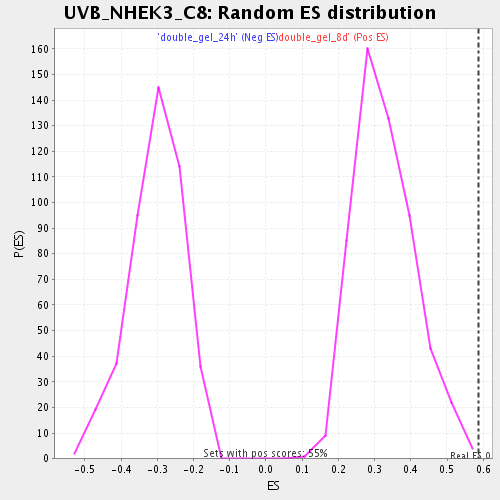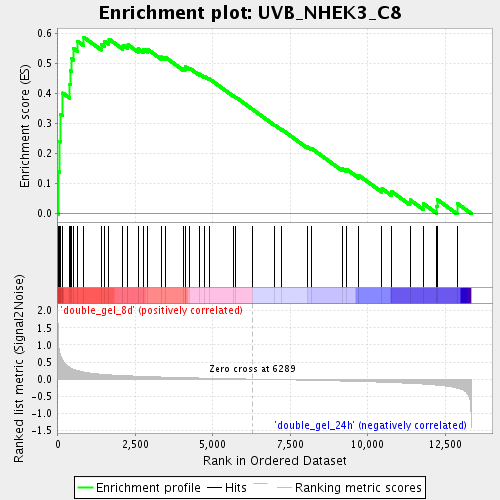
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | UVB_NHEK3_C8 |
| Enrichment Score (ES) | 0.5868601 |
| Normalized Enrichment Score (NES) | 1.7904748 |
| Nominal p-value | 0.0018115942 |
| FDR q-value | 0.030503485 |
| FWER p-Value | 0.956 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 21 | 1.039 | 0.1398 | Yes |
| 2 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 61 | 0.759 | 0.2402 | Yes |
| 3 | PTPRZ1 | PTPRZ1 Entrez, Source | protein tyrosine phosphatase, receptor-type, Z polypeptide 1 | 88 | 0.682 | 0.3310 | Yes |
| 4 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 156 | 0.554 | 0.4014 | Yes |
| 5 | HK1 | HK1 Entrez, Source | hexokinase 1 | 385 | 0.345 | 0.4312 | Yes |
| 6 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 401 | 0.337 | 0.4759 | Yes |
| 7 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 442 | 0.313 | 0.5155 | Yes |
| 8 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 507 | 0.282 | 0.5491 | Yes |
| 9 | GJB2 | GJB2 Entrez, Source | gap junction protein, beta 2, 26kDa (connexin 26) | 628 | 0.250 | 0.5741 | Yes |
| 10 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 834 | 0.207 | 0.5869 | Yes |
| 11 | PON2 | PON2 Entrez, Source | paraoxonase 2 | 1402 | 0.143 | 0.5637 | No |
| 12 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 1507 | 0.134 | 0.5741 | No |
| 13 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 1647 | 0.125 | 0.5807 | No |
| 14 | FASN | FASN Entrez, Source | fatty acid synthase | 2099 | 0.104 | 0.5609 | No |
| 15 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 2261 | 0.097 | 0.5620 | No |
| 16 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 2593 | 0.084 | 0.5485 | No |
| 17 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 2756 | 0.078 | 0.5470 | No |
| 18 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 2893 | 0.074 | 0.5469 | No |
| 19 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 3344 | 0.061 | 0.5214 | No |
| 20 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3479 | 0.058 | 0.5193 | No |
| 21 | PTPRF | PTPRF Entrez, Source | protein tyrosine phosphatase, receptor type, F | 4050 | 0.043 | 0.4823 | No |
| 22 | GLG1 | GLG1 Entrez, Source | golgi apparatus protein 1 | 4112 | 0.042 | 0.4834 | No |
| 23 | FDFT1 | FDFT1 Entrez, Source | farnesyl-diphosphate farnesyltransferase 1 | 4118 | 0.042 | 0.4888 | No |
| 24 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 4257 | 0.038 | 0.4836 | No |
| 25 | ST6GALNAC2 | ST6GALNAC2 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 2 | 4568 | 0.033 | 0.4647 | No |
| 26 | THBS2 | THBS2 Entrez, Source | thrombospondin 2 | 4738 | 0.029 | 0.4560 | No |
| 27 | PRDX6 | PRDX6 Entrez, Source | peroxiredoxin 6 | 4883 | 0.026 | 0.4488 | No |
| 28 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 5660 | 0.012 | 0.3920 | No |
| 29 | R3HDM1 | R3HDM1 Entrez, Source | R3H domain containing 1 | 5745 | 0.010 | 0.3871 | No |
| 30 | PFN2 | PFN2 Entrez, Source | profilin 2 | 6281 | 0.000 | 0.3469 | No |
| 31 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 7002 | -0.012 | 0.2944 | No |
| 32 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 7229 | -0.017 | 0.2797 | No |
| 33 | PGM1 | PGM1 Entrez, Source | phosphoglucomutase 1 | 8051 | -0.032 | 0.2223 | No |
| 34 | TMED10 | TMED10 Entrez, Source | transmembrane emp24-like trafficking protein 10 (yeast) | 8177 | -0.035 | 0.2177 | No |
| 35 | TOMM20 | TOMM20 Entrez, Source | translocase of outer mitochondrial membrane 20 homolog (yeast) | 9181 | -0.056 | 0.1498 | No |
| 36 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 9312 | -0.058 | 0.1480 | No |
| 37 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 9713 | -0.068 | 0.1272 | No |
| 38 | FBN2 | FBN2 Entrez, Source | fibrillin 2 (congenital contractural arachnodactyly) | 10457 | -0.089 | 0.0835 | No |
| 39 | PTPRK | PTPRK Entrez, Source | protein tyrosine phosphatase, receptor type, K | 10767 | -0.100 | 0.0738 | No |
| 40 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 11364 | -0.123 | 0.0458 | No |
| 41 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 11795 | -0.144 | 0.0330 | No |
| 42 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 12221 | -0.173 | 0.0246 | No |
| 43 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 12249 | -0.175 | 0.0464 | No |
| 44 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 12884 | -0.262 | 0.0344 | No |