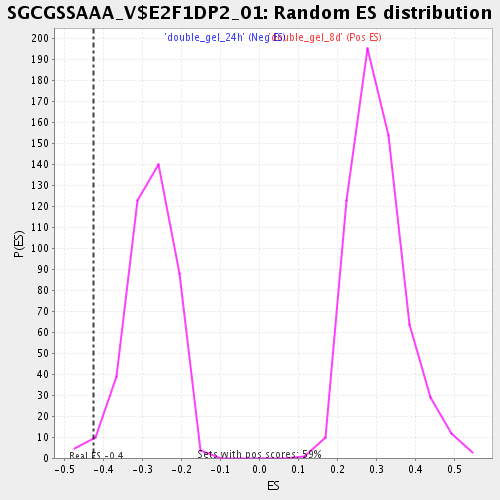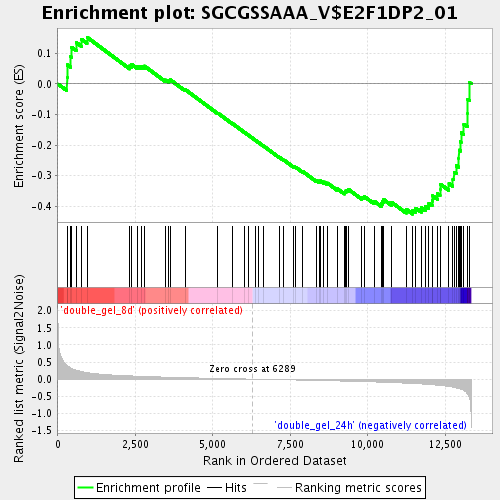
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | SGCGSSAAA_V$E2F1DP2_01 |
| Enrichment Score (ES) | -0.42547062 |
| Normalized Enrichment Score (NES) | -1.5160028 |
| Nominal p-value | 0.017114915 |
| FDR q-value | 0.22194488 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 293 | 0.407 | 0.0207 | No |
| 2 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 300 | 0.402 | 0.0625 | No |
| 3 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 412 | 0.330 | 0.0889 | No |
| 4 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 442 | 0.313 | 0.1195 | No |
| 5 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 597 | 0.258 | 0.1350 | No |
| 6 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 764 | 0.223 | 0.1459 | No |
| 7 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 940 | 0.189 | 0.1526 | No |
| 8 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 2319 | 0.095 | 0.0587 | No |
| 9 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 2376 | 0.093 | 0.0643 | No |
| 10 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 2578 | 0.084 | 0.0580 | No |
| 11 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 2696 | 0.080 | 0.0576 | No |
| 12 | SLC9A5 | SLC9A5 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 5 | 2790 | 0.077 | 0.0587 | No |
| 13 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3479 | 0.058 | 0.0130 | No |
| 14 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 3580 | 0.055 | 0.0113 | No |
| 15 | PRPF4B | PRPF4B Entrez, Source | PRP4 pre-mRNA processing factor 4 homolog B (yeast) | 3623 | 0.054 | 0.0138 | No |
| 16 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 4102 | 0.042 | -0.0178 | No |
| 17 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 5162 | 0.021 | -0.0953 | No |
| 18 | GABRB3 | GABRB3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 3 | 5640 | 0.012 | -0.1300 | No |
| 19 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 6034 | 0.005 | -0.1591 | No |
| 20 | ACBD6 | ACBD6 Entrez, Source | acyl-Coenzyme A binding domain containing 6 | 6149 | 0.002 | -0.1675 | No |
| 21 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 6380 | -0.001 | -0.1847 | No |
| 22 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 6471 | -0.003 | -0.1911 | No |
| 23 | PHF5A | PHF5A Entrez, Source | PHD finger protein 5A | 6646 | -0.006 | -0.2036 | No |
| 24 | KBTBD7 | KBTBD7 Entrez, Source | kelch repeat and BTB (POZ) domain containing 7 | 7148 | -0.015 | -0.2398 | No |
| 25 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7291 | -0.018 | -0.2486 | No |
| 26 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 7606 | -0.024 | -0.2698 | No |
| 27 | EFNA5 | EFNA5 Entrez, Source | ephrin-A5 | 7672 | -0.025 | -0.2721 | No |
| 28 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 7897 | -0.029 | -0.2859 | No |
| 29 | HCN3 | HCN3 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 3 | 8351 | -0.038 | -0.3160 | No |
| 30 | GATA1 | GATA1 Entrez, Source | GATA binding protein 1 (globin transcription factor 1) | 8429 | -0.040 | -0.3176 | No |
| 31 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 8474 | -0.041 | -0.3166 | No |
| 32 | HNRPR | HNRPR Entrez, Source | heterogeneous nuclear ribonucleoprotein R | 8586 | -0.043 | -0.3204 | No |
| 33 | HOXC10 | HOXC10 Entrez, Source | homeobox C10 | 8695 | -0.045 | -0.3238 | No |
| 34 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9009 | -0.052 | -0.3419 | No |
| 35 | FMO4 | FMO4 Entrez, Source | flavin containing monooxygenase 4 | 9249 | -0.057 | -0.3539 | No |
| 36 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 9277 | -0.058 | -0.3499 | No |
| 37 | FANCC | FANCC Entrez, Source | Fanconi anemia, complementation group C | 9324 | -0.059 | -0.3472 | No |
| 38 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 9382 | -0.060 | -0.3452 | No |
| 39 | RET | RET Entrez, Source | ret proto-oncogene (multiple endocrine neoplasia and medullary thyroid carcinoma 1, Hirschsprung disease) | 9808 | -0.071 | -0.3697 | No |
| 40 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 9887 | -0.073 | -0.3679 | No |
| 41 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 10205 | -0.082 | -0.3832 | No |
| 42 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10438 | -0.088 | -0.3914 | No |
| 43 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 10466 | -0.089 | -0.3840 | No |
| 44 | UGCGL1 | UGCGL1 Entrez, Source | UDP-glucose ceramide glucosyltransferase-like 1 | 10515 | -0.091 | -0.3781 | No |
| 45 | RPS20 | RPS20 Entrez, Source | ribosomal protein S20 | 10766 | -0.100 | -0.3865 | No |
| 46 | TRIM39 | TRIM39 Entrez, Source | tripartite motif-containing 39 | 11239 | -0.117 | -0.4097 | No |
| 47 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | -0.4121 | Yes |
| 48 | NUP62 | NUP62 Entrez, Source | nucleoporin 62kDa | 11546 | -0.131 | -0.4056 | Yes |
| 49 | GLRA3 | GLRA3 Entrez, Source | glycine receptor, alpha 3 | 11725 | -0.140 | -0.4043 | Yes |
| 50 | KCNS2 | KCNS2 Entrez, Source | potassium voltage-gated channel, delayed-rectifier, subfamily S, member 2 | 11867 | -0.148 | -0.3994 | Yes |
| 51 | HTF9C | HTF9C Entrez, Source | - | 11970 | -0.155 | -0.3908 | Yes |
| 52 | XTP3TPA | XTP3TPA Entrez, Source | - | 12079 | -0.163 | -0.3818 | Yes |
| 53 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 12095 | -0.164 | -0.3657 | Yes |
| 54 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 12249 | -0.175 | -0.3588 | Yes |
| 55 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 12334 | -0.182 | -0.3460 | Yes |
| 56 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 12339 | -0.183 | -0.3272 | Yes |
| 57 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 12621 | -0.214 | -0.3259 | Yes |
| 58 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 12748 | -0.232 | -0.3110 | Yes |
| 59 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 12783 | -0.238 | -0.2885 | Yes |
| 60 | NCL | NCL Entrez, Source | nucleolin | 12857 | -0.256 | -0.2672 | Yes |
| 61 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 12940 | -0.274 | -0.2445 | Yes |
| 62 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 12945 | -0.275 | -0.2159 | Yes |
| 63 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 12994 | -0.291 | -0.1890 | Yes |
| 64 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 13014 | -0.296 | -0.1593 | Yes |
| 65 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13101 | -0.328 | -0.1314 | Yes |
| 66 | DCK | DCK Entrez, Source | deoxycytidine kinase | 13222 | -0.428 | -0.0954 | Yes |
| 67 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 13227 | -0.440 | -0.0496 | Yes |
| 68 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13288 | -0.553 | 0.0041 | Yes |