3 | SERPINB3 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 3 | 3795 | 0.050 | 0.3913 | No |
| 82 | ATP6V0D1 | ATP6V0D1 Entrez, Source | ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 | 3820 | 0.050 | 0.3908 | No |
| 83 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4036 | 0.044 | 0.3756 | No |
| 84 | SLC9A3R1 | SLC9A3R1 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 1 | 4072 | 0.043 | 0.3741 | No |
| 85 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 4125 | 0.042 | 0.3713 | No |
| 86 | SLC25A1 | SLC25A1 Entrez, Source | solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 | 4469 | 0.035 | 0.3461 | No |
| 87 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 4488 | 0.034 | 0.3457 | No |
| 88 | TFF3 | TFF3 Entrez, Source | trefoil factor 3 (intestinal) | 4594 | 0.032 | 0.3385 | No |
| 89 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 4595 | 0.032 | 0.3394 | No |
| 90 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 4630 | 0.031 | 0.3377 | No |
| 91 | NDUFV1 | NDUFV1 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 1, 51kDa | 4642 | 0.031 | 0.3377 | No |
| 92 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 4656 | 0.031 | 0.3375 | No |
| 93 | TMEM158 | TMEM158 Entrez, Source | transmembrane protein 158 | 4671 | 0.031 | 0.3373 | No |
| 94 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 4827 | 0.027 | 0.3262 | No |
| 95 | ASNS | ASNS Entrez, Source | asparagine synthetase | 5217 | 0.020 | 0.2972 | No |
| 96 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 5264 | 0.019 | 0.2942 | No |
| 97 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 5285 | 0.019 | 0.2932 | No |
| 98 | CLU | CLU Entrez, Source | clusterin | 5443 | 0.016 | 0.2817 | No |
| 99 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 5502 | 0.015 | 0.2776 | No |
| 100 | LGALS3BP | LGALS3BP Entrez, Source | lectin, galactoside-binding, soluble, 3 binding protein | 5525 | 0.014 | 0.2764 | No |
| 101 | PPIB | PPIB Entrez, Source | peptidylprolyl isomerase B (cyclophilin B) | 5568 | 0.013 | 0.2735 | No |
| 102 | GRIP2 | GRIP2 Entrez, Source | glutamate receptor interacting protein 2 | 5756 | 0.010 | 0.2596 | No |
| 103 | ACBD3 | ACBD3 Entrez, Source | acyl-Coenzyme A binding domain containing 3 | 5799 | 0.009 | 0.2566 | No |
| 104 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 5816 | 0.009 | 0.2556 | No |
| 105 | PFKL | PFKL Entrez, Source | phosphofructokinase, liver | 5889 | 0.007 | 0.2503 | No |
| 106 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 5900 | 0.007 | 0.2498 | No |
| 107 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 6062 | 0.004 | 0.2376 | No |
| 108 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 6151 | 0.002 | 0.2310 | No |
| 109 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 6293 | -0.000 | 0.2203 | No |
| 110 | LASP1 | LASP1 Entrez, Source | LIM and SH3 protein 1 | 6351 | -0.001 | 0.2159 | No |
| 111 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 6435 | -0.002 | 0.2097 | No |
| 112 | EMP2 | EMP2 Entrez, Source | epithelial membrane protein 2 | 6473 | -0.003 | 0.2070 | No |
| 113 | HSPD1 | HSPD1 Entrez, Source | heat shock 60kDa protein 1 (chaperonin) | 6540 | -0.004 | 0.2021 | No |
| 114 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 6550 | -0.005 | 0.2015 | No |
| 115 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 6630 | -0.006 | 0.1957 | No |
| 116 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 6652 | -0.006 | 0.1942 | No |
| 117 | MT1A | MT1A Entrez, Source | metallothionein 1A (functional) | 6765 | -0.008 | 0.1859 | No |
| 118 | ACTR2 | ACTR2 Entrez, Source | ARP2 actin-related protein 2 homolog (yeast) | 7164 | -0.015 | 0.1561 | No |
| 119 | CLEC11A | CLEC11A Entrez, Source | C-type lectin domain family 11, member A | 7186 | -0.016 | 0.1549 | No |
| 120 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7236 | -0.017 | 0.1516 | No |
| 121 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 7341 | -0.019 | 0.1442 | No |
| 122 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 7366 | -0.019 | 0.1429 | No |
| 123 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 7503 | -0.022 | 0.1331 | No |
| 124 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 7863 | -0.028 | 0.1066 | No |
| 125 | MRPL49 | MRPL49 Entrez, Source | mitochondrial ribosomal protein L49 | 7943 | -0.030 | 0.1014 | No |
| 126 | SH2B3 | SH2B3 Entrez, Source | SH2B adaptor protein 3 | 8077 | -0.033 | 0.0921 | No |
| 127 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 8108 | -0.034 | 0.0907 | No |
| 128 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 8233 | -0.036 | 0.0823 | No |
| 129 | MDK | MDK Entrez, Source | midkine (neurite growth-promoting factor 2) | 8318 | -0.038 | 0.0769 | No |
| 130 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 8374 | -0.039 | 0.0738 | No |
| 131 | UBC | UBC Entrez, Source | ubiquitin C | 8543 | -0.042 | 0.0621 | No |
| 132 | EPOR | EPOR Entrez, Source | erythropoietin receptor | 8567 | -0.043 | 0.0615 | No |
| 133 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 8582 | -0.043 | 0.0616 | No |
| 134 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8685 | -0.045 | 0.0551 | No |
| 135 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 8722 | -0.046 | 0.0536 | No |
| 136 | ARF6 | ARF6 Entrez, Source | ADP-ribosylation factor 6 | 8733 | -0.046 | 0.0541 | No |
| 137 | GRN | GRN Entrez, Source | granulin | 8784 | -0.047 | 0.0515 | No |
| 138 | LUM | LUM Entrez, Source | lumican | 8919 | -0.050 | 0.0427 | No |
| 139 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 8999 | -0.052 | 0.0381 | No |
| 140 | MUC1 | MUC1 Entrez, Source | mucin 1, cell surface associated | 9275 | -0.058 | 0.0187 | No |
| 141 | HSPE1 | HSPE1 Entrez, Source | heat shock 10kDa protein 1 (chaperonin 10) | 9296 | -0.058 | 0.0187 | No |
| 142 | EFNA3 | EFNA3 Entrez, Source | ephrin-A3 | 9558 | -0.064 | 0.0006 | No |
| 143 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 9615 | -0.066 | -0.0019 | No |
| 144 | GPR56 | GPR56 Entrez, Source | G protein-coupled receptor 56 | 9622 | -0.066 | -0.0006 | No |
| 145 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 9721 | -0.069 | -0.0062 | No |
| 146 | JUP | JUP Entrez, Source | junction plakoglobin | 9963 | -0.075 | -0.0225 | No |
| 147 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 9995 | -0.076 | -0.0228 | No |
| 148 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10112 | -0.079 | -0.0295 | No |
| 149 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 10156 | -0.080 | -0.0306 | No |
| 150 | SNRPN /// SNURF | 10197 | -0.082 | -0.0314 | No | ||
| 151 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 10248 | -0.083 | -0.0330 | No |
| 152 | DNAJA1 | DNAJA1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 1 | 10249 | -0.083 | -0.0308 | No |
| 153 | RPS15A | RPS15A Entrez, Source | ribosomal protein S15a | 10391 | -0.087 | -0.0391 | No |
| 154 | FAP | FAP Entrez, Source | fibroblast activation protein, alpha | 10470 | -0.089 | -0.0427 | No |
| 155 | SIAHBP1 | SIAHBP1 Entrez, Source | - | 10678 | -0.096 | -0.0558 | No |
| 156 | AP2A2 | AP2A2 Entrez, Source | adaptor-related protein complex 2, alpha 2 subunit | 10778 | -0.100 | -0.0607 | No |
| 157 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 10955 | -0.107 | -0.0712 | No |
| 158 | EEF1A2 | EEF1A2 Entrez, Source | eukaryotic translation elongation factor 1 alpha 2 | 11120 | -0.113 | -0.0806 | No |
| 159 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 11359 | -0.123 | -0.0954 | No |
| 160 | SERBP1 | SERBP1 Entrez, Source | SERPINE1 mRNA binding protein 1 | 11399 | -0.125 | -0.0950 | No |
| 161 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | -0.0953 | No |
| 162 | NXF1 | NXF1 Entrez, Source | nuclear RNA export factor 1 | 11487 | -0.128 | -0.0947 | No |
| 163 | LY6G6C | LY6G6C Entrez, Source | lymphocyte antigen 6 complex, locus G6C | 11508 | -0.129 | -0.0927 | No |
| 164 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 11683 | -0.137 | -0.1023 | No |
| 165 | IL1RN | IL1RN Entrez, Source | interleukin 1 receptor antagonist | 11709 | -0.139 | -0.1005 | No |
| 166 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 11721 | -0.139 | -0.0975 | No |
| 167 | NRD1 | NRD1 Entrez, Source | nardilysin (N-arginine dibasic convertase) | 11728 | -0.140 | -0.0942 | No |
| 168 | EIF3S9 | EIF3S9 Entrez, Source | eukaryotic translation initiation factor 3, subunit 9 eta, 116kDa | 11737 | -0.140 | -0.0911 | No |
| 169 | EEF1D | EEF1D Entrez, Source | eukaryotic translation elongation factor 1 delta (guanine nucleotide exchange protein) | 11758 | -0.142 | -0.0888 | No |
| 170 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 11793 | -0.144 | -0.0875 | No |
| 171 | BLCAP | BLCAP Entrez, Source | bladder cancer associated protein | 11823 | -0.146 | -0.0858 | No |
| 172 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 11943 | -0.153 | -0.0907 | No |
| 173 | AURKB | AURKB Entrez, Source | aurora kinase B | 11946 | -0.153 | -0.0867 | No |
| 174 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 12008 | -0.158 | -0.0871 | No |
| 175 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 12067 | -0.162 | -0.0872 | No |
| 176 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 12220 | -0.173 | -0.0941 | No |
| 177 | ANP32B | ANP32B Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member B | 12233 | -0.174 | -0.0903 | No |
| 178 | AURKA | AURKA Entrez, Source | aurora kinase A | 12238 | -0.175 | -0.0859 | No |
| 179 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 12249 | -0.175 | -0.0819 | No |
| 180 | MSN | MSN Entrez, Source | moesin | 12335 | -0.182 | -0.0835 | No |
| 181 | GSTO1 | GSTO1 Entrez, Source | glutathione S-transferase omega 1 | 12484 | -0.197 | -0.0894 | No |
| 182 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 12528 | -0.202 | -0.0873 | No |
| 183 | XBP1 | XBP1 Entrez, Source | X-box binding protein 1 | 12599 | -0.210 | -0.0869 | No |
| 184 | PPID | PPID Entrez, Source | peptidylprolyl isomerase D (cyclophilin D) | 12627 | -0.215 | -0.0832 | No |
| 185 | STIP1 | STIP1 Entrez, Source | stress-induced-phosphoprotein 1 (Hsp70/Hsp90-organizing protein) | 12759 | -0.234 | -0.0869 | No |
| 186 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12761 | -0.235 | -0.0806 | No |
| 187 | TUBB | TUBB Entrez, Source | tubulin, beta | 12860 | -0.256 | -0.0812 | No |
| 188 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 12874 | -0.260 | -0.0752 | No |
| 189 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 12884 | -0.262 | -0.0688 | No |
| 190 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 12892 | -0.265 | -0.0622 | No |
| 191 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 12951 | -0.277 | -0.0592 | No |
| 192 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 12967 | -0.282 | -0.0527 | No |
| 193 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12971 | -0.283 | -0.0453 | No |
| 194 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 13021 | -0.298 | -0.0410 | No |
| 195 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 13080 | -0.318 | -0.0369 | No |
| 196 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13101 | -0.328 | -0.0296 | No |
| 197 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 13125 | -0.341 | -0.0222 | No |
| 198 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 13159 | -0.370 | -0.0147 | No |
| 199 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 13267 | -0.519 | -0.0089 | No |
| 200 | CRIP2 | CRIP2 Entrez, Source | cysteine-rich protein 2 | 13280 | -0.538 | 0.0047 | No |