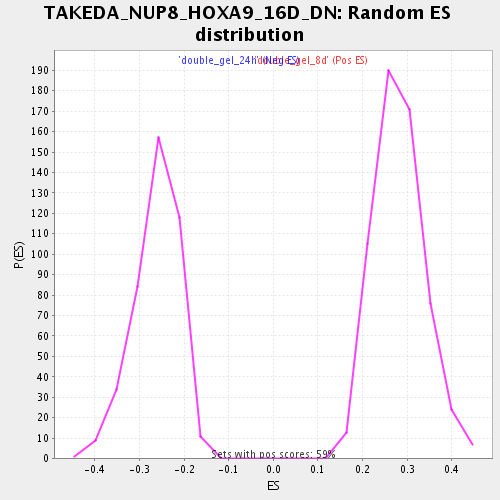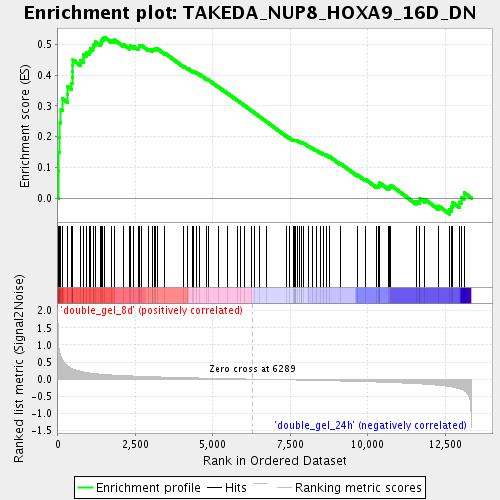
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | TAKEDA_NUP8_HOXA9_16D_DN |
| Enrichment Score (ES) | 0.52369654 |
| Normalized Enrichment Score (NES) | 1.8665956 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.016636593 |
| FWER p-Value | 0.698 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PGLYRP1 | PGLYRP1 Entrez, Source | peptidoglycan recognition protein 1 | 9 | 1.343 | 0.0885 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 31 | 0.935 | 0.1490 | Yes |
| 3 | PROCR | PROCR Entrez, Source | protein C receptor, endothelial (EPCR) | 59 | 0.763 | 0.1976 | Yes |
| 4 | CD24 | CD24 Entrez, Source | CD24 molecule | 66 | 0.745 | 0.2467 | Yes |
| 5 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 98 | 0.666 | 0.2886 | Yes |
| 6 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 140 | 0.582 | 0.3242 | Yes |
| 7 | ARG1 | ARG1 Entrez, Source | arginase, liver | 307 | 0.397 | 0.3380 | Yes |
| 8 | TSPAN2 | TSPAN2 Entrez, Source | tetraspanin 2 | 319 | 0.384 | 0.3627 | Yes |
| 9 | PQLC1 | PQLC1 Entrez, Source | PQ loop repeat containing 1 | 444 | 0.312 | 0.3741 | Yes |
| 10 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 458 | 0.303 | 0.3932 | Yes |
| 11 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 474 | 0.294 | 0.4116 | Yes |
| 12 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 480 | 0.292 | 0.4307 | Yes |
| 13 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 484 | 0.291 | 0.4498 | Yes |
| 14 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 724 | 0.231 | 0.4471 | Yes |
| 15 | CD59 | CD59 Entrez, Source | CD59 molecule, complement regulatory protein | 818 | 0.210 | 0.4540 | Yes |
| 16 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 823 | 0.209 | 0.4676 | Yes |
| 17 | ITGA9 | ITGA9 Entrez, Source | integrin, alpha 9 | 923 | 0.192 | 0.4729 | Yes |
| 18 | SDS | SDS Entrez, Source | serine dehydratase | 1007 | 0.182 | 0.4787 | Yes |
| 19 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 1052 | 0.177 | 0.4871 | Yes |
| 20 | HEXA | HEXA Entrez, Source | hexosaminidase A (alpha polypeptide) | 1133 | 0.167 | 0.4922 | Yes |
| 21 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1157 | 0.164 | 0.5013 | Yes |
| 22 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 1203 | 0.161 | 0.5086 | Yes |
| 23 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 1379 | 0.144 | 0.5050 | Yes |
| 24 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 1411 | 0.142 | 0.5121 | Yes |
| 25 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 1427 | 0.140 | 0.5203 | Yes |
| 26 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 1501 | 0.135 | 0.5237 | Yes |
| 27 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 1736 | 0.120 | 0.5140 | No |
| 28 | LRP3 | LRP3 Entrez, Source | low density lipoprotein receptor-related protein 3 | 1822 | 0.116 | 0.5153 | No |
| 29 | SFXN5 | SFXN5 Entrez, Source | sideroflexin 5 | 2105 | 0.104 | 0.5009 | No |
| 30 | KBTBD10 | KBTBD10 Entrez, Source | kelch repeat and BTB (POZ) domain containing 10 | 2314 | 0.095 | 0.4915 | No |
| 31 | HP | HP Entrez, Source | haptoglobin | 2342 | 0.094 | 0.4957 | No |
| 32 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 2449 | 0.090 | 0.4936 | No |
| 33 | RBP1 | RBP1 Entrez, Source | retinol binding protein 1, cellular | 2591 | 0.084 | 0.4885 | No |
| 34 | ADORA3 | ADORA3 Entrez, Source | adenosine A3 receptor | 2617 | 0.083 | 0.4922 | No |
| 35 | APOE | APOE Entrez, Source | apolipoprotein E | 2619 | 0.083 | 0.4976 | No |
| 36 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 2686 | 0.081 | 0.4980 | No |
| 37 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2930 | 0.073 | 0.4845 | No |
| 38 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 3040 | 0.070 | 0.4809 | No |
| 39 | SCGB3A1 | SCGB3A1 Entrez, Source | secretoglobin, family 3A, member 1 | 3058 | 0.069 | 0.4842 | No |
| 40 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 3107 | 0.068 | 0.4852 | No |
| 41 | NPTN | NPTN Entrez, Source | neuroplastin | 3133 | 0.068 | 0.4878 | No |
| 42 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 3208 | 0.065 | 0.4865 | No |
| 43 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 3449 | 0.059 | 0.4723 | No |
| 44 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 4063 | 0.043 | 0.4289 | No |
| 45 | SULT1C1 | SULT1C1 Entrez, Source | sulfotransferase family, cytosolic, 1C, member 1 | 4188 | 0.040 | 0.4222 | No |
| 46 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 4354 | 0.037 | 0.4122 | No |
| 47 | DNAH11 | DNAH11 Entrez, Source | dynein, axonemal, heavy chain 11 | 4390 | 0.036 | 0.4119 | No |
| 48 | CEACAM1 | CEACAM1 Entrez, Source | carcinoembryonic antigen-related cell adhesion molecule 1 (biliary glycoprotein) | 4467 | 0.035 | 0.4085 | No |
| 49 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 4578 | 0.032 | 0.4023 | No |
| 50 | CORO2A | CORO2A Entrez, Source | coronin, actin binding protein, 2A | 4794 | 0.028 | 0.3879 | No |
| 51 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 4845 | 0.027 | 0.3860 | No |
| 52 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 5173 | 0.021 | 0.3627 | No |
| 53 | SLC16A10 | SLC16A10 Entrez, Source | solute carrier family 16, member 10 (aromatic amino acid transporter) | 5484 | 0.015 | 0.3403 | No |
| 54 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 5792 | 0.009 | 0.3177 | No |
| 55 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 5890 | 0.007 | 0.3108 | No |
| 56 | OLIG1 | OLIG1 Entrez, Source | oligodendrocyte transcription factor 1 | 6022 | 0.005 | 0.3013 | No |
| 57 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 6245 | 0.001 | 0.2845 | No |
| 58 | DPP4 | DPP4 Entrez, Source | dipeptidyl-peptidase 4 (CD26, adenosine deaminase complexing protein 2) | 6342 | -0.001 | 0.2773 | No |
| 59 | MMP8 | MMP8 Entrez, Source | matrix metallopeptidase 8 (neutrophil collagenase) | 6500 | -0.004 | 0.2657 | No |
| 60 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 6724 | -0.008 | 0.2494 | No |
| 61 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 7364 | -0.019 | 0.2024 | No |
| 62 | GGTA1 | GGTA1 Entrez, Source | glycoprotein, alpha-galactosyltransferase 1 | 7462 | -0.021 | 0.1965 | No |
| 63 | CENTB2 | CENTB2 Entrez, Source | centaurin, beta 2 | 7595 | -0.023 | 0.1881 | No |
| 64 | CAMK2G | CAMK2G Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma | 7620 | -0.024 | 0.1878 | No |
| 65 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 7645 | -0.024 | 0.1877 | No |
| 66 | CLTC | CLTC Entrez, Source | clathrin, heavy chain (Hc) | 7654 | -0.025 | 0.1887 | No |
| 67 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 7676 | -0.025 | 0.1888 | No |
| 68 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 7733 | -0.026 | 0.1862 | No |
| 69 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 7794 | -0.027 | 0.1835 | No |
| 70 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 7859 | -0.028 | 0.1806 | No |
| 71 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 7870 | -0.029 | 0.1817 | No |
| 72 | ZFAND3 | ZFAND3 Entrez, Source | zinc finger, AN1-type domain 3 | 7940 | -0.030 | 0.1785 | No |
| 73 | SCN11A | SCN11A Entrez, Source | sodium channel, voltage-gated, type XI, alpha | 8099 | -0.033 | 0.1688 | No |
| 74 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 8218 | -0.036 | 0.1622 | No |
| 75 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 8330 | -0.038 | 0.1564 | No |
| 76 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 8458 | -0.041 | 0.1495 | No |
| 77 | CEBPE | CEBPE Entrez, Source | CCAAT/enhancer binding protein (C/EBP), epsilon | 8579 | -0.043 | 0.1433 | No |
| 78 | SYNJ2 | SYNJ2 Entrez, Source | synaptojanin 2 | 8657 | -0.045 | 0.1404 | No |
| 79 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 8752 | -0.047 | 0.1364 | No |
| 80 | MLL5 | MLL5 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia 5 (trithorax homolog, Drosophila) | 9119 | -0.054 | 0.1124 | No |
| 81 | NCOR1 | NCOR1 Entrez, Source | nuclear receptor co-repressor 1 | 9668 | -0.067 | 0.0755 | No |
| 82 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 9927 | -0.075 | 0.0609 | No |
| 83 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 10293 | -0.084 | 0.0390 | No |
| 84 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 10344 | -0.086 | 0.0409 | No |
| 85 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 10385 | -0.087 | 0.0436 | No |
| 86 | CCDC52 | CCDC52 Entrez, Source | coiled-coil domain containing 52 | 10386 | -0.087 | 0.0494 | No |
| 87 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 10665 | -0.096 | 0.0348 | No |
| 88 | LRRFIP2 | LRRFIP2 Entrez, Source | leucine rich repeat (in FLII) interacting protein 2 | 10707 | -0.097 | 0.0382 | No |
| 89 | CSTB | CSTB Entrez, Source | cystatin B (stefin B) | 10747 | -0.099 | 0.0418 | No |
| 90 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 11560 | -0.132 | -0.0108 | No |
| 91 | GGA1 | GGA1 Entrez, Source | golgi associated, gamma adaptin ear containing, ARF binding protein 1 | 11668 | -0.137 | -0.0098 | No |
| 92 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 11683 | -0.137 | -0.0018 | No |
| 93 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 11835 | -0.147 | -0.0034 | No |
| 94 | PAPD4 | PAPD4 Entrez, Source | PAP associated domain containing 4 | 12288 | -0.178 | -0.0257 | No |
| 95 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 12638 | -0.216 | -0.0377 | No |
| 96 | EDD1 | EDD1 Entrez, Source | E3 ubiquitin protein ligase, HECT domain containing, 1 | 12711 | -0.226 | -0.0282 | No |
| 97 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12737 | -0.231 | -0.0147 | No |
| 98 | TMEM37 | TMEM37 Entrez, Source | transmembrane protein 37 | 12955 | -0.278 | -0.0126 | No |
| 99 | RNF141 | RNF141 Entrez, Source | ring finger protein 141 | 13018 | -0.297 | 0.0024 | No |
| 100 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 13108 | -0.331 | 0.0177 | No |