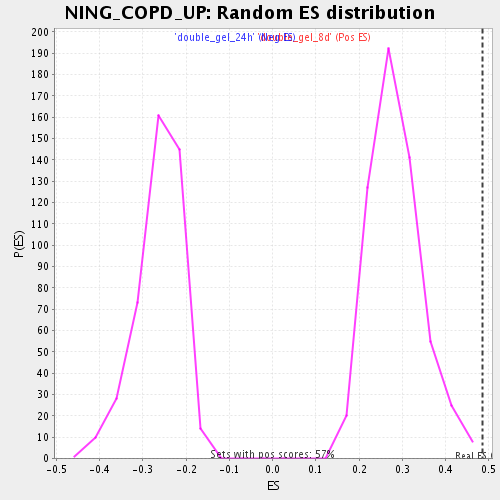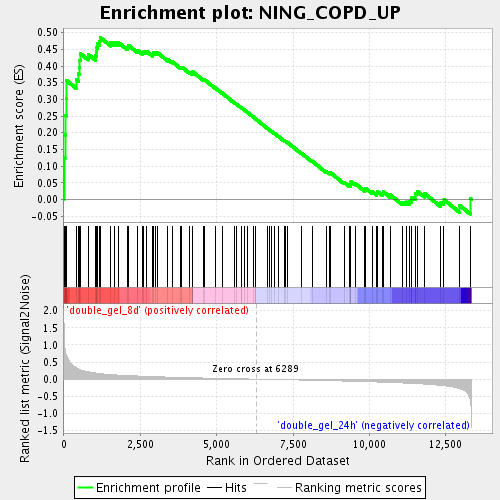
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | NING_COPD_UP |
| Enrichment Score (ES) | 0.4862242 |
| Normalized Enrichment Score (NES) | 1.7086638 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.05257883 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.1267 | Yes |
| 2 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 37 | 0.888 | 0.1936 | Yes |
| 3 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 60 | 0.762 | 0.2516 | Yes |
| 4 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 82 | 0.687 | 0.3037 | Yes |
| 5 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 89 | 0.680 | 0.3564 | Yes |
| 6 | CDC42EP2 | CDC42EP2 Entrez, Source | CDC42 effector protein (Rho GTPase binding) 2 | 399 | 0.337 | 0.3595 | Yes |
| 7 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 476 | 0.294 | 0.3767 | Yes |
| 8 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 507 | 0.282 | 0.3966 | Yes |
| 9 | RAMP3 | RAMP3 Entrez, Source | receptor (calcitonin) activity modifying protein 3 | 518 | 0.280 | 0.4177 | Yes |
| 10 | RARRES2 | RARRES2 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 2 | 544 | 0.271 | 0.4370 | Yes |
| 11 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 796 | 0.215 | 0.4348 | Yes |
| 12 | TLE2 | TLE2 Entrez, Source | transducin-like enhancer of split 2 (E(sp1) homolog, Drosophila) | 1029 | 0.179 | 0.4313 | Yes |
| 13 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.4425 | Yes |
| 14 | FEZ2 | FEZ2 Entrez, Source | fasciculation and elongation protein zeta 2 (zygin II) | 1075 | 0.173 | 0.4552 | Yes |
| 15 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 1103 | 0.170 | 0.4665 | Yes |
| 16 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 1163 | 0.164 | 0.4749 | Yes |
| 17 | WFS1 | WFS1 Entrez, Source | Wolfram syndrome 1 (wolframin) | 1182 | 0.163 | 0.4862 | Yes |
| 18 | MAF1 | MAF1 Entrez, Source | MAF1 homolog (S. cerevisiae) | 1532 | 0.133 | 0.4703 | No |
| 19 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 1666 | 0.124 | 0.4699 | No |
| 20 | VAMP8 | VAMP8 Entrez, Source | vesicle-associated membrane protein 8 (endobrevin) | 1782 | 0.118 | 0.4705 | No |
| 21 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 2076 | 0.105 | 0.4565 | No |
| 22 | FASN | FASN Entrez, Source | fatty acid synthase | 2099 | 0.104 | 0.4630 | No |
| 23 | VWF | VWF Entrez, Source | von Willebrand factor | 2400 | 0.092 | 0.4475 | No |
| 24 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 2564 | 0.085 | 0.4419 | No |
| 25 | APOE | APOE Entrez, Source | apolipoprotein E | 2619 | 0.083 | 0.4443 | No |
| 26 | GPX4 | GPX4 Entrez, Source | glutathione peroxidase 4 (phospholipid hydroperoxidase) | 2713 | 0.080 | 0.4435 | No |
| 27 | INPP1 | INPP1 Entrez, Source | inositol polyphosphate-1-phosphatase | 2901 | 0.074 | 0.4352 | No |
| 28 | PNRC1 | PNRC1 Entrez, Source | proline-rich nuclear receptor coactivator 1 | 2915 | 0.074 | 0.4400 | No |
| 29 | NBL1 | NBL1 Entrez, Source | neuroblastoma, suppression of tumorigenicity 1 | 2982 | 0.072 | 0.4406 | No |
| 30 | MAPKAPK3 | MAPKAPK3 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 3 | 3069 | 0.069 | 0.4395 | No |
| 31 | CSK | CSK Entrez, Source | c-src tyrosine kinase | 3403 | 0.060 | 0.4191 | No |
| 32 | CTSF | CTSF Entrez, Source | cathepsin F | 3540 | 0.056 | 0.4133 | No |
| 33 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 3822 | 0.050 | 0.3959 | No |
| 34 | GTF2I | GTF2I Entrez, Source | general transcription factor II, i | 3859 | 0.048 | 0.3970 | No |
| 35 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 4125 | 0.042 | 0.3803 | No |
| 36 | G3BP | G3BP Entrez, Source | - | 4201 | 0.039 | 0.3777 | No |
| 37 | CAPNS1 | CAPNS1 Entrez, Source | calpain, small subunit 1 | 4212 | 0.039 | 0.3800 | No |
| 38 | MTCH1 | MTCH1 Entrez, Source | mitochondrial carrier homolog 1 (C. elegans) | 4213 | 0.039 | 0.3831 | No |
| 39 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 4578 | 0.032 | 0.3582 | No |
| 40 | TFF3 | TFF3 Entrez, Source | trefoil factor 3 (intestinal) | 4594 | 0.032 | 0.3595 | No |
| 41 | A2M | A2M Entrez, Source | alpha-2-macroglobulin | 4951 | 0.025 | 0.3346 | No |
| 42 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 5173 | 0.021 | 0.3196 | No |
| 43 | CPA3 | CPA3 Entrez, Source | carboxypeptidase A3 (mast cell) | 5582 | 0.013 | 0.2898 | No |
| 44 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 5658 | 0.012 | 0.2851 | No |
| 45 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 5816 | 0.009 | 0.2739 | No |
| 46 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 5827 | 0.009 | 0.2738 | No |
| 47 | ARCN1 | ARCN1 Entrez, Source | archain 1 | 5911 | 0.007 | 0.2681 | No |
| 48 | NDUFA5 | NDUFA5 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5, 13kDa | 6009 | 0.005 | 0.2612 | No |
| 49 | EGFL7 | EGFL7 Entrez, Source | EGF-like-domain, multiple 7 | 6196 | 0.001 | 0.2472 | No |
| 50 | PNMT | PNMT Entrez, Source | phenylethanolamine N-methyltransferase | 6269 | 0.000 | 0.2418 | No |
| 51 | NCKAP1 | NCKAP1 Entrez, Source | NCK-associated protein 1 | 6284 | 0.000 | 0.2408 | No |
| 52 | DVL1 | DVL1 Entrez, Source | dishevelled, dsh homolog 1 (Drosophila) | 6668 | -0.007 | 0.2124 | No |
| 53 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 6718 | -0.007 | 0.2093 | No |
| 54 | MSLN | MSLN Entrez, Source | mesothelin | 6799 | -0.009 | 0.2039 | No |
| 55 | HIF3A | HIF3A Entrez, Source | hypoxia inducible factor 3, alpha subunit | 6901 | -0.011 | 0.1971 | No |
| 56 | CDC2L1 | CDC2L1 Entrez, Source | cell division cycle 2-like 1 (PITSLRE proteins) | 7029 | -0.013 | 0.1886 | No |
| 57 | DCN | DCN Entrez, Source | decorin | 7217 | -0.016 | 0.1757 | No |
| 58 | NPC2 | NPC2 Entrez, Source | Niemann-Pick disease, type C2 | 7253 | -0.017 | 0.1744 | No |
| 59 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 7323 | -0.018 | 0.1706 | No |
| 60 | RPLP1 | RPLP1 Entrez, Source | ribosomal protein, large, P1 | 7767 | -0.027 | 0.1393 | No |
| 61 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 8120 | -0.034 | 0.1154 | No |
| 62 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 8582 | -0.043 | 0.0839 | No |
| 63 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8685 | -0.045 | 0.0798 | No |
| 64 | BCAP31 | BCAP31 Entrez, Source | B-cell receptor-associated protein 31 | 8715 | -0.046 | 0.0812 | No |
| 65 | RPS15 | RPS15 Entrez, Source | ribosomal protein S15 | 9166 | -0.055 | 0.0515 | No |
| 66 | ACTN4 | ACTN4 Entrez, Source | actinin, alpha 4 | 9335 | -0.059 | 0.0435 | No |
| 67 | BLVRA | BLVRA Entrez, Source | biliverdin reductase A | 9379 | -0.060 | 0.0449 | No |
| 68 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 9390 | -0.060 | 0.0489 | No |
| 69 | RPL32 | RPL32 Entrez, Source | ribosomal protein L32 | 9393 | -0.060 | 0.0534 | No |
| 70 | GRINA | GRINA Entrez, Source | glutamate receptor, ionotropic, N-methyl D-asparate-associated protein 1 (glutamate binding) | 9530 | -0.063 | 0.0481 | No |
| 71 | FGG | FGG Entrez, Source | fibrinogen gamma chain | 9841 | -0.072 | 0.0304 | No |
| 72 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 9875 | -0.073 | 0.0336 | No |
| 73 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 10089 | -0.079 | 0.0237 | No |
| 74 | RPL4 | RPL4 Entrez, Source | ribosomal protein L4 | 10242 | -0.083 | 0.0187 | No |
| 75 | RPL13 | RPL13 Entrez, Source | ribosomal protein L13 | 10259 | -0.083 | 0.0240 | No |
| 76 | RPS23 | RPS23 Entrez, Source | ribosomal protein S23 | 10436 | -0.088 | 0.0176 | No |
| 77 | C1R | C1R Entrez, Source | complement component 1, r subcomponent | 10455 | -0.089 | 0.0233 | No |
| 78 | CALM1 | CALM1 Entrez, Source | calmodulin 1 (phosphorylase kinase, delta) | 10674 | -0.096 | 0.0143 | No |
| 79 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 11080 | -0.111 | -0.0075 | No |
| 80 | RABGGTB | RABGGTB Entrez, Source | Rab geranylgeranyltransferase, beta subunit | 11200 | -0.116 | -0.0074 | No |
| 81 | DDX52 | DDX52 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 52 | 11297 | -0.120 | -0.0053 | No |
| 82 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 11364 | -0.123 | -0.0006 | No |
| 83 | RPL22 | RPL22 Entrez, Source | ribosomal protein L22 | 11388 | -0.124 | 0.0073 | No |
| 84 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 11506 | -0.129 | 0.0086 | No |
| 85 | RPS3 | RPS3 Entrez, Source | ribosomal protein S3 | 11507 | -0.129 | 0.0187 | No |
| 86 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 11560 | -0.132 | 0.0251 | No |
| 87 | RPL14 | RPL14 Entrez, Source | ribosomal protein L14 | 11799 | -0.144 | 0.0184 | No |
| 88 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 12334 | -0.182 | -0.0076 | No |
| 89 | SDC1 | SDC1 Entrez, Source | syndecan 1 | 12438 | -0.192 | -0.0004 | No |
| 90 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 12940 | -0.274 | -0.0168 | No |
| 91 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13299 | -0.601 | 0.0032 | No |