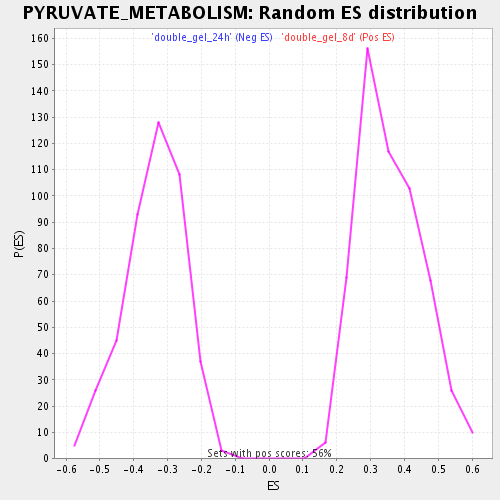
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | PYRUVATE_METABOLISM |
| Enrichment Score (ES) | 0.6590275 |
| Normalized Enrichment Score (NES) | 1.8414096 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.020095544 |
| FWER p-Value | 0.812 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LDHB | LDHB Entrez, Source | lactate dehydrogenase B | 15 | 1.102 | 0.2131 | Yes |
| 2 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 127 | 0.610 | 0.3233 | Yes |
| 3 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.4182 | Yes |
| 4 | PC | PC Entrez, Source | pyruvate carboxylase | 322 | 0.380 | 0.4820 | Yes |
| 5 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 411 | 0.331 | 0.5397 | Yes |
| 6 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 526 | 0.277 | 0.5849 | Yes |
| 7 | PDHA2 | PDHA2 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 2 | 1026 | 0.179 | 0.5823 | Yes |
| 8 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 1343 | 0.147 | 0.5872 | Yes |
| 9 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 1428 | 0.140 | 0.6082 | Yes |
| 10 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 1482 | 0.136 | 0.6307 | Yes |
| 11 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1647 | 0.125 | 0.6428 | Yes |
| 12 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 2249 | 0.097 | 0.6166 | Yes |
| 13 | LDHD | LDHD Entrez, Source | lactate dehydrogenase D | 2272 | 0.096 | 0.6337 | Yes |
| 14 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 2544 | 0.086 | 0.6300 | Yes |
| 15 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 2674 | 0.081 | 0.6361 | Yes |
| 16 | MDH2 | MDH2 Entrez, Source | malate dehydrogenase 2, NAD (mitochondrial) | 2860 | 0.075 | 0.6369 | Yes |
| 17 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 2893 | 0.074 | 0.6490 | Yes |
| 18 | ACACA | ACACA Entrez, Source | acetyl-Coenzyme A carboxylase alpha | 2948 | 0.073 | 0.6590 | Yes |
| 19 | HAGH | HAGH Entrez, Source | hydroxyacylglutathione hydrolase | 3639 | 0.054 | 0.6177 | No |
| 20 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 3710 | 0.052 | 0.6225 | No |
| 21 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 3755 | 0.051 | 0.6291 | No |
| 22 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 4375 | 0.036 | 0.5897 | No |
| 23 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 6088 | 0.003 | 0.4618 | No |
| 24 | GLO1 | GLO1 Entrez, Source | glyoxalase I | 7131 | -0.014 | 0.3863 | No |
| 25 | HAGHL | HAGHL Entrez, Source | hydroxyacylglutathione hydrolase-like | 9367 | -0.060 | 0.2301 | No |
| 26 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 9934 | -0.075 | 0.2021 | No |
| 27 | LDHC | LDHC Entrez, Source | lactate dehydrogenase C | 10001 | -0.077 | 0.2120 | No |
| 28 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 12501 | -0.200 | 0.0632 | No |