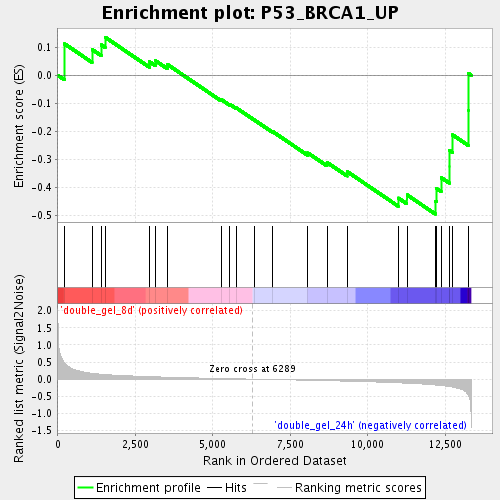
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | P53_BRCA1_UP |
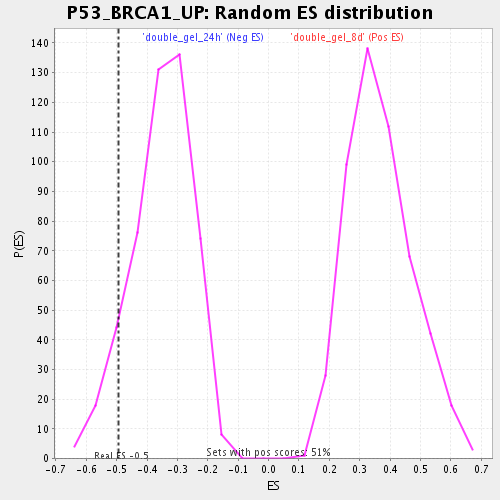
| Enrichment Score (ES) | -0.49562857 |
| Normalized Enrichment Score (NES) | -1.4021006 |
| Nominal p-value | 0.087576374 |
| FDR q-value | 0.29164943 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 216 | 0.473 | 0.1126 | No |
| 2 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 1103 | 0.170 | 0.0926 | No |
| 3 | CTSH | CTSH Entrez, Source | cathepsin H | 1401 | 0.143 | 0.1092 | No |
| 4 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 1525 | 0.133 | 0.1362 | No |
| 5 | ENPP2 | ENPP2 Entrez, Source | ectonucleotide pyrophosphatase/phosphodiesterase 2 (autotaxin) | 2952 | 0.072 | 0.0489 | No |
| 6 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 3153 | 0.067 | 0.0522 | No |
| 7 | CTSF | CTSF Entrez, Source | cathepsin F | 3540 | 0.056 | 0.0385 | No |
| 8 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 5270 | 0.019 | -0.0861 | No |
| 9 | COX6B2 | COX6B2 Entrez, Source | cytochrome c oxidase subunit VIb polypeptide 2 (testis) | 5546 | 0.014 | -0.1030 | No |
| 10 | FXYD3 | FXYD3 Entrez, Source | FXYD domain containing ion transport regulator 3 | 5757 | 0.010 | -0.1160 | No |
| 11 | PTPRV | PTPRV Entrez, Source | protein tyrosine phosphatase, receptor type, V (pseudogene) | 6358 | -0.001 | -0.1608 | No |
| 12 | ROBO1 | ROBO1 Entrez, Source | roundabout, axon guidance receptor, homolog 1 (Drosophila) | 6932 | -0.011 | -0.2008 | No |
| 13 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 8065 | -0.033 | -0.2769 | No |
| 14 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8685 | -0.045 | -0.3111 | No |
| 15 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 9332 | -0.059 | -0.3436 | No |
| 16 | PHLDA3 | PHLDA3 Entrez, Source | pleckstrin homology-like domain, family A, member 3 | 10983 | -0.108 | -0.4381 | No |
| 17 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 11266 | -0.119 | -0.4268 | No |
| 18 | PQLC3 | PQLC3 Entrez, Source | PQ loop repeat containing 3 | 12183 | -0.170 | -0.4492 | Yes |
| 19 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 12217 | -0.173 | -0.4046 | Yes |
| 20 | EPHX1 | EPHX1 Entrez, Source | epoxide hydrolase 1, microsomal (xenobiotic) | 12370 | -0.186 | -0.3654 | Yes |
| 21 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 12632 | -0.215 | -0.3263 | Yes |
| 22 | APOBEC1 | APOBEC1 Entrez, Source | apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 | 12633 | -0.215 | -0.2676 | Yes |
| 23 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 12738 | -0.231 | -0.2125 | Yes |
| 24 | SRXN1 | SRXN1 Entrez, Source | sulfiredoxin 1 homolog (S. cerevisiae) | 13244 | -0.464 | -0.1241 | Yes |
| 25 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 13250 | -0.482 | 0.0069 | Yes |