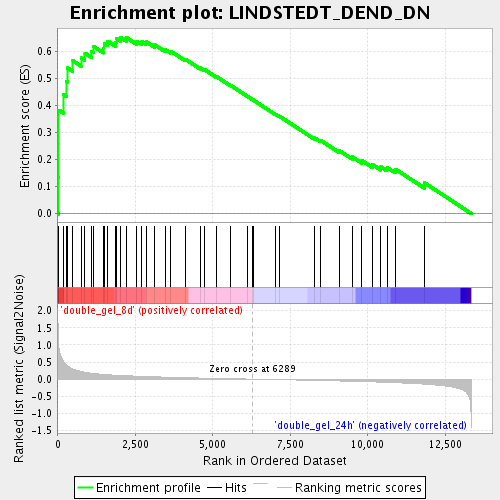
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | LINDSTEDT_DEND_DN |
| Enrichment Score (ES) | 0.6515983 |
| Normalized Enrichment Score (NES) | 2.0141325 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0044725384 |
| FWER p-Value | 0.118 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 25 | 0.995 | 0.1321 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 31 | 0.935 | 0.2576 | Yes |
| 3 | DAB2 | DAB2 Entrez, Source | disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila) | 32 | 0.925 | 0.3822 | Yes |
| 4 | F13A1 | F13A1 Entrez, Source | coagulation factor XIII, A1 polypeptide | 186 | 0.510 | 0.4394 | Yes |
| 5 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 279 | 0.415 | 0.4883 | Yes |
| 6 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.5384 | Yes |
| 7 | HLA-DMB | HLA-DMB Entrez, Source | major histocompatibility complex, class II, DM beta | 483 | 0.292 | 0.5649 | Yes |
| 8 | IFNGR1 | IFNGR1 Entrez, Source | interferon gamma receptor 1 | 750 | 0.226 | 0.5754 | Yes |
| 9 | IGSF6 | IGSF6 Entrez, Source | immunoglobulin superfamily, member 6 | 872 | 0.199 | 0.5931 | Yes |
| 10 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 1082 | 0.173 | 0.6007 | Yes |
| 11 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1157 | 0.164 | 0.6172 | Yes |
| 12 | ACP5 | ACP5 Entrez, Source | acid phosphatase 5, tartrate resistant | 1475 | 0.137 | 0.6118 | Yes |
| 13 | SLC1A5 | SLC1A5 Entrez, Source | solute carrier family 1 (neutral amino acid transporter), member 5 | 1487 | 0.136 | 0.6293 | Yes |
| 14 | ITGAM | ITGAM Entrez, Source | integrin, alpha M (complement component 3 receptor 3 subunit) | 1608 | 0.128 | 0.6375 | Yes |
| 15 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 1866 | 0.113 | 0.6334 | Yes |
| 16 | FCGRT | FCGRT Entrez, Source | Fc fragment of IgG, receptor, transporter, alpha | 1897 | 0.112 | 0.6462 | Yes |
| 17 | SLC6A2 | SLC6A2 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, noradrenalin), member 2 | 2018 | 0.107 | 0.6516 | Yes |
| 18 | ALOX15 | ALOX15 Entrez, Source | arachidonate 15-lipoxygenase | 2217 | 0.099 | 0.6500 | No |
| 19 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 2541 | 0.086 | 0.6373 | No |
| 20 | RCBTB2 | RCBTB2 Entrez, Source | regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 | 2710 | 0.080 | 0.6354 | No |
| 21 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 2848 | 0.076 | 0.6353 | No |
| 22 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 3128 | 0.068 | 0.6234 | No |
| 23 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 3460 | 0.059 | 0.6065 | No |
| 24 | MAN2B1 | MAN2B1 Entrez, Source | mannosidase, alpha, class 2B, member 1 | 3637 | 0.054 | 0.6005 | No |
| 25 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 4125 | 0.042 | 0.5695 | No |
| 26 | FUT1 | FUT1 Entrez, Source | fucosyltransferase 1 (galactoside 2-alpha-L-fucosyltransferase, H blood group) | 4598 | 0.032 | 0.5383 | No |
| 27 | LST1 | LST1 Entrez, Source | leukocyte specific transcript 1 | 4729 | 0.030 | 0.5325 | No |
| 28 | RAP1GAP | RAP1GAP Entrez, Source | RAP1 GTPase activating protein | 5106 | 0.022 | 0.5072 | No |
| 29 | P2RY6 | P2RY6 Entrez, Source | pyrimidinergic receptor P2Y, G-protein coupled, 6 | 5559 | 0.013 | 0.4750 | No |
| 30 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 6122 | 0.003 | 0.4332 | No |
| 31 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 6293 | -0.000 | 0.4204 | No |
| 32 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 6298 | -0.000 | 0.4201 | No |
| 33 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 7017 | -0.013 | 0.3678 | No |
| 34 | CCL22 | CCL22 Entrez, Source | chemokine (C-C motif) ligand 22 | 7142 | -0.015 | 0.3605 | No |
| 35 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 8284 | -0.037 | 0.2797 | No |
| 36 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 8474 | -0.041 | 0.2710 | No |
| 37 | NPR1 | NPR1 Entrez, Source | natriuretic peptide receptor A/guanylate cyclase A (atrionatriuretic peptide receptor A) | 9076 | -0.053 | 0.2330 | No |
| 38 | AQP3 | AQP3 Entrez, Source | aquaporin 3 (Gill blood group) | 9501 | -0.063 | 0.2096 | No |
| 39 | LAPTM4A | LAPTM4A Entrez, Source | lysosomal-associated protein transmembrane 4 alpha | 9810 | -0.071 | 0.1960 | No |
| 40 | ITGB2 | ITGB2 Entrez, Source | integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) | 10160 | -0.081 | 0.1806 | No |
| 41 | QSCN6 | QSCN6 Entrez, Source | quiescin Q6 | 10424 | -0.088 | 0.1727 | No |
| 42 | CD99 | CD99 Entrez, Source | CD99 molecule | 10623 | -0.094 | 0.1705 | No |
| 43 | CCNH | CCNH Entrez, Source | cyclin H | 10890 | -0.105 | 0.1646 | No |
| 44 | PTGER3 | PTGER3 Entrez, Source | prostaglandin E receptor 3 (subtype EP3) | 11830 | -0.146 | 0.1137 | No |

